90:
513:
In mice, these IELs are the most abundant at birth and with age their numbers decrease. In humans, these cells are present during gestation but are very rare in adulthood. TCRαβ IELs develop in thymus where they undergo agonist positive selection and thereby are self-reactive. Nevertheless, they have
213:
Their role in immune system is crucial because IELs provide a first line of defense at this extensive barrier with the outside world. All IEL T cells are antigen-experienced T cells, which typically display a cytotoxic functional phenotype. IELs mediate antigen-specific delayed-type hypersensitivity
546:
TCRγδ IELs develop outside of thymus and their maintenance and function in the intestinal epithelium is influenced by a cross-talk with enterocytes. Moreover, they can migrate through the epithelium with the help of interactions with epithelial cells. Most of these cells express Vγ7 in mice and Vγ4
547:
in humans. Their function resides in the protection of the intestinal barrier against pathogens early in the infection and later they quench the inflammation and protect the barrier from tissue damage. The mechanism is not clear, but TCRγδ IELs have cytotoxic properties and can produce cytokines
176:
and in intestinal epithelium. Interaction between TL and CD8αα does not serve for migration of IELs into the epithelium, but it is important for modulating immune response of IELs. It has been suggested that cross-talk between TL and CD8αα might regulate IELs survival and proliferation. More
200:
Development and cytolytic activation are independent of live micro-organisms but they become cytolytic in response to the exogenous antigenic substances other than live micro-organisms in the gut. IEL T cells acquire their activated memory phenotype post-thymically, in response to
81:
In mice both groups are retained in almost equal proportions. In humans, the majority of IELs are alpha beta T cells. 15% of IELs are gamma delta T cells and thus represent a minor component of human IELs. However, IELs significantly increase under certain conditions, such as
412:
These DP IELs are subset of induced IELs, which are CD4 IELs with some functions of CD8 IELs and under physiological condition their number in the intestine is very small. During the intestinal inflammation, levels of DP IELs significantly increase.
250:, IEL elevation throughout the small intestine is one of many specific markers. IELs have heightened activated status that can lead to inflammatory disease such as IBD, promote cancer development and progression, or become the malignant cells in
361:. Function of TCRαβ CD4 CD8αα IELs is unclear. Even though they express granzymes and have cytolytic properties, it has been suggested that they can also have regulatory properties in the context of chronic intestinal inflammation.
431:
transcription factor, because CD4CD8aa IELs have low levels of ThPOK expression while the expression of Runx3 is very high. T-bet inducing environment is also required for the Runx3 upregulation, most likely containing
504:
Also termed unconventional IELs, express either TCRαβ or TCRγδ and do not express either CD4 or CD8αβ, but express CD8αα homodimers. In contrast to induced TCR IELs lack expression of CD2, CD5, CD28, LFA-1, and Thy1.
293:
Also termed conventional IELs, express TCRαβ together with CD4 or CD8αβ and are derived from antigen-experienced T cells that home to intraepithelial space. Contrary to natural IELs, induced IELs are the progeny of
78:(αE integrin), that is distinct from the conventional T cells in the intestine. IELs are mainly T cells with mixture of subsets. They are divided into two groups – conventional and unconventional IELs.
110:. IELs can be divided into two major subsets based on their CD8 coreceptor expression. One subset of IELs typically express activation marker CD8αα and some IELs express CD8αβ marker (CD8αβ promotes
654:
Hopper AD, Hurlstone DP, Leeds JS, McAlindon ME, Dube AK, Stephenson TJ, Sanders DS (November 2006). "The occurrence of terminal ileal histological abnormalities in patients with coeliac disease".
492:
DP IELs probably play role in intestinal homeostasis because of their immunosuppressive function. But for their cytotoxic responses they may play an important role in the pathological process of
369:
These IELs emerge from peripherally activated conventional CD8 T-cells and home to the intestinal epithelium, where they function as effector or memory cells. They continuously express
1163:
Cheroutre H (2015-01-01). "Chapter 35 - Intraepithelial TCRαβ T Cells in Health and
Disease". In Mestecky J, Lefrancois L, Strober W, Russell MW, Kelsall BL (eds.).
