457:. The distinction between the monoderm and diderm bacteria is supported by conserved signature indels in a number of important proteins (viz. DnaK, GroEL). Of these two structurally distinct groups of bacteria, monoderms are indicated to be ancestral. Based upon a number of observations including that the gram-positive bacteria are the major producers of antibiotics and that, in general, gram-negative bacteria are resistant to them, it has been proposed that the outer cell membrane in gram-negative bacteria (diderms) has evolved as a protective mechanism against
165:
272:
467:, which stain gram-positive due to the presence of a thick peptidoglycan layer and also possess an outer cell membrane are suggested as intermediates in the transition between monoderm (gram-positive) and diderm (gram-negative) bacteria. The diderm bacteria can also be further differentiated between simple diderms lacking lipopolysaccharide, the archetypical diderm bacteria where the outer cell membrane contains lipopolysaccharide, and the diderm bacteria where outer cell membrane is made up of
2536:
433:
necessarily coalesce for some bacterial species. The gram-positive and gram-negative staining response is also not a reliable characteristic as these two kinds of bacteria do not form phylogenetic coherent groups. However, although Gram staining response is an empirical criterion, its basis lies in the marked differences in the ultrastructure and chemical composition of the bacterial cell wall, marked by the absence or presence of an outer lipid membrane.
628:
157:
384:
46:
2546:
20:
1924:
418:
262:, Gram staining is a rapid method used to differentiate bacterial species. Such staining, together with growth requirement and antibiotic susceptibility testing, and other macroscopic and physiologic tests, forms a basis for practical classification and subdivision of the bacteria (e.g., see figure and pre-1990 versions of
448:
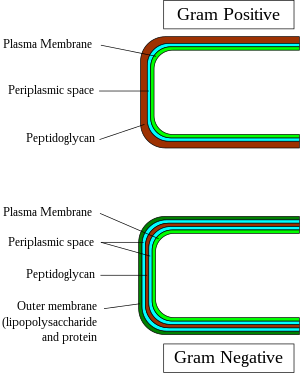
In contrast to gram-positive bacteria, all typical gram-negative bacteria are bounded by a cytoplasmic membrane and an outer cell membrane; they contain only a thin layer of peptidoglycan (2–3 nm) between these membranes. The presence of inner and outer cell membranes defines a new compartment
436:
All gram-positive bacteria are bounded by a single-unit lipid membrane, and, in general, they contain a thick layer (20–80 nm) of peptidoglycan responsible for retaining the Gram stain. A number of other bacteria—that are bounded by a single membrane, but stain gram-negative due to either lack
432:
Although bacteria are traditionally divided into two main groups, gram-positive and gram-negative, based on their Gram stain retention property, this classification system is ambiguous as it refers to three distinct aspects (staining result, envelope organization, taxonomic group), which do not
1381:"Negativicoccus succinicivorans gen. Nov., sp. Nov., isolated from human clinical samples, emended description of the family Veillonellaceae and description of Negativicutes classis nov., Selenomonadales ord. nov. and Acidaminococcaceae fam. nov. In the bacterial phylum Firmicutes"
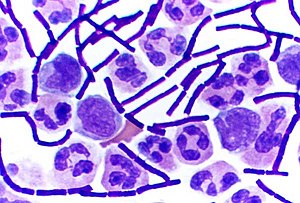
1813:"Analysis of Whole-Genome Sequences of Pathogenic Gram-Positive and Gram-Negative Isolates from the Same Hospital Environment to Investigate Common Evolutionary Trends Associated with Horizontal Gene Exchange, Mutations and DNA Methylation Patterning"
116:
used in this stage degrades the outer membrane of gram-negative cells, making the cell wall more porous and incapable of retaining the crystal violet stain. Their peptidoglycan layer is much thinner and sandwiched between an
441:, or their inability to retain the Gram stain because of their cell wall composition—also show close relationship to the gram-positive bacteria. For the bacterial cells bounded by a single cell membrane, the term
619:)-containing gram-negative bacterial phyla provides evidence that these phyla of bacteria form a monophyletic clade and that no loss of the outer membrane from any species from this group has occurred.
773:; the number might be an overestimate since several of the reports are supported by single papers. Transformation among gram-positive bacteria has been studied in medically important species such as
1928:
1269:
Gupta, R.S. (1998). "What are archaebacteria: life's third domain or monoderm prokaryotes related to Gram-positive bacteria? A new proposal for the classification of prokaryotic organisms".
766:
virus into a recipient host bacterium). In transformation, the genetic material passes through the intervening medium, and uptake is completely dependent on the recipient bacterium.
967:
1430:
sp. nov., isolated from fallen leaves on geothermal soils, and description of
Thermogemmatisporaceae fam. nov. and Thermogemmatisporales ord. Nov. Within the class Ktedonobacteria"
86:
The Gram stain is used by microbiologists to place bacteria into two main categories, Gram-positive (+) and Gram-negative (-). Gram-positive bacteria have a thick layer of
547:) that are either part of the phylum Bacillota or branch in its proximity are found to possess a diderm cell structure. However, a conserved signature indel (CSI) in the
4487:
3971:
1327:"Origin of diderm (gram-negative) bacteria: antibiotic selection pressure rather than endosymbiosis likely led to the evolution of bacterial cells with two membranes"
4423:
4063:
3335:
4248:
3455:
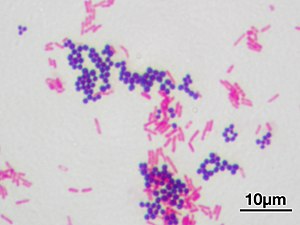
3435:
2919:
3887:
3788:
1768:
Johnston, C.; Martin, B.; Fichant, G.; Polard, P; Claverys, J.P. (2014). "Bacterial transformation: distribution, shared mechanisms and divergent control".
3781:
3762:
3589:
3509:
3288:
1187:
Desvaux, M.; Hébraud, M.; Talon, R.; Henderson, I.R. (2009). "Secretion and subcellular localizations of bacterial proteins: A semantic awareness issue".
250:
in the cell wall. Some of these are lipoteichoic acids, which have a lipid component in the cell membrane that can assist in anchoring the peptidoglycan.
4349:
4175:
4151:
4122:
4009:
3933:
3817:
3795:
3036:
4055:
3899:
3521:
3484:
3462:
3371:
3359:
4255:
4137:
4031:
3347:
3276:
2870:
2833:
2781:
2582:
3921:
3750:
3540:
3502:
4480:
4430:
3872:
3802:
3414:
1514:"Investigation of Candidate Division TM7, a Recently Recognized Major Lineage of the Domain Bacteria with No Known Pure-Culture Representatives"
4411:
4337:
3769:
242:. In gram-positive bacteria, the S-layer is attached to the peptidoglycan layer. Gram-negative bacteria's S-layer is attached directly to the
238:
rings to support them, whereas gram-negative have four. Both gram-positive and gram-negative bacteria commonly have a surface layer called an
4446:
4163:
3955:
3860:
3552:
3402:
3386:
2948:
2724:
1065:"Protein phylogenies and signature sequences: A reappraisal of evolutionary relationships among archaebacteria, eubacteria and eukaryotes"
4908:
318:
of the gram-positive bacteria was challenged, with major implications for the therapeutic and general study of these organisms. Based on
1950:
4473:
4110:
3577:
1955:
1167:
841:
3113:
769:
As of 2014 about 80 species of bacteria were known to be capable of transformation, about evenly divided between gram-positive and
721:
are three gram-positive genera that cause plant disease. Gram-positive bacteria are capable of causing serious and sometimes fatal
1933:
4764:
2575:
1635:
750:, in which exogenous genetic material passes from a donor bacterium to a recipient bacterium, the other two processes being
4898:
4454:
2475:
1704:
Lau, S.K.P.; McNabb, A.; Woo, G.K.S.; Hoang, L.; Fung, A.M.Y.; Chung, L.M.W.; Woo, P.C.Y.; Yuen, K.-Y. (22 November 2006).
4788:
1469:
Sutcliffe, I.C. (2011). "Cell envelope architecture in the
Chloroflexi: A shifting frontline in a phylogenetic turf war".
1811:
Korotetskiy I, Shilov S, Kuznetsova T, Kerimzhanova B, Korotetskaya N, Ivanova L, Zubenko N, Parenova R, Reva O (2023).
2242:
1982:
1379:
Marchandin, H.; Teyssier, C.; Campos, J.; Jean-Pierre, H.; Roger, F.; Gay, B.; Carlier, J.-P.; Jumas-Bilak, E. (2009).
888:
5134:
4515:
4465:
1687:
500:
79:
test, which is traditionally used to quickly classify bacteria into two broad categories according to their type of
2620:
2568:
259:
1706:"Catabacter hongkongensis gen. nov., sp. nov., Isolated from Blood Cultures of Patients from Hong Kong and Canada"
5310:
4739:
4719:
4687:
4668:
2615:
2232:
2159:
370:
genera. The (low G + C) Bacillota, have a 45–60% GC content, but this is lower than that of the
Actinomycetota.
2356:
2087:
698:
4642:
2699:
2689:
2672:
510:
have gram-positive stains, although they are structurally similar to gram-negative bacteria with two layers.
5277:
4951:
4560:
517:
have a single layer, yet (with some exceptions) stain negative. Two related phyla to the
Chloroflexi, the
4780:
4659:
4937:
2197:
599:, etc.) from these other atypical diderm bacteria, as well as other phyla of monoderm bacteria (e.g.,
4692:
3104:
2549:
1569:
Cavaletti, L.; Monciardini, P.; Bamonte, R.; Schumann, P.; Rohde, M.; Sosio, M.; Donadio, S. (2006).
743:
1138:
5172:
5166:
4873:
4733:
4579:
2351:
2274:
2237:
2033:
1862:
Michod, R.E.; Bernstein, H.; Nedelcu, A.M. (2008). "Adaptive value of sex in microbial pathogens".
775:
747:
727:
641:
In the classical sense, six gram-positive genera are typically pathogenic in humans. Two of these,
402:
243:
122:
5123:
4834:
4629:
4267:
2704:
793:
759:
319:
140:
Despite their thicker peptidoglycan layer, gram-positive bacteria are more receptive to certain
5229:
5022:
4681:
4636:
4585:
2427:
2346:
1975:
1133:
770:
311:
279:
109:
1225:
Sutcliffe, I.C. (2010). "A phylum level perspective on bacterial cell envelope architecture".
105:
after it is washed away from the rest of the sample, in the decolorization stage of the test.
5320:
5250:
5243:
5082:
4892:
4828:
4603:
2667:
2467:
2109:
787:
751:
725:
in newborn infants. Novel species of clinically relevant gram-positive bacteria also include
1118:
5256:
5235:
4903:
4842:
4622:
4615:
4318:
3253:
2307:
2182:
2165:
2003:
1582:
1525:
1478:
781:
690:
426:
141:
101:. This is because the thick layer of peptidoglycan in the bacterial cell wall retains the
8:
4798:
4758:
4608:
4511:
4324:
4079:
3258:
2432:
2317:
2312:
2205:
2175:
2028:
2013:
862:
821:
422:
392:
118:
113:
30:
1586:
1529:
1482:
921:
825:
5096:
4979:
4373:
2684:
2662:
2650:
2625:
2560:
2462:
2361:
2170:
2082:
2075:
1943:
1839:
1812:
1793:
1740:
1705:
1603:
1570:
1351:
1326:
1294:
1159:
938:
905:
722:
616:
264:
98:
25:
4495:
1897:
1025:
1000:
525:
Some
Bacillota species are not gram-positive. The class Negativicutes, which includes
483:
whereas gram-negative bacteria are diderms and have two bilayers. Exceptions include:
5315:
5268:
4999:
4823:
4793:
3671:
2694:
2657:
2645:
2635:
2539:
2402:
2368:
2335:
2192:
2187:
2145:
2092:
1968:
1879:
1844:
1785:
1745:
1727:
1683:
1631:
1608:
1551:
1546:
1513:
1494:
1490:
1451:
1402:
1356:
1286:
1282:
1242:
1204:
1151:
1094:
1089:
1080:
1064:
1030:
943:
925:
884:
799:
706:
584:
531:, are diderm and stain gram-negative. Additionally, a number of bacterial taxa (viz.
496:
450:
303:
219:
201:
200:
Peptidoglycan chains are cross-linked to form rigid cell walls by a bacterial enzyme
190:
90:
within the cell wall, and Gram-negative bacteria have a thin layer of peptidoglycan.
1797:
1298:
1163:
373:
5220:
4329:
4087:
4042:
3995:
3833:
2999:
2609:
2254:
2130:
2065:
1951:
3D structures of proteins associated with plasma membrane of gram-positive bacteria
1871:
1834:
1829:
1824:
1777:
1735:
1717:
1598:
1590:
1541:
1533:
1486:
1441:
1392:
1346:
1338:
1278:
1234:
1196:
1143:
1084:
1076:
1020:
1016:
1012:
976:
933:
917:
164:
97:
stain used in the test, and then appear to be purple-coloured when seen through an
57:
34:
1956:
3D structures of proteins associated with outer membrane of gram-positive bacteria
1537:
4993:
4707:
4534:
4380:
4284:
4097:
4092:
3846:
3648:
3626:
3441:
3154:
3125:
3075:
2910:
2805:
2773:
2480:
2379:
2264:
2140:
2070:
2023:
1875:
1667:. The First Iranian International Congress of Medical Bacteriology. Tabriz, Iran.
667:
596:
588:
544:
454:
348:
323:
172:
In general, the following characteristics are present in gram-positive bacteria:
102:
615:, etc.). The presence of this CSI in all sequenced species of conventional LPS (
5151:
4360:
4299:
4290:
4187:
4143:
3951:
3914:
3828:
3720:
3661:
3164:
3141:
2844:
2757:
2744:
2739:
2640:
2135:
2099:
689:
produce spores. The spore-forming bacteria can again be divided based on their
649:
600:
580:
556:
555:) protein distinguishes all traditional phyla of gram-negative bacteria (e.g.,
536:
339:
287:
271:
223:
186:
94:
38:
1512:
Hugenholtz, P.; Tyson, G.W.; Webb, R.I.; Wagner, A.M.; Blackall, L.L. (2001).
1342:
1238:
1200:
1147:
5304:
4697:
4550:
4366:
4020:
3739:
3679:
3656:
3595:
3382:
3265:
3243:
3212:
3186:
3087:
2901:
2792:
2768:
2749:
2720:
2489:
2374:
2330:
2302:
2293:
2283:
2259:
2155:
1731:
1571:"New Lineage of Filamentous, Spore-Forming, Gram-Positive Bacteria from Soil"
929:
763:
643:
612:
592:
576:
532:
514:
492:
480:
354:
247:
180:
87:
54:
981:
962:
326:. Two of these were gram-positive and were divided on the proportion of the
5129:
4814:
4226:
4216:
3940:
3732:
3704:
3613:
3533:
3469:
3324:
3299:
3222:
2818:
2800:
2519:
2504:
2384:
2125:
2043:
1991:
1883:
1848:
1789:
1749:
1612:
1555:
1498:
1455:
1446:
1421:
1406:
1397:
1380:
1360:
1246:
1208:
1155:
947:
608:
568:
540:
507:
468:
366:
295:
126:
64:
1290:
1098:
1034:
5283:
5058:
5028:
4198:
4102:
3709:
3636:
2407:
1960:
1722:
1594:
837:
685:
633:
572:
564:
527:
463:
145:
1781:
5211:
5196:
5185:
5048:
4929:
4857:
4702:
4676:
4502:
4311:
4203:
3982:
3714:
3641:
2978:
2944:
2824:
2591:
2514:
2499:
2417:
2150:
1810:
732:
560:
488:
458:
453:
or the periplasmic compartment. These bacteria have been designated as
438:
397:
307:
299:
235:
227:
148:
than gram-negative bacteria, due to the absence of the outer membrane.
76:
904:
Brown, Stephanie; Santa Maria, John P.; Walker, Suzanne (2013-09-08).
302:(neutral in staining) and Mendocutes (variable in staining). Based on
5156:
5063:
4920:
4883:
4728:
4506:
4498:
3694:
2960:
2509:
2340:
2298:
2210:
2104:
662:
627:
604:
343:
315:
291:
211:
194:
80:
19:
5088:
5012:
4969:
4862:
4569:
4539:
2679:
2595:
2018:
1995:
1568:
1378:
679:
673:
479:
In general, gram-positive bacteria are monoderms and have a single
360:
331:
231:
134:
130:
72:
1680:
Avery's
Neonatology: Pathophysiology and Management of the Newborn
661:(rod-shaped) and can be subdivided based on their ability to form
5102:
4985:
4525:
2966:
2448:
2412:
2048:
1434:
International
Journal of Systematic and Evolutionary Microbiology
1385:
International
Journal of Systematic and Evolutionary Microbiology
968:
International
Journal of Systematic and Evolutionary Microbiology
755:
658:
374:
Importance of the outer cell membrane in bacterial classification
327:
239:
156:
45:
2494:
2249:
654:
283:
1420:
Yabe, S.; Aiba, Y.; Sakai, Y.; Hazaka, M.; Yokota, A. (2010).
112:
cannot retain the violet stain after the decolorization step;
2422:
1651:
Sahebnasagh, R.; Saderi, H.; Owlia, P. (4–7 September 2011).
552:
548:
50:
1657:
strains from clinical samples in Tehran by detection of the
1186:
1939:
1767:
16:
Bacteria that give a positive result in the Gram stain test
1119:"The natural evolutionary relationships among prokaryotes"
417:
1511:
631:
Colonies of a gram-positive pathogen of the oral cavity,
518:
335:
2590:
246:. Specific to gram-positive bacteria is the presence of
1861:
1650:
1182:
1180:
903:
275:
Species identification hierarchy in clinical settings
1419:
1177:
758:between two bacterial cells in direct contact) and
160:
Gram-positive and gram-negative cell wall structure
963:"Proposals Concerning the Higher Taxa of Bacteria"
521:clade and the Ktedonobacteria, are also monoderms.
1703:
1220:
1218:
5302:
1694:Access provided by the University of Pittsburgh.
487:Some taxa lack peptidoglycan (such as the class
306:phylogenetic studies of the late microbiologist
197:agents, and also for certain types of adherence.
1625:
1374:
1372:
1370:
906:"Wall Teichoic Acids of Gram-Positive Bacteria"
879:Madigan, Michael T.; Martinko, John M. (2006).
878:
844:), if any, governs the document being written.
1628:Clinical Microbiology Made Ridiculously Simple
1215:
322:of the 16S sequences, Woese recognised twelve
4481:
2576:
1976:
1908:. Centers for Disease Control and Prevention.
1671:
960:
860:
828:, their initial letter can be either capital
731:, which is an emerging pathogen belonging to
657:(sphere-shaped). The remaining organisms are
1367:
994:
992:
421:The structure of peptidoglycan, composed of
1804:
1763:
1761:
1759:
738:
461:selection pressure. Some bacteria, such as
391:It has been suggested that this section be
338:. The high G + C phylum was made up of the
4488:
4474:
2583:
2569:
1990:
1983:
1969:
1320:
1318:
1316:
1314:
1312:
1310:
1308:
1264:
1262:
1260:
1258:
1256:
1069:Microbiology and Molecular Biology Reviews
265:Bergey's Manual of Systematic Bacteriology
1838:
1828:
1739:
1721:
1677:
1602:
1545:
1468:
1445:
1396:
1350:
1224:
1137:
1112:
1110:
1108:
1088:
1058:
1056:
1054:
1052:
1050:
1048:
1046:
1044:
1024:
989:
980:
937:
342:, and the low G + C phylum contained the
1756:
883:(11th ed.). Pearson Prentice Hall.