427:
DP IELs induction is directed by the transcriptional regulation. During the development of IELs, CD4 T cells downregulate ThPOK and instead start to express
630:
449:
74:. Due to their constant exposure to of antigens at mucosal barrier, they have unique antigen-experienced activated phenotypes and they constantly express
481:
expression to prevent pathogens from invading and to maintain integrity of the intestinal epithelial barrier. Their CD4 phenotype is responsible for
58:
Intestinal IELs are long-lived resistant effector cells spread along the entire length of intestine, where they patrol the space between intestinal
610:
These innate lymphocytes express homodimer CD8αα and CD3 and develop outside of thymus. They have cytotoxic and phagocytic properties, express
285:
IELs can be divided into different subpopulations based on molecular markers expression, mainly by expression of TCR and CD8αα, and by origin.
345:
In mice, up to 50% of these IELs can express CD8αα homodimer, which they acquire in the intestinal epithelium after external stimuli such as
614:
and thereby can present antigens to conventional CD4 T-cells. iCD8α protect against bacterial infections and promotes experimental colitis.
273:(hereditary non-polyposis colon cancer <HNPCC>). IELs themselves can, when chronically activated, undergo mutation that can lead to
622:
These cells are very similar to iCD8α population and it is unclear if this is a different subset of cells or only precursors of iCD8α.
246:
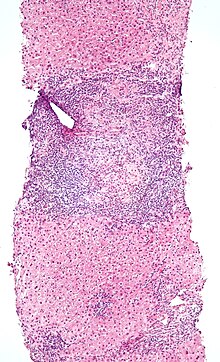
An elevated IEL population, as determined by biopsy, typically indicates ongoing inflammation within the mucosa. In diseases such as
460:(AhR). AhR is ligand-dependent transcription factor, and its activation is responsible for ThPOK downregulation. AhR is activated by
578:, where these cells seem to have pathogenic role at the beginning, whereas later they protect the epithelium against tissue damage.
489:
secretion that prevents Th1-induced inflammation in the intestine, therefore their role can be complementary to T regulatory cells.
251:
698:
145:
Expression of CD8αα is an important phenotypic marker of IELs, but not all IELs subpopulations express this molecule. CD8αα
1371:"Intestinal intraepithelial lymphocytes: Maintainers of intestinal immune tolerance and regulators of intestinal immunity"
1282:
Bellizzi AM, Frankel WL (November 2009). "Colorectal cancer due to deficiency in DNA mismatch repair function: a review".
302:-restricted CD4 naïve T cells that further undergo a post-thymic differentiation. These cells express activation markers (
1180:
1147:
1093:
961:
424:. Their migration into the intestinal epithelium depends mainly on the luminal bacteria and the dietary antigens.
548:
486:
346:
257:
Alternatively, elevated IEL populations can be a marker for developing neoplasia in the tissue such as found in
157:. CD8αα is mainly expressed by effector or mature antigen-experienced cells in the gut. This molecule can bind
52:
1239:
Chander U, Leeman-Neill RJ, Bhagat G (August 2018). "Pathogenesis of
Enteropathy-Associated T Cell Lymphoma".
1564:
Cervantes-Barragan L, Chai JN, Tianero MD, Di Luccia B, Ahern PP, Merriman J, et al. (August 2017).
493:
219:
1617:
Sujino T, London M, Hoytema van
Konijnenburg DP, Rendon T, Buch T, Silva HM, et al. (June 2016).
1734:
1676:"Intestinal Epithelial and Intraepithelial T Cell Crosstalk Mediates a Dynamic Response to Infection"
457:
1674:
Hoytema van
Konijnenburg DP, Reis BS, Pedicord VA, Farache J, Victora GD, Mucida D (November 2017).