626:
416:
310:and collaborators and colleagues at the
270:
163:
155:
44:
18:
4765:Cutaneous Streptococcus iniae infection
1305:
1253:
874:
872:
762:(injection of donor bacterial DNA by a
5303:
1855:
1630:. Miami, FL: MedMaster. pp. 4–5.
1626:Gladwin, Mark; Trattler, Bill (2007).
1575:Applied and Environmental Microbiology
1518:Applied and Environmental Microbiology
1105:
1041:
961:Gibbons, N.E.; Murray, R.G.E. (1978).
495:, and the insect-endosymbionts of the
437:of the peptidoglycan layer, as in the
4469:
2564:
1964:
1324:
1268:
1116:
1062:
998:
395:out into another article titled
4899:Staphylococcal scalded skin syndrome
2545:
1682:. Philadelphia, PA: Wolters Kluwer.
869:
807:
797:and in gram-positive soil bacterium
377:
214:than that in gram-negative bacteria.
168:Structure of gram-positive cell wall
1653:Detection of methicillin-resistant
1505:
922:10.1146/annurev-micro-092412-155620
93:Gram-positive bacteria take up the
75:that give a positive result in the
13:
1562:
290:based primarily on Gram staining:
151:
14:
5332:
5135:Clostridial necrotizing enteritis
1916:
1864:Infection, Genetics and Evolution
346:. The Actinomycetota include the
253:
189:and lipoids are present, forming
2544:
2535:
2534:
1927: This article incorporates
1922:
1710:Journal of Clinical Microbiology
1491:10.1111/j.1462-2920.2010.02339.x
1283:10.1046/j.1365-2958.1998.00978.x
1173:from the original on 2013-06-25.
1126:Critical Reviews in Microbiology
1081:10.1128/MMBR.62.4.1435-1491.1998
622:
382:
4740:Group B streptococcal infection
4688:Group A streptococcal infection
2233:Bacterial cellular morphologies
1890:
1697:
1644:
1619:
1462:
1413:
881:Brock Biology of Microorganisms
861:Basic Biology (18 March 2016).
1830:10.3390/microorganisms11020323
1424:Thermogemmatispora onikobensis
1017:10.1128/MMBR.51.2.221-271.1987
954:
897:
854:
746:is one of three processes for
226:. Also, only some species are
125:, causing them to take up the
1:
1538:10.1128/AEM.67.1.411-419.2001
910:Annual Review of Microbiology
847:
474:
49:Violet-stained gram-positive
5278:Erysipelothrix rhusiopathiae
1900:Emerging Infectious Diseases
1876:10.1016/j.meegid.2008.01.002
1770:Nature Reviews. Microbiology
665:. The non-spore formers are
33:sample stand out from round
7:
1428:Thermogemmatispora foliorum
820:derive from the surname of
677:(a coccobacillus), whereas
10:
5337:
2476:Bacteria (classifications)
2198:Primary nutritional groups
1678:MacDonald, Mhairi (2015).
1471:Environmental Microbiology
176:Cytoplasmic lipid membrane
137:) and appear red or pink.
5267:
5219:
5210:
5183:
5149:
5113:
5072:
5056:
5047:
5010:
4967:
4919:
4882:
4871:
4856:
4812:
4779:
4750:
4718:
4693:Streptococcal pharyngitis
4667:
4658:
4596:
4568:
4559:
4548:
4533:
4524:
4440:
4403:
4277:
4078:
3977:"Acidulodesulfobacteriia"
3963:
3950:
3693:
3625:
3567:
3494:
3394:
3381:
3317:
3236:
3205:
3198:
3103:
2943:
2891:
2732:
2719:
2603:
2530:
2461:
2441:
2393:
2282:
2273:
2225:
2118:
2056:
2042:
2002:
1343:10.1007/s10482-011-9616-8
1239:10.1016/j.tim.2010.06.005
1201:10.1016/j.tim.2009.01.004
1148:10.1080/10408410091154219
218:Only some species have a
210:A much smaller volume of
23:Rod-shaped gram-positive
5173:Pseudomembranous colitis
5167:Clostridioides difficile
4234:"Thermodesulfovibrionia"
3972:Acidulodesulfobacteriota
2987:"Thermosediminibacteria"
2925:"Marinamargulisbacteria"
2352:Bacterial outer membrane
1426:gen. nov., sp. nov. And
776:Streptococcus pneumoniae
748:horizontal gene transfer
739:Bacterial transformation
728:Catabacter hongkongensis
298:(negative in staining),
294:(positive in staining),
230:, and when they do have
222:, usually consisting of
123:bacterial outer membrane
37:, which also accept the
5124:Clostridium perfringens
4835:Urinary tract infection
4268:Thermodesulfobacteriota
4069:"Thermosulfidibacteria"
2928:"Riflemargulisbacteria"
1331:Antonie van Leeuwenhoek
1005:Microbiological Reviews
982:10.1099/00207713-28-1-1
794:Streptococcus sanguinis
5311:Gram-positive bacteria
5230:Ureaplasma urealyticum
5023:Listeria monocytogenes
4727:bacitracin resistant,
4586:Pneumococcal infection
4424:Salinosulfoleibacteria
4242:"Lambdaproteobacteria"
4064:Thermosulfidibacterota
3341:"Macinerneyibacteriia"
2984:"Thermoanaerobacteria"
2911:Vampirovibrionophyceae
2347:Gram-negative bacteria
2326:Gram-positive bacteria
1929:public domain material
1447:10.1099/ijs.0.024877-0
1398:10.1099/ijs.0.013102-0
1271:Molecular Microbiology
1227:Trends in Microbiology
1189:Trends in Microbiology
771:gram-negative bacteria
638:
491:, some members of the
429:
312:University of Illinois
286:was divided into four
276:
169:
161:
110:gram-negative bacteria
69:gram-positive bacteria
60:
42:
5251:Mycoplasma pneumoniae
5244:Mycoplasma genitalium
5083:Clostridium botulinum
4829:Enterococcus faecalis
4604:Viridans streptococci
3336:Macinerneyibacteriota
2202:Substrate preference
1655:Staphylococcus aureus
1001:"Bacterial evolution"
836:, depending on which
788:Staphylococcus aureus
630:
420:
274:
167:
159:
48:
22:
5257:Mycoplasma pneumonia
5236:Ureaplasma infection
4946:novobiocin resistant
4904:Toxic shock syndrome
4843:Enterococcus faecium
4319:Polarisedimenticolia
4307:"Guanabaribacteriia"
4249:Schekmaniibacteriota
3990:"Desulfurobacteriia"
3905:"Handelsmanbacteria"
3893:"Krumholzibacteriia"
3456:Firestoneibacteriota
3436:Desantisiibacteriota
3254:Coprothermobacterota
3134:"Absconditabacteria"
3051:"Desulfitobacteriia"
2920:Margulisiibacteriota
2183:Microbial metabolism
1902:Journal Style Guide"
1723:10.1128/jcm.01831-06
1595:10.1128/AEM.00132-06
1325:Gupta, R.S. (2011).
1117:Gupta, R.S. (2000).
1063:Gupta, R.S. (1998).
999:Woese, C.R. (1987).
826:eponymous adjectives
782:Streptococcus mutans
699:facultative anaerobe
449:in these cells: the
427:N-acetylmuramic acid
5189:(non-spore forming)
4799:Streptococcus bovis
4759:Streptococcus iniae
4507:Infectious diseases
4453:Alternative views:
4325:Thermoanaerobaculia
4128:"Anaeroferrophilia"
4080:Deltaproteobacteria
4052:Desulfobacterota G
3915:Neomarinimicrobiota
3888:Krumholzibacteriota
3789:Edwardsiibacteriota
3515:"Hinthialibacteria"
3294:"Thermodesulfobiia"
3259:Coprothermobacteria
3071:"Halanaerobiaeota"
3019:"Thermaerobacteria"
2906:"Sericytochromatia"
2433:Non-motile bacteria
2029:Pathogenic bacteria
1782:10.1038/nrmicro3199
1587:2006ApEnM..72.4360C
1530:2001ApEnM..67..411H
1483:2011EnvMi..13..279S
840:(e.g., that of the
822:Hans Christian Gram
445:has been proposed.