342:. These cells migrate into the intestinal epithelium as effector or tissue-resident memory T cells.
177:
accurately, TL prevents proliferation of IELs, when there is co-occurrence of weak TCR stimulation.
531:
420:
and contrary to natural IELs, these cells increase with age, especially when they are exposed to
1131:
1122:
568:
36:
165:
towards antigens. Thus, when recognizing MHC I, CD8αα functions as a repressor of activation.
552:
470:
448:(RA). RA have the ability to induce an expression of the intetsine-homing receptors, such as
1619:"Tissue adaptation of regulatory and intraepithelial CD4⁺ T cells controls gut inflammation"
474:. Therefore, the DP IELs induction is dependent on the microbiota composition and the diet.
1729:
1630:
330:. Upon the entry into the intestinal epithelium, these cells can start express also CD8αα.
8:
595:
534:
and epithelial cells in the intestine recognizes microbes and triggers the production of
441:
215:
1634:
953:
1700:
1675:
1651:
1618:
1594:
1565:
1500:
1476:"Transcription factor T-bet regulates intraepithelial lymphocyte functional maturation"
1475:
1451:
1426:
1395:
1370:
1307:
1264:
1221:
1172:
1050:
1025:
921:
896:
867:
842:
815:
790:
757:
732:
599:
40:
1085:
1705:
1656:
1599:
1546:
1505:
1456:
1400:
1348:
1299:
1256:
1213:
1176:
1143:
1099:
1089:
1055:
1003:
957:
926:
872:
820:
762:
694:
671:
421:
222:
when transferred to semiallogeneic hosts. IELs are also able to produce a variety of
63:
1311:
1268:
1225:
89:
51:
and cause killing of infected target cells. In the GI tract, they are components of
1695:
1687:
1646:
1638:
1589:
1581:
1536:
1495:
1487:
1446:
1438:
1390:
1382:
1338:
1291:
1248:
1205:
1196:
Ondrejka S, Jagadeesh D (December 2016). "Enteropathy-Associated T-Cell
Lymphoma".
1168:
1135:
1081:
1045:
1037:
993:
949:
916:
908:
862:
854:
810:
802:
752:
744:
663:
556:
433:
394:
390:
350:
154:
103:
1541:
1524:
1491:
1442:
1343:
1326:
1295:
998:
981:
401:
370:
262:
258:
190:
162:
111:
83:
67:
47:, IELs do not need priming. Upon encountering antigens, they immediately release
32:
1041:
1691:
564:
560:
535:
482:
437:
354:
270:
1523:
Iwata M, Hirakiyama A, Eshima Y, Kagechika H, Kato C, Song SY (October 2004).
1474:
Reis BS, Hoytema van
Konijnenburg DP, Grivennikov SI, Mucida D (August 2014).
1252:
1209:
912:
858:
806:
667:
400:
Some of these cells also express CD8αα homodimer and can be pathogenic during
1723:
1473:
748:
445:
358:
339:
315:
1642:
1585:
1386:
1709:
1673:
1660:
1603:
1550:
1509:
1460:
1404:
1352:
1303:
1260:
1217:
1139:
1059:
1007:
930:
876:
824:
766:
675:
611:
299:
266:
247:
186:
1616:
1427:"Lineage re-commitment of CD4CD8αα intraepithelial lymphocytes in the gut"
1103:
944:
Lambolez F, Mayans S, Cheroutre H (2013). "Lymphocytes: Intraepithelial".
791:"Diverse developmental pathways of intestinal intraepithelial lymphocytes"
733:"Intestinal Intraepithelial Lymphocytes: Sentinels of the Mucosal Barrier"
456:). Another transcription factor responsible for DP IELs induction is the
295:
169:
158:
519:
518:
in animal experiments. These cells are influenced by normal intestinal
478:
465:
374:
126:
71:
59:
28:
24:
1563:
161:, but, opposed to the function of CD8αβ, CD8αα reduces sensitivity of
523:
146:
48:
274:
223:
130:
386:
168:
CD8αα can also recognize thymus leukemia (TL) antigen, which is a
575:
515:
214:(DTH) responses, exhibit virus-specific CTL function, to express
202:
477:
The function of CD4CD8aa IELs is due to their CD8 phenotype and
137:
expression is lower in comparison with effector memory T cells.
93:
primary biliary cirrhosis. Bile duct intraepithelial lymphocytes
461:
417:
235:
194:
173:
44:
1522:
653:
428:
407:
378:
323:
134:
118:
75:
1324:
1023:
730:
527:
453:
382:
327:
319:
307:
303:
122:
1525:"Retinoic acid imprints gut-homing specificity on T cells"
1080:. Advances in Immunology. Vol. 58. pp. 297–343.
1238:
691:
The Immune
Response in Infection and Inflammatory Disease
688:
311:
150:
107:
571:, all of which can contribute to the diverse functions.
193:. TCRαβ CD8αα (natural IELs) cells differentiate in the
982:"Doubting the TCR coreceptor function of CD8alphaalpha"
943:
1026:"TL and CD8αα: Enigmatic partners in mucosal immunity"
117:
In both humans and mice IELs express higher levels of
1167:(Fourth ed.). Academic Press. pp. 733–748.
788:
254:, a lymphoma that is a complication of celiac sprue.
1325:
Meresse B, Malamut G, Cerf-Bensussan N (June 2012).
574:
Similar functions have been found in the context of
1024:Olivares-Villagómez D, Van Kaer L (November 2010).
843:"Intraepithelial lymphocytes: to serve and protect"
153:αβ heterodimer, which is expressed on conventional
114:activation, whereas CD8αα suppresses TCR signals).
1121:
840:
1195:
1078:Intraepithelial lymphocytes and the immune system
979:
602:and in mice, they are pathogenic during colitis.
338:TCRαβCD4 IELs arise from conventional peripheral
1721:
789:McDonald BD, Jabri B, Bendelac A (August 2018).
731:Olivares-Villagómez D, Van Kaer L (April 2018).
1281:
693:. London: New Science Press. pp. 218–219.
185:Induced IELs (TCRαβ+ CD8αβ) are generated from
689:DeFranco AL, Locksley RM, Robertson M (2007).
649:
647:
645:
1570:induces gut intraepithelial CD4CD8αα T cells"
1232:
1189:
894:
1424:
841:Sheridan BS, Lefrançois L (December 2010).
642:
234:2-type cells and can also provide help for
66:(the intraepithelial space). Epithelium of
1123:"Mucosa-Associated Lymphoid Tissue (MALT)"
538:cytokine, which promotes TCRαβCD8αα IELs.
514:regulatory properties and protect against
226:which are characteristically produced by T
1699:
1650:
1593:
1540:
1499:
1450:
1394:
1368:
1342:
1327:"Celiac disease: an immunological jigsaw"
1162:
1049:
997:
980:Cheroutre H, Lambolez F (February 2008).
920:
866:
814:
756:
1425:Park Y, Moon SJ, Lee SW (January 2016).
1275:
88:
617:
408:Double positive (DP) TCRαβCD4CD8αα IELs
218:(NK)-like activity and produce a local
149:is an alternative isoform to classical
1722:
1241:Current Hematologic Malignancy Reports
1198:Current Hematologic Malignancy Reports
1130:(Second ed.). Elsevier. pp.
1119:
598:. In humans, they are elevated during
252:enteropathy-associated T-cell lymphoma
1420:
1418:
1416:
1414:
1369:Ma H, Qiu Y, Yang H (February 2021).
1364:
1362:
1115:
1113:
1071:
1069:
895:Mayassi T, Jabri B (September 2018).