423:N-acetylglucosamine
119:inner cell membrane
31:cerebrospinal fluid
5097:Clostridium tetani
4980:Bacillus anthracis
4597:optochin resistant
4512:Bacterial diseases
4181:"Methylomirabilia"
4131:"Deferrimicrobiia"
4015:"Calescibacteriia"
3841:"Raymondbacteriia"
3823:"Fermentibacteria"
3782:Delongiibacteriota
3763:Coatesiibacteriota
3590:Lindowiibacteriota
3510:Hinthialibacterota
3289:Thermodesulfobiota
3137:"Andersenbacteria"
3063:"Syntrophomonadia"
3054:"Desulfotomaculia"
3045:"Carboxydothermia"
2876:"Eudoremicrobiia"
2861:"Heboniibacteriia"
2362:Lipopolysaccharide
639:
617:lipopolysaccharide
501:gram-indeterminate
430:
277:
191:lipoteichoic acids
170:
162:
99:optical microscope
61:
43:
26:Bacillus anthracis
5298:
5297:
5294:
5293:
5269:Anaeroplasmatales
5206:
5205:
5145:
5144:
5043:
5042:
5039:
5038:
4963:
4962:
4852:
4851:
4808:
4807:
4775:
4774:
4654:
4653:
4463:
4462:
4399:
4398:
4395:
4394:
4350:Leptospirillaeota
4304:"Fischerbacteria"
4176:Methylomirabilota
4152:Desulfuromonadota
4123:Deferrimicrobiota
4047:"Deferribacteria"
4010:Calescibacteriota
4000:"Campylobacteria"
3934:Tianyaibacteriota
3908:"Latescibacteria"
3818:Fermentibacterota
3796:Eiseniibacteriota
3698:(Sphingobacteria)
3672:Verrucomicrobiota
3630:(Planctobacteria)
3527:"Hydrogenedentia"
3429:"Tritonobacteria"
3313:
3312:
3179:"Saccharimonadia"
3160:"Howlettbacteria"
3099:
3098:
3083:"Selenobacteria"
3048:"Dehalobacteriia"
3042:"Carboxydocellia"
3037:Desulfotomaculota
3027:"Hydrogenisporia"
3016:"Symbiobacteriia"
3005:"Proteinivoracia"
2849:"Abditibacteriia"
2762:"Geothermincolia"
2558:
2557:
2457:
2456:
2403:Bacterial capsule
2369:Periplasmic space
2336:Lipoteichoic acid
2221:
2220:
2193:Microbial ecology
2188:Nitrogen fixation
1637:978-0-940780-81-1
808:Orthographic note
802:, Bacillus cereus
800:Bacillus subtilis
707:obligate anaerobe
585:Verrucomicrobiota
497:Enterobacteriales
451:periplasmic space
443:monoderm bacteria
415:
414:
410:
334:content in their
320:molecular studies
304:16S ribosomal RNA
204:
193:, which serve as
53:and pink-stained
35:white blood cells
5328:
5221:Mycoplasmataceae
5217:
5216:
5197:Finegoldia magna
5070:
5069:
5054:
5053:
4952:S. saprophyticus
4880:
4879:
4869:
4868:
4665:
4664:
4566:
4565:
4557:
4556:
4546:
4545:
4531:
4530:
4490:
4483:
4476:
4467:
4466:
4355:"Leptospirillia"
4330:Vicinamibacteria
4261:"Entotheonellia"
4157:Desulfuromonadia
4088:Bdellovibrionota
4056:Syntrophorhabdia
4043:Deferribacterota
3996:Campylobacterota
3961:
3960:
3900:Latescibacterota
3834:Chitinivibrionia
3811:"Tariuqbacteria"
3684:Verrucomicrobiia
3522:Hydrogenedentota
3485:Ratteibacteriota
3478:"Velamenicoccia"
3463:Goldiibacteriota
3426:"Erginobacteria"
3420:"Ancaeobacteria"
3392:
3391:
3372:Walliibacteriota
3360:Rifleibacteriota
3353:"Muiribacteriia"
3282:"Lithacetigenia"
3203:
3202:
3173:"Microgenomatia"
3150:"Doudnabacteria"
2995:"Dethiobacteria"
2954:
2934:"Termititenacia"
2898:Cyanobacteriota
2882:"Eremiobacteria"
2813:"Limnocylindria"
2787:"Bipolaricaulia"
2730:
2729:
2632:
2612:
2585:
2578:
2571:
2562:
2561:
2548:
2547:
2538:
2537:
2486:Former groupings
2280:
2279:
2131:Human microbiome
2054:
2053:
1985:
1978:
1971:
1962:
1961:
1947:
1942:. Archived from
1926:
1925:
1910:
1909:
1894:
1888:
1887:
1859:
1853:
1852:
1842:
1832:
1808:
1802:
1801:
1765:
1754:
1753:
1743:
1725:
1701:
1695:
1693:
1675:
1669:
1668:
1648:
1642:
1641:
1623:
1617:
1616:
1606:
1581:(6): 4360–4369.
1566:
1560:
1559:
1549:
1509:
1503:
1502:
1466:
1460:
1459:
1449:
1417:
1411:
1410:
1400:
1391:(6): 1271–1279.
1376:
1365:
1364:
1354:
1322:
1303:
1302:
1266:
1251:
1250:
1222:
1213:
1212:
1184:
1175:
1174:
1172:
1141:
1123:
1114:
1103:
1102:
1092:
1075:(4): 1435–1491.
1060:
1039:
1038:
1028:
996:
987:
986:
984:
958:
952:
951:
941:
901:
895:
894:
876:
867:
866:
858:
756:genetic material
406:
386:
385:
378:
234:, have only two
202:
5336:
5335:
5331:
5330:
5329:
5327:
5326:
5325:
5301:
5300:
5299:
5290:
5263:
5202:
5179:
5141:
5109:
5035:
5006:
4994:Bacillus cereus
4959:
4915:
4860:
4848:
4804:
4771:
4746:
4714:
4708:Rheumatic fever
4650:
4592:
4537:
4535:Lactobacillales
4520:
4494:
4464:
4459:
4447:Bergey's Manual
4436:
4391:
4386:"Pseudomonadia"
4296:"Aminicenantia"
4285:Acidobacteriota
4273:
4256:Tectimicrobiota
4138:Deferrisomatota
4098:Bdellovibrionia
4093:Bacteriovoracia
4074:
4032:Dadaibacteriota
3954:
3946:
3941:Zixiibacteriota
3927:"Marinisomatia"
3881:"Stahlbacteria"
3854:"Glassbacteria"
3847:Gemmatimonadota
3756:"Cloacimonadia"
3721:Ignavibacteriia
3697:
3689:
3676:Kiritimatiellia
3666:"Uabimicrobiia"
3649:Planctomycetota
3629:
3621:
3570:bacteriobiontes
3569:
3563:
3546:"Saltatorellae"
3490:
3442:Elusimicrobiota
3385:
3377:
3365:"Ozemibacteria"
3348:Muiribacteriota
3309:
3277:Lithacetigenota
3266:Dictyoglomerota
3232:
3194:
3182:"Torokbacteria"
3176:"Paceibacteria"
3170:"Kazanbacteria"
3155:Gracilibacteria
3147:"Dojkabacteria"
3126:Patescibacteria
3119:"Elulimicrobia"
3095:
3013:"Sulfobacillia"
2952:
2951:
2947:
2939:
2931:"Saganbacteria"
2892:Cyanoprokaryota
2887:
2871:Eremiobacterota
2839:"Dormibacteria"
2834:Dormiibacterota
2810:Ktedonobacteria
2806:Dehalococcoidia
2782:Bipolaricaulota
2774:Thermoleophilia
2765:"Humimicrobiia"
2723:
2715:
2714:
2630:
2608:
2599:
2589:
2559:
2554:
2526:
2481:Bacterial phyla
2465:
2453:
2437:
2395:
2389:
2380:Arabinogalactan
2285:
2269:
2217:
2114:
2058:
2046:
2038:
2024:Lysogenic cycle
2005:
1998:
1989:
1932:
1923:
1919:
1914:
1913:
1896:
1895:
1891:
1860:
1856:
1809:
1805:
1766:
1757:
1702:
1698:
1690:
1676:
1672:
1649:
1645:
1638:
1624:
1620:
1567:
1563:
1510:
1506:
1467:
1463:
1418:
1414:
1377:
1368:
1323:
1306:
1267:
1254:
1233:(10): 464–470.
1223:
1216:
1185:
1178:
1170:
1139:10.1.1.496.1356
1121:
1115:
1106:
1061:
1042:
997:
990:
959:
955:
902:
898:
891:
877:
870:
859:
855:
850:
812:The adjectives
810:
741:
668:Corynebacterium
625:
597:Acidobacteriota
589:Planctomycetota
545:Elusimicrobiota
477:
455:diderm bacteria
411:
408:(November 2023)
387:
383:
376:
349:Corynebacterium
324:bacterial phyla
256:
224:polysaccharides
205:-transpeptidase
154:
152:Characteristics
17:
12:
11:
5:
5334:
5324:
5323:
5318:
5313:
5296:
5295:
5292:
5291:
5289:
5288:
5287:
5286:
5273:
5271:
5265:
5264:
5262:
5261:
5260:
5259:
5247:
5240:
5239:
5238:
5225:
5223:
5214:
5208:
5207:
5204:
5203:
5201:
5200:
5192:
5190:
5181:
5180:
5178:
5177:
5176:
5175:
5162:
5160:
5152:Clostridioides
5147:
5146:
5143:
5142:
5140:
5139:
5138:
5137:
5132:
5119:
5117:
5111:
5110:
5108:
5107:
5106:
5105:
5093:
5092:
5091:
5078:
5076:
5067:
5051:
5045:
5044:
5041:
5040:
5037:
5036:
5034:
5033:
5032:
5031:
5018:
5016:
5008:
5007:
5005:
5004:
5003:
5002:
5000:Food poisoning
4990:
4989:
4988:
4975:
4973:
4965:
4964:
4961:
4960:
4958:
4957:
4956:
4955:
4943:
4942:
4941:
4938:S. epidermidis
4925:
4923:
4917:
4916:
4914:
4913:
4912:
4911:
4906:
4901:
4888:
4886:
4877:
4874:Staphylococcus
4866:
4854:
4853:
4850:
4849:
4847:
4846:
4839:
4838:
4837:
4820:
4818:
4810:
4809:
4806:
4805:
4803:
4802:
4791:
4785:
4783:
4777:
4776:
4773:
4772:
4770:
4769:
4768:
4767:
4754:
4752:
4748:
4747:
4745:
4744:
4743:
4742:
4724:
4722:
4716:
4715:
4713:
4712:
4711:
4710:
4705:
4700:
4695:
4690:
4673:
4671:
4662:
4656:
4655:
4652:
4651:
4649:
4648:
4640:
4633:
4626:
4619:
4612:
4600:
4598:
4594:
4593:
4591:
4590:
4589:
4588:
4575:
4573:
4563:
4554:
4543:
4528:
4522:
4521:
4519:
4518:
4509:
4493:
4492:
4485:
4478:
4470:
4461:
4460:
4458:
4457:
4451:
4441:
4438:
4437:
4435:
4434:
4427:
4420:
4419:
4418:
4407:
4405:
4401:
4400:
4397:
4396:
4393:
4392:
4390:
4389:
4388:
4387:
4384:
4377:
4370:
4361:Pseudomonadota
4358:
4357:
4356:
4346:
4345:
4344:
4334:
4333:
4332:
4327:
4322:
4315:
4308:
4305:
4302:
4300:Blastocatellia
4297:
4294:
4291:Acidobacteriia
4281:
4279:
4275:
4274:
4272:
4271:
4264:
4263:
4262:
4252:
4245:
4244:
4243:
4237:
4236:
4235:
4232:
4224:
4223:
4222:
4214:
4213:
4212:
4209:
4208:"Bradymonadia"
4206:
4196:
4195:
4194:
4193:"Moduliflexia"
4184:
4183:
4182:
4172:
4171:
4170:
4160:
4159:
4158:
4148:
4147:
4146:
4144:Deferrisomatia
4134:
4133:
4132:
4129:
4119:
4118:
4117:
4107:
4106:
4105:
4100:
4095:
4084:
4082:
4076:
4075:
4073:
4072:
4071:
4070:
4060:
4059:
4058:
4050:
4049:
4048:
4040:
4039:
4038:
4037:"Dadabacteria"
4028:
4027:
4026:
4018:
4017:
4016:
4006:
4005:
4004:
4001:
3993:
3992:
3991:
3988:
3980:
3979:
3978:
3967:
3965:
3958:
3952:Proteobacteria
3948:
3947:
3945:
3944:
3937:
3930:
3929:
3928:
3922:Marinisomatota
3918:
3911:
3910:
3909:
3906:
3896:
3895:
3894:
3884:
3883:
3882:
3879:
3878:"Hydrothermia"
3869:
3868:
3867:
3857:
3856:
3855:
3852:
3851:Gemmatimonadia
3844:
3843:
3842:
3839:
3836:
3829:Fibrobacterota
3826:
3825:
3824:
3814:
3813:
3812:
3809:
3808:"Electryoneia"
3799:
3792:
3785:
3778:
3777:
3776:
3775:"Cosmopolitia"
3766:
3759:
3758:
3757:
3751:Cloacimonadota
3747:
3746:
3745:
3737:
3736:
3735:
3730:
3727:
3726:"Kapabacteria"
3724:
3717:
3712:
3701:
3699:
3691:
3690:
3688:
3687:
3686:
3685:
3682:
3677:
3669:
3668:
3667:
3664:
3662:Planctomycetia
3659:
3654:
3646:
3645:
3644:
3633:
3631:
3623:
3622:
3620:
3619:
3618:
3617:
3610:
3607:
3604:
3603:"Brachyspirae"
3601:
3600:"Brevinematia"
3593:
3586:
3585:
3584:
3573:
3571:
3565:
3564:
3562:
3561:
3560:
3559:
3549:
3548:
3547:
3541:Saltatorellota
3537:
3534:Poribacteriota
3530:
3529:
3528:
3518:
3517:
3516:
3506:
3503:Abyssubacteria
3498:
3496:
3492:
3491:
3489:
3488:
3481:
3480:
3479:
3476:
3466:
3459:
3452:
3451:
3450:
3447:
3446:Elusimicrobiia
3439:
3432:
3431:
3430:
3427:
3424:
3423:"Auribacteria"
3421:
3411:
3410:
3409:
3398:
3396:
3389:
3379:
3378:
3376:
3375:
3368:
3367:
3366:
3356:
3355:
3354:
3344:
3343:
3342:
3332:
3331:
3330:
3325:Fusobacteriota
3321:
3319:
3315:
3314:
3311:
3310:
3308:
3307:
3306:
3305:
3297:
3296:
3295:
3285:
3284:
3283:
3273:
3272:
3271:
3270:Dictyoglomeria
3263:
3262:
3261:
3251:
3250:
3249:
3240:
3238:
3234:
3233:
3231:
3230:
3229:
3228:
3220:
3219:
3218:
3209:
3207:
3200:
3196:
3195:
3193:
3192:
3191:
3190:
3183:
3180:
3177:
3174:
3171:
3168:
3165:Katanobacteria
3161:
3158:
3151:
3148:
3145:
3142:Berkelbacteria
3138:
3135:
3132:
3122:
3121:
3120:
3109:
3107:
3101:
3100:
3097:
3096:
3094:
3093:
3092:
3091:
3081:
3080:
3079:
3069:
3068:
3067:
3066:"Thermincolia"
3064:
3061:
3058:
3055:
3052:
3049:
3046:
3043:
3033:
3032:
3031:
3028:
3022:
3021:
3020:
3017:
3014:
3008:
3007:
3006:
3003:
3000:Natranaerobiia
2996:
2990:
2989:
2988:
2985:
2982:
2972:
2971:
2970:
2957:
2955:
2941:
2940:
2938:
2937:
2936:
2935:
2932:
2929:
2926:
2916:
2915:
2914:
2907:
2904:
2895:
2893:
2889:
2888:
2886:
2885:
2884:
2883:
2880:
2874:
2867:
2866:
2865:
2862:
2859:
2858:Fimbriimonadia
2856:
2855:Chthonomonadia
2853:
2850:
2845:Armatimonadota
2842:
2841:
2840:
2830:
2829:
2828:
2816:
2815:
2814:
2811:
2808:
2803:
2798:
2790:
2789:
2788:
2778:
2777:
2776:
2771:
2766:
2763:
2760:
2758:Coriobacteriia
2755:
2754:"Aquicultoria"
2752:
2747:
2745:Acidimicrobiia
2740:Actinomycetota
2736:
2734:
2727:
2717:
2716:
2713:
2712:
2711:
2710:
2709:
2708:
2705:Mesomycetozoea
2702:
2697:
2687:
2677:
2676:
2675:
2670:
2665:
2660:
2655:
2654:
2653:
2641:Diaphoretickes
2638:
2633:
2628:
2623:
2618:
2613:
2605:
2604:
2601:
2600:
2598:classification
2588:
2587:
2580:
2573:
2565:
2556:
2555:
2553:
2552:
2542:
2531:
2528:
2527:
2525:
2524:
2523:
2522:
2517:
2512:
2507:
2497:
2492:
2483:
2478:
2472:
2470:
2459:
2458:
2455:
2454:
2452:
2451:
2445:
2443:
2439:
2438:
2436:
2435:
2430:
2425:
2420:
2415:
2410:
2405:
2399:
2397:
2391:
2390:
2388:
2387:
2382:
2371:
2366:
2365:
2364:
2359:
2343:
2338:
2333:
2322:
2321:
2320:
2315:
2310:
2296:
2290:
2288:
2277:
2271:
2270:
2268:
2267:
2262:
2257:
2252:
2247:
2246:
2245:
2240:
2238:cell structure
2229:
2227:
2223:
2222:
2219:
2218:
2216:
2215:
2214:
2213:
2211:Saccharophilic
2208:
2200:
2195:
2190:
2185:
2180:
2179:
2178:
2173:
2168:
2163:
2153:
2148:
2143:
2138:
2128:
2122:
2120:
2116:
2115:
2113:
2112:
2107:
2102:
2100:Microaerophile
2097:
2096:
2095:
2090:
2080:
2079:
2078:
2073:
2062:
2060:
2051:
2040:
2039:
2037:
2036:
2031:
2026:
2021:
2016:
2010:
2008:
2000:
1999:
1988:
1987:
1980:
1973:
1965:
1959:
1958:
1953:
1948:
1946:on 2009-12-08.
1935:Science Primer
1918:
1917:External links
1915:
1912:
1911:
1889:
1870:(3): 267–285.
1854:
1817:Microorganisms
1803:
1776:(3): 181–196.
1755:
1716:(2): 395–401.
1696:
1688:
1670:
1643:
1636:
1618:
1561:
1524:(1): 411–419.
1504:
1477:(2): 279–282.
1461:
1440:(4): 903–910.
1412:
1366:
1337:(2): 171–182.
1304:
1277:(3): 695–707.
1252:
1214:
1195:(4): 139–145.
1176:
1132:(2): 111–131.
1104:
1040:
1011:(2): 221–271.
988:
953:
916:(1): 313–336.
896:
890:978-0131443297
889:
868:
852:
851:
849:
846:
832:or lower-case
809:
806:
744:Transformation
740:
737:
650:Staphylococcus
624:
621:
601:Actinomycetota
581:Fibrobacterota
557:Pseudomonadota
537:Fusobacteriota
523:
522:
511:
504:
476:
473:
413:
412:
390:
388:
381:
375:
372:
340:Actinobacteria
282:, the kingdom
255:
254:Classification
252:
248:teichoic acids
244:outer membrane
216:
215:
208:
198:
187:Teichoic acids
184:
177:
153:
150:
95:crystal violet
39:crystal violet
29:bacteria in a
15:
9:
6:
4:
3:
2:
5333:
5322:
5319:
5317:
5314:
5312:
5309:
5308:
5306:
5285:
5282:
5281:
5280:
5279:
5275:
5274:
5272:
5270:
5266:
5258:
5255:
5254:
5253:
5252:
5248:
5246:
5245:
5241:
5237:
5234:
5233:
5232:
5231:
5227:
5226:
5224:
5222:
5218:
5215:
5213:
5209:
5199:
5198:
5194:
5193:
5191:
5188:
5187:
5182:
5174:
5171:
5170:
5169:
5168:
5164:
5163:
5161:
5158:
5154:
5153:
5148:
5136:
5133:
5131:
5128:
5127:
5126:
5125:
5121:
5120:
5118:
5116:
5112:
5104:
5101:
5100:
5099:
5098:
5094:
5090:
5087:
5086:
5085:
5084:
5080:
5079:
5077:
5075:
5071:
5068:
5065:
5061:
5060:
5055:
5052:
5050:
5046:
5030:
5027:
5026:
5025:
5024:
5020:
5019:
5017:
5015:
5014:
5009:
5001:
4998:
4997:
4996:
4995:
4991:
4987:
4984:
4983:
4982:
4981:
4977:
4976:
4974:
4972:
4971:
4966:
4954:
4953:
4949:
4948:
4947:
4944:
4940:
4939:
4935:
4934:
4933:
4931:
4927:
4926:
4924:
4922:
4918:
4910:
4907:
4905:
4902:
4900:
4897:
4896:
4895:
4894:
4890:
4889:
4887:
4885:
4881:
4878:
4876:
4875:
4870:
4867:
4864:
4859:
4855:
4845:
4844:
4840:
4836:
4833:
4832:
4831:
4830:
4825:
4822:
4821:
4819:
4817:
4816:
4811:
4801:
4800:
4795:
4792:
4790:
4787:
4786:
4784:
4782:
4778:
4766:
4763:
4762:
4761:
4760:
4756:
4755:
4753:
4749:
4741:
4738:
4737:
4736:
4735:
4734:S. agalactiae
4730:
4726:
4725:
4723:
4721:
4717:
4709:
4706:
4704:
4701:
4699:
4698:Scarlet fever
4696:
4694:
4691:
4689:
4686:
4685:
4684:
4683:
4679:susceptible:
4678:
4675:
4674:
4672:
4670:
4666:
4663:
4661:
4657:
4647:
4645:
4641:
4639:
4638:
4634:
4632:
4631:
4627:
4625:
4624:
4620:
4618:
4617:
4613:
4611:
4610:
4605:
4602:
4601:
4599:
4595:
4587:
4584:
4583:
4582:
4581:
4580:S. pneumoniae
4577:
4576:
4574:
4571:
4567:
4564:
4562:
4558:
4555:
4553:
4552:
4551:Streptococcus
4547:
4544:
4541:
4536:
4532:
4529:
4527:
4523:
4517:
4513:
4510:
4508:
4504:
4500:
4497:
4496:
4491:
4486:
4484:
4479:
4477:
4472:
4471:
4468:
4456:
4452:
4449:
4448:
4443:
4442:
4439:
4432:
4431:Teskebacteria
4428:
4425:
4421:
4417:"Qinglongiia"
4416:
4415:
4413:
4409:
4408:
4406:
4402:
4385:
4382:
4381:Magnetococcia
4378:
4375:
4374:Mariprofundia
4371:
4368:
4367:Caulobacteria
4364:
4363:
4362:
4359:
4354:
4353:
4351:
4347:
4343:"Canglongiia"
4342:
4341:
4339:
4335:
4331:
4328:
4326:
4323:
4320:
4316:
4313:
4309:
4306:
4303:
4301:
4298:
4295:
4292:
4288:
4287:
4286:
4283:
4282:
4280:
4276:
4269:
4265:
4260:
4259:
4257:
4253:
4250:
4246:
4241:
4240:
4238:
4233:
4230:
4229:
4228:
4225:
4220:
4219:
4218:
4215:
4210:
4207:
4205:
4202:
4201:
4200:
4197:
4192:
4191:
4189:
4188:Moduliflexota
4185:
4180:
4179:
4177:
4173:
4168:
4167:
4165:
4161:
4156:
4155:
4153:
4149:
4145:
4142:
4141:
4139:
4135:
4130:
4127:
4126:
4124:
4120:
4115:
4114:
4112:
4108:
4104:
4101:
4099:
4096:
4094:
4091:
4090:
4089:
4086:
4085:
4083:
4081:
4077:
4068:
4067:
4065:
4061:
4057:
4054:
4053:
4051:
4046:
4045:
4044:
4041:
4036:
4035:
4033:
4029:
4024:
4023:
4022:
4021:Chrysiogenota
4019:
4014:
4013:
4011:
4007:
4003:Desulfurellia
4002:
3999:
3998:
3997:
3994:
3989:
3986:
3985:
3984:
3981:
3976:
3975:
3973:
3969:
3968:
3966:
3962:
3959:
3957:
3953:
3949:
3942:
3938:
3935:
3931:
3926:
3925:
3923:
3919:
3916:
3912:
3907:
3904:
3903:
3901:
3897:
3892:
3891:
3889:
3885:
3880:
3877:
3876:
3874:
3873:Hydrothermota
3870:
3865:
3864:
3862:
3858:
3853:
3850:
3849:
3848:
3845:
3840:
3838:Fibrobacteria
3837:
3835:
3832:
3831:
3830:
3827:
3822:
3821:
3819:
3815:
3810:
3807:
3806:
3804:
3803:Electryoneota
3800:
3797:
3793:
3790:
3786:
3783:
3779:
3774:
3773:
3771:
3767:
3764:
3760:
3755:
3754:
3752:
3748:
3743:
3742:
3741:
3740:Calditrichota
3738:
3734:
3731:
3728:
3725:
3722:
3718:
3716:
3713:
3711:
3708:
3707:
3706:
3703:
3702:
3700:
3696:
3692:
3683:
3681:
3680:Lentisphaeria
3678:
3675:
3674:
3673:
3670:
3665:
3663:
3660:
3658:
3657:Phycisphaeria
3655:
3652:
3651:
3650:
3647:
3643:
3640:
3639:
3638:
3635:
3634:
3632:
3628:
3624:
3615:
3611:
3609:"Leptospiria"
3608:
3606:"Exilispiria"
3605:
3602:
3599:
3598:
3597:
3596:Spirochaetota
3594:
3591:
3587:
3582:
3581:
3579:
3575:
3574:
3572:
3566:
3557:
3556:
3554:
3550:
3545:
3544:
3542:
3538:
3535:
3531:
3526:
3525:
3523:
3519:
3514:
3513:
3511:
3507:
3504:
3500:
3499:
3497:
3493:
3486:
3482:
3477:
3475:"Omnitrophia"
3474:
3473:
3471:
3467:
3464:
3460:
3457:
3453:
3449:Endomicrobiia
3448:
3445:
3444:
3443:
3440:
3437:
3433:
3428:
3425:
3422:
3419:
3418:
3416:
3415:Auribacterota
3412:
3407:
3406:
3404:
3400:
3399:
3397:
3393:
3390:
3388:
3384:
3383:Hydrobacteria
3380:
3373:
3369:
3364:
3363:
3361:
3357:
3352:
3351:
3349:
3345:
3340:
3339:
3337:
3333:
3329:Fusobacteriia
3328:
3327:
3326:
3323:
3322:
3320:
3318:Fusobacterida
3316:
3304:"Thermotogia"
3303:
3302:
3301:
3298:
3293:
3292:
3290:
3286:
3281:
3280:
3278:
3274:
3269:
3268:
3267:
3264:
3260:
3257:
3256:
3255:
3252:
3247:
3246:
3245:
3244:Caldisericota
3242:
3241:
3239:
3235:
3226:
3225:
3224:
3221:
3216:
3215:
3214:
3213:Atribacterota
3211:
3210:
3208:
3206:Synergistetes
3204:
3201:
3197:
3188:
3187:Wirthbacteria
3184:
3181:
3178:
3175:
3172:
3169:
3166:
3162:
3159:
3156:
3152:
3149:
3146:
3143:
3139:
3136:
3133:
3130:
3129:
3127:
3123:
3118:
3117:
3115:
3111:
3110:
3108:
3106:
3102:
3089:
3088:Selenomonadia
3085:
3084:
3082:
3077:
3076:Halanaerobiia
3073:
3072:
3070:
3065:
3062:
3060:"Peptococcia"
3059:
3056:
3053:
3050:
3047:
3044:
3041:
3040:
3038:
3034:
3029:
3026:
3025:
3023:
3018:
3015:
3012:
3011:
3009:
3004:
3001:
2997:
2994:
2993:
2991:
2986:
2983:
2980:
2976:
2975:
2973:
2968:
2964:
2963:
2962:
2959:
2958:
2956:
2950:
2946:
2942:
2933:
2930:
2927:
2924:
2923:
2921:
2917:
2912:
2908:
2905:
2903:
2902:Cyanobacteria
2900:
2899:
2897:
2896:
2894:
2890:
2881:
2878:
2877:
2875:
2872:
2868:
2863:
2860:
2857:
2854:
2852:Armatimonadia
2851:
2848:
2847:
2846:
2843:
2838:
2837:
2835:
2831:
2826:
2822:
2821:
2820:
2817:
2812:
2809:
2807:
2804:
2802:
2799:
2797:"Caldilineia"
2796:
2795:
2794:
2793:Chloroflexota
2791:
2786:
2785:
2783:
2779:
2775:
2772:
2770:
2769:Rubrobacteria
2767:
2764:
2761:
2759:
2756:
2753:
2751:
2750:Actinomycetia
2748:
2746:
2743:
2742:
2741:
2738:
2737:
2735:
2731:
2728:
2726:
2725:BV1, BV3, BV5
2722:
2721:Terrabacteria
2718:
2706:
2703:
2701:
2698:
2696:
2693:
2692:
2691:
2688:
2686:
2683:
2682:
2681:
2678:
2674:
2671:
2669:
2668:Stramenopiles
2666:
2664:
2661:
2659:
2656:
2652:
2649:
2648:
2647:
2644:
2643:
2642:
2639:
2637:
2634:
2631:(major groups
2629:
2627:
2624:
2622:
2619:
2617:
2614:
2611:
2607:
2606:
2602:
2597:
2593:
2586:
2581:
2579:
2574:
2572:
2567:
2566:
2563:
2551:
2543:
2541:
2533:
2532:
2529:
2521:
2518:
2516:
2513:
2511:
2508:
2506:
2503:
2502:
2501:
2498:
2496:
2493:
2491:
2490:Schizomycetes
2487:
2484:
2482:
2479:
2477:
2474:
2473:
2471:
2469:
2464:
2460:
2450:
2447:
2446:
2444:
2440:
2434:
2431:
2429:
2426:
2424:
2421:
2419:
2416:
2414:
2411:
2409:
2406:
2404:
2401:
2400:
2398:
2392:
2386:
2383:
2381:
2378:
2376:
2372:
2370:
2367:
2363:
2360:
2358:
2355:
2354:
2353:
2350:
2348:
2344:
2342:
2339:
2337:
2334:
2332:
2331:Teichoic acid
2329:
2327:
2323:
2319:
2316:
2314:
2311:
2309:
2306:
2305:
2304:
2303:Peptidoglycan
2300:
2297:
2295:
2294:Cell membrane
2292:
2291:
2289:
2287:
2281:
2278:
2276:
2272:
2266:
2263:
2261:
2258:
2256:
2253:
2251:
2248:
2244:
2241:
2239:
2236:
2235:
2234:
2231:
2230:
2228:
2224:
2212:
2209:
2207:
2204:
2203:
2201:
2199:
2196:
2194:
2191:
2189:
2186:
2184:
2181:
2177:
2174:
2172:
2169:
2167:
2164:
2161:
2157:
2154:
2152:
2149:
2147:
2144:
2142:
2139:
2137:
2134:
2133:
2132:
2129:
2127:
2124:
2123:
2121:
2117:
2111:
2108:
2106:
2103:
2101:
2098:
2094:
2091:
2089:
2086:
2085:
2084:
2081:
2077:
2074:
2072:
2069:
2068:
2067:
2064:
2063:
2061:
2055:
2052:
2050:
2045:
2041:
2035:
2032:
2030:
2027:
2025:
2022:
2020:
2017:
2015:
2012:
2011:
2009:
2007:
2001:
1997:
1993:
1986:
1981:
1979:
1974:
1972:
1967:
1966:
1963:
1957:
1954:
1952:
1949:
1945:
1941:
1937:
1936:
1930:
1921:
1920:
1907:
1903:
1901:
1893:
1885:
1881:
1877:
1873:
1869:
1865:
1858:
1850:
1846:
1841:
1836:
1831:
1826:
1822:
1818:
1814:
1807:
1799:
1795:
1791:
1787:
1783:
1779:
1775:
1771:
1764:
1762:
1760:
1751:
1747:
1742:
1737:
1733:
1729:
1724:
1719:
1715:
1711:
1707:
1700:
1691:
1689:9781451192681
1685:
1681:
1674:
1666:
1662:
1658:
1654:
1647:
1639:
1633:
1629:
1622:
1614:
1610:
1605:
1600:
1596:
1592:
1588:
1584:
1580:
1576:
1572:
1565:
1557:
1553:
1548:
1543:
1539:
1535:
1531:
1527:
1523:
1519:
1515:
1508:
1500:
1496:
1492:
1488:
1484:
1480:
1476:
1472:
1465:
1457:
1453:
1448:
1443:
1439:
1435:
1431:
1429:
1425:
1416:
1408:
1404:
1399:
1394:
1390:
1386:
1382:
1375:
1373:
1371:
1362:
1358:
1353:
1348:
1344:
1340:
1336:
1332:
1328:
1321:
1319:
1317:
1315:
1313:
1311:
1309:
1300:
1296:
1292:
1288:
1284:
1280:
1276:
1272:
1265:
1263:
1261:
1259:
1257:
1248:
1244:
1240:
1236:
1232:
1228:
1221:
1219:
1210:
1206:
1202:
1198:
1194:
1190:
1183:
1181:
1169:
1165:
1161:
1157:
1153:
1149:
1145:
1140:
1135:
1131:
1127:
1120:
1113:
1111:
1109:
1100:
1096:
1091:
1086:
1082:
1078:
1074:
1070:
1066:
1059:
1057:
1055:
1053:
1051:
1049:
1047:
1045:
1036:
1032:
1027:
1022:
1018:
1014:
1010:
1006:
1002:
995:
993:
983:
978:
974:
970:
969:
964:
957:
949:
945:
940:
935:
931:
927:
923:
919:
915:
911:
907:
900:
892:
886:
882:
875:
873:
864:
857:
853:
845:
843:
839:
835:
831:
827:
823:
819:
818:gram-negative
815:
814:gram-positive
805:
803:
801:
796:
795:
790:
789:
784:
783:
778:
777:
772:
767:
765:
764:bacteriophage
761:
757:
754:(transfer of
753:
749:
745:
736:
734:
730:
729:
724:
720:
716:
712:
708:
704:
700:
696:
692:
688:
687:
682:
681:
676:
675:
670:
669:
664:
660:
656:
652:
651:
646:
645:
644:Streptococcus
636:
635:
629:
623:Pathogenicity
620:
618:
614:
613:Chloroflexota
610:
606:
602:
598:
594:
593:Spirochaetota
590:
586:
582:
578:
577:Cyanobacteria
574:
570:
566:
562:
558:
554:
550:
546:
542:
538:
534:
533:Negativicutes
530:
529:
520:
516:
515:Chloroflexota
512:
509:
505:
502:
498:
494:
493:Rickettsiales
490:
486:
485:
484:
482:
481:lipid bilayer
472:
470:
466:
465:
460:
456:
452:
446:
444:
440:
434:
428:
424:
419:
409:
404:
400:
399:
394:
389:
380:
379:
371:
369:
368:
363:
362:
357:
356:
355:Mycobacterium
351:
350:
345:
341:
337:
333:
329:
325:
321:
317:
313:
309:
305:
301:
297:
293:
289:
285:
281:
273:
269:
267:
266:
261:
251:
249:
245:
241:
237:
233:
229:
225:
221:
213:
209:
206:
199:
196:
192:
188:
185:
182:
181:peptidoglycan
178:
175:
174:
173:
166:
158:
149:
147:
143:
138:
136:
132:
128:
124:
120:
115:
111:
106:
104:
100:
96:
91:
89:
88:peptidoglycan
84:
82:
78:
74:
70:
66:
59:
56:
55:gram-negative
52:
47:
40:
36:
32:
28:
27:
21:
5321:Bacteriology
5276:
5249:
5242:
5228:
5195:
5184:
5165:
5150:
5130:Gas gangrene
5122:
5114:
5095:
5081:
5073:
5057:
5021:
5011:
4992:
4978:
4968:
4950:
4945:
4936:
4928:
4891:
4872:
4841:
4827:
4815:Enterococcus
4813:
4797:
4757:
4732:
4680:
4644:S. anginosus
4643:
4635:
4630:S. sanguinis
4628:
4621:
4614:
4607:
4578:
4549:
4445:
4412:Qinglongiota
4338:Canglongiota
4227:Nitrospirota
4217:Nitrospinota
4169:"Lernaellia"
4025:Chrysiogenia
3866:"Heilongiia"
3770:Cosmopoliota
3744:Calditrichia
3733:Rhodothermia
3729:"Kryptoniia"
3705:Bacteroidota
3614:Spirochaetia
3558:"Sumerlaeia"
3470:Omnitrophota
3408:"Aerophobia"
3300:Thermotogota
3248:Caldisericia
3223:Synergistota
3217:Atribacteria
3199:Thermotogida
3030:Limnochordia
3024:Bacillota G
3010:Bacillota E
2992:Bacillota D
2974:Bacillota A
2864:"Zipacnadia"
2819:Deinococcota
2801:Chloroflexia
2690:Opisthokonta
2520:Mendosicutes
2505:Gracilicutes
2485:
2385:Mycolic acid
2375:Mycobacteria
2373:
2345:
2325:
2324:
2260:Coccobacilli
2160:in pregnancy
2126:Extremophile
2110:Aerotolerant
2044:Biochemistry
2006:microbiology
1992:Microbiology
1944:the original
1934:
1905:
1899:
1892:
1867:
1863:
1857:
1820:
1816:
1806:
1773:
1769:
1713:
1709:
1699:
1679:
1673:
1664:
1660:
1656:
1652:
1646:
1627:
1621:
1578:
1574:
1564:
1521:
1517:
1507:
1474:
1470:
1464:
1437:
1433:
1427:
1423:
1415:
1388:
1384:
1334:
1330:
1274:
1270:
1230:
1226:
1192:
1188:
1129:
1125:
1072:
1068:
1008:
1004:
972:
966:
956:
913:
909:
899:
880:
856:
833:
829:
817:
813:
811:
798:
792:
786:
780:
774:
768:
760:transduction
742:
726:
718:
714:
710:
702:
694:
684:
678:
672:
666:
648:
642:
640:
632:
609:Thermotogota
569:Bacteroidota
541:Synergistota
526:
524:
508:Deinococcota
478:
469:mycolic acid
462:
447:
442:
435:
431:
407:
396:
367:Streptomyces
365:
359:
353:
347:
296:Gracilicutes
280:Historically
278:
263:
257:
217:
171:
139:
127:counterstain
108:Conversely,
107:
92:
85:
68:
65:bacteriology
62:
24:
5284:Erysipeloid
5059:Clostridium
5029:Listeriosis
4932:susceptible
4682:S. pyogenes
4637:S. sobrinus
4572:susceptible
4455:Wikispecies
4450:(2001–2012)
4231:Nitrospiria
4221:Nitrospinia
4199:Myxococcota
4164:Lernaellota
4103:Oligoflexia
3861:Heilongiota
3710:Bacteroidia
3653:"Brocadiia"
3637:Chlamydiota
3568:Spirochaeto
3553:Sumerlaeota
3403:Aerophobota
3237:Thermocalda
3227:Synergistia
3057:"Moorellia"
2979:Clostridiia
2825:Deinococcia
2592:Prokaryotes
2408:Slime layer
2088:Facultative
2076:Facultative
838:style guide
752:conjugation
719:Clavibacter
711:Rathybacter
703:Clostridium
691:respiration
686:Clostridium
634:Actinomyces
573:Chlorobiota
565:Chlamydiota
528:Selenomonas
464:Deinococcus
439:mycoplasmas
258:Along with
228:flagellates
146:antibiotics
144:–targeting
5305:Categories
5212:Mollicutes
5186:Finegoldia
5115:nonmotile:
5049:Clostridia
4930:novobiocin
4858:Bacillales
4703:Erysipelas
4677:bacitracin
4312:Holophagia
4211:Polyangiia
4204:Myxococcia
3983:Aquificota
3964:Aquificida
3715:Chlorobiia
3642:Chlamydiia
3583:"Babeliae"
2945:Firmicutes
2879:"Xenobiia"
2515:Mollicutes
2510:Firmicutes
2500:Prokaryota
2418:Glycocalyx
2243:plasticity
2206:Lipophilic
2059:preference
2034:Resistance
1823:(2): 323.
975:(1): 1–6.
863:"Bacteria"
848:References
723:infections
561:Aquificota
499:) and are
489:Mollicutes
475:Exceptions
459:antibiotic
398:Gram stain
344:Firmicutes
308:Carl Woese
300:Mollicutes
260:cell shape
236:basal body
77:Gram stain
5159:-forming)
5066:-forming)
4893:S. aureus
4751:ungrouped
4729:CAMP test
4623:S. oralis
4616:S. mutans
4499:Bacillota
4116:"Binatia"
3987:Aquificia
3695:FCB group
3627:PVC group
3105:CPR group
2961:Bacillota
2685:Amoebozoa
2663:Alveolata
2651:Cryptista
2626:Eukaryota
2468:evolution
2442:Composite
2341:Endospore
2299:Cell wall
2275:Structure
2166:Placental
2105:Nanaerobe
2083:Anaerobic
2014:Infection
1732:0095-1137
1134:CiteSeerX
930:0066-4227
733:Bacillota
715:Leifsonia
605:Bacillota
316:monophyly
292:Bacillota
288:divisions
212:periplasm
195:chelating
142:cell wall
81:cell wall
5316:Staining
5089:Botulism
5013:Listeria
4970:Bacillus
4609:S. mitis
4570:optochin
4444:Source:
4111:Binatota
3578:Babelota
3387:BV2, BV4
2967:Bacillia
2680:Amorphea
2658:Rhizaria
2646:Hacrobia
2636:Excavata
2621:Bacteria
2596:Bacteria
2540:Category
2463:Taxonomy
2396:envelope
2286:envelope
2176:Salivary
2093:Obligate
2071:Obligate
2019:Exotoxin
1996:Bacteria
1884:18295550
1849:36838287
1798:23559881
1790:24509783
1750:17122022
1613:16751552
1556:11133473
1499:20860732
1456:20495028
1407:19667386
1361:21717204
1299:41206658
1247:20637628
1209:19299134
1168:Archived
1164:30541897
1156:10890353
948:24024634
709:. Also,
701:, while
695:Bacillus
680:Bacillus
674:Listeria
361:Nocardia
332:cytosine
232:flagella
135:fuchsine
131:safranin
73:bacteria
5103:Tetanus
5074:motile:
4986:Anthrax
4526:Bacilli
4239:SAR324
3495:Clade 2
3395:Clade 1
3114:Elulota
2695:Animals
2616:Archaea
2550:Commons
2449:Biofilm
2428:Fimbria
2413:S-layer
2394:Outside
2255:Bacilli
2171:Uterine
2156:Vaginal
2066:Aerobic
2049:ecology
2004:Medical
1906:CDC.gov
1840:9961978
1741:1829005
1604:1489649
1583:Bibcode
1526:Bibcode
1479:Bibcode
1352:3133647
1291:9723910
1099:9841678
1035:2439888
939:3883102
659:bacilli
403:Discuss
328:guanine
240:S-layer
220:capsule
114:alcohol
58:bacilli
4404:others
4278:others
2953:Low GC
2733:others
2673:Plants
2610:Domain
2495:Monera
2265:Spiral
2057:Oxygen
1882:
1847:
1837:
1796:
1788:
1748:
1738:
1730:
1686:
1634:
1611:
1601:
1554:
1544:
1497:
1454:
1405:
1359:
1349:
1297:
1289:
1245:
1207:
1162:
1154:
1136:
1097:
1087:
1033:
1026:373105
1023:
946:
936:
928:
887:
717:, and
705:is an
663:spores
653:, are
543:, and
314:, the
284:Monera
179:Thick
121:and a
41:stain.
5157:spore
5064:spore
4646:group
4501:(low-
2700:Fungi
2423:Pilus
2377:only:
2357:Porin
2349:only:
2328:only:
2250:Cocci
2226:Shape
2146:Mouth
2119:Other
1931:from
1794:S2CID
1665:genes
1547:92593
1295:S2CID
1171:(PDF)
1160:S2CID
1122:(PDF)
1090:98952
824:; as
697:is a
655:cocci
553:GroEL
549:HSP60
393:split
183:layer
103:stain
51:cocci
4909:MRSA
4863:Cat+
4540:Cat-
3131:ABY1
2466:and
2284:Cell
2151:Skin
2141:Lung
2047:and
1940:NCBI
1880:PMID
1845:PMID
1786:PMID
1746:PMID
1728:ISSN
1684:ISBN
1661:and
1659:mecA
1632:ISBN
1609:PMID
1552:PMID
1495:PMID
1452:PMID
1403:PMID
1357:PMID
1287:PMID
1243:PMID
1205:PMID
1152:PMID
1095:PMID
1031:PMID
944:PMID
926:ISSN
885:ISBN
816:and
791:and
683:and
671:and
647:and
513:The
506:The
425:and
364:and
330:and
71:are
4921:Cg-
4884:Cg+
4826:+:
4824:BEA
4796:+:
4794:BEA
4731:+:
4503:G+C
3956:BV2
2949:BV3
2318:DAP
2313:NAG
2308:NAM
2136:Gut
1872:doi
1835:PMC
1825:doi
1778:doi
1736:PMC
1718:doi
1663:nuc
1599:PMC
1591:doi
1542:PMC
1534:doi
1487:doi
1442:doi
1393:doi
1347:PMC
1339:doi
1335:100
1279:doi
1235:doi
1197:doi
1144:doi
1085:PMC
1077:doi
1021:PMC
1013:doi
977:doi
934:PMC
918:doi
842:CDC
637:sp.
579:",
575:, "
519:TM7
401:. (
336:DNA
268:).
133:or
63:In
5307::
4606::
4516:G+
4514::
4505:)
4414:"
4352:"
4340:"
4258:"
4190:"
4178:"
4166:"
4154:"
4140:"
4125:"
4113:"
4066:"
4034:"
4012:"
3974:"
3924:"
3902:"
3890:"
3875:"
3863:"
3820:"
3805:"
3772:"
3753:"
3580:"
3555:"
3543:"
3524:"
3512:"
3472:"
3417:"
3405:"
3362:"
3350:"
3338:"
3291:"
3279:"
3128:"
3116:"
3039:"
2922:"
2836:"
2784:"
2594::
2488::
2301::
1994::
1938:.
1904:.
1878:.
1866:.
1843:.
1833:.
1821:11
1819:.
1815:.
1792:.
1784:.
1774:12
1772:.
1758:^
1744:.
1734:.
1726:.
1714:45
1712:.
1708:.
1607:.
1597:.
1589:.
1579:72
1577:.
1573:.
1550:.
1540:.
1532:.
1522:67
1520:.
1516:.
1493:.
1485:.
1475:13
1473:.
1450:.
1438:61
1436:.
1432:.
1401:.
1389:60
1387:.
1383:.
1369:^
1355:.
1345:.
1333:.
1329:.
1307:^
1293:.
1285:.
1275:29
1273:.
1255:^
1241:.
1231:18
1229:.
1217:^
1203:.
1193:17
1191:.
1179:^
1166:.
1158:.
1150:.
1142:.
1130:26
1128:.
1124:.
1107:^
1093:.
1083:.
1073:62
1071:.
1067:.
1043:^
1029:.
1019:.
1009:51
1007:.
1003:.
991:^
973:28
971:.
965:.
942:.
932:.
924:.
914:67
912:.
908:.
871:^
804:.
785:,
779:,
735:.
713:,
693::
611:,
607:,
603:,
595:,
591:,
587:,
583:,
571:,
567:,
563:,
559:,
539:,
535:,
471:.
405:)
358:,
352:,
203:DD
83:.
67:,
5155:(
5062:(
4865:)
4861:(
4789:D
4781:γ
4720:B
4669:A
4660:β
4561:α
4542:)
4538:(
4489:e
4482:t
4475:v
4433:"
4429:"
4426:"
4422:"
4410:"
4383:"
4379:"
4376:"
4372:"
4369:"
4365:"
4348:"
4336:"
4321:"
4317:"
4314:"
4310:"
4293:"
4289:"
4270:"
4266:"
4254:"
4251:"
4247:"
4186:"
4174:"
4162:"
4150:"
4136:"
4121:"
4109:"
4062:"
4030:"
4008:"
3970:"
3943:"
3939:"
3936:"
3932:"
3920:"
3917:"
3913:"
3898:"
3886:"
3871:"
3859:"
3816:"
3801:"
3798:"
3794:"
3791:"
3787:"
3784:"
3780:"
3768:"
3765:"
3761:"
3749:"
3723:"
3719:"
3616:"
3612:"
3592:"
3588:"
3576:"
3551:"
3539:"
3536:"
3532:"
3520:"
3508:"
3505:"
3501:"
3487:"
3483:"
3468:"
3465:"
3461:"
3458:"
3454:"
3438:"
3434:"
3413:"
3401:"
3374:"
3370:"
3358:"
3346:"
3334:"
3287:"
3275:"
3189:"
3185:"
3167:"
3163:"
3157:"
3153:"
3144:"
3140:"
3124:"
3112:"
3090:"
3086:"
3078:"
3074:"
3035:"
3002:"
2998:"
2981:"
2977:"
2969:"
2965:"
2918:"
2913:"
2909:"
2873:"
2869:"
2832:"
2827:"
2823:"
2780:"
2707:)
2584:e
2577:t
2570:v
2162:)
2158:(
1984:e
1977:t
1970:v
1898:"
1886:.
1874::
1868:8
1851:.
1827::
1800:.
1780::
1752:.
1720::
1692:.
1640:.
1615:.
1593::
1585::
1558:.
1536::
1528::
1501:.
1489::
1481::
1458:.
1444::
1422:"
1409:.
1395::
1363:.
1341::
1301:.
1281::
1249:.
1237::
1211:.
1199::
1146::
1101:.
1079::
1037:.
1015::
985:.
979::
950:.
920::
893:.
865:.
834:g
830:G
551:(
503:.
207:.
129:(
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.