416:DP IELs develop independently of the
310:) and unlike natural TCRIELs express
269:, particularly those associated with
106:, and over 75% of these also express
1019:
1017:
975:
973:
890:
888:
886:
836:
834:
784:
782:
780:
778:
776:
726:
724:
722:
720:
718:
716:
714:
712:
710:
499:
364:
288:
70:contains approximately 1 IEL per 10
1075:
954:10.1002/9780470015902.a0001197.pub3
897:"Human intraepithelial lymphocytes"
13:
1411:
1359:
1173:10.1016/b978-0-12-415847-4.00035-5
1110:
1066:
333:
14:
1746:
1014:
970:
883:
831:
773:
707:
280:
220:graft-versus-host reaction (GVHR)
847:Current Gastroenterology Reports
589:
1667:
1610:
1557:
1516:
1467:
1318:
1156:
594:These cells show properties of
468:induced by microbiota, such as
393:as opposed to the conventional
102:The majority of IELs (80%) are
1284:Advances in Anatomic Pathology
937:
682:
586:IELs that do not express TCR.
541:
508:
205:encountered in the periphery.
180:
172:molecule that is expressed in
53:gut-associated lymphoid tissue
1:
1086:10.1016/s0065-2776(08)60622-7
635:
452:and CC-chemokine receptor 9 (
385:and produce lower amounts of
1542:10.1016/j.immuni.2004.08.011
1492:10.1016/j.immuni.2014.06.017
1443:10.5483/BMBRep.2016.49.1.242
1375:Journal of Leukocyte Biology
1344:10.1016/j.immuni.2012.06.006
1296:10.1097/PAP.0b013e3181bb6bdc
999:10.1016/j.immuni.2008.01.005
581:
241:
97:
7:
1042:10.1016/j.imlet.2010.09.004
948:. American Cancer Society.
656:Digestive and Liver Disease
625:
208:
37:gastrointestinal (GI) tract
17:Intraepithelial lymphocytes
10:
1751:
1692:10.1016/j.cell.2017.08.046
1128:Encyclopedia of Immunology
795:Nature Reviews. Immunology
1253:10.1007/s11899-018-0459-5
1210:10.1007/s11899-016-0357-7
913:10.1038/s41385-018-0016-5
859:10.1007/s11894-010-0148-6
807:10.1038/s41577-018-0013-7
668:10.1016/j.dld.2006.04.003
458:Aryl hydrocarbon receptor
265:cancers, as well as some
1120:McGhee JR (1998-01-01).
749:10.1016/j.it.2017.11.003
605:
532:antigen presenting cells
140:
43:. However, unlike other
1643:10.1126/science.aaf3892
1586:10.1126/science.aah5825
1387:10.1002/JLB.3RU0220-111
1140:10.1006/rwei.1999.0448
1126:. In Delves PJ (ed.).
569:antimicrobial peptides
530:receptor expressed by
94:
1568:Lactobacillus reuteri
1076:Sim GK (1995-01-01).
471:Lactobacillus reuteri
298:-restricted CD8αβ or
92:
737:Trends in Immunology
133:cytolytic granules.
121:, activation marker
1635:2016Sci...352.1581S
1629:(6293): 1581–1586.
631:IEL of the GI tract
170:non-classical MHC I
31:layer of mammalian
1686:(4): 783–794.e13.
1165:Mucosal Immunology
1030:Immunology Letters
901:Mucosal Immunology
422:exogenous antigens
267:colorectal cancers
95:
41:reproductive tract
1580:(6353): 806–810.
700:978-0-19-920614-8
618:TCRiCD3CD8αα IELs
64:basement membrane
1742:
1735:Digestive system
1714:
1713:
1703:
1671:
1665:
1664:
1654:
1614:
1608:
1607:
1597:
1561:
1555:
1554:
1544:
1520:
1514:
1513:
1503:
1471:
1465:
1464:
1454:
1422:
1409:
1408:
1398:
1366:
1357:
1356:
1346:
1322:
1316:
1315:
1279:
1273:
1272:
1236:
1230:
1229:
1193:
1187:
1186:
1160:
1154:
1153:
1125:
1117:
1108:
1107:
1073:
1064:
1063:
1053:
1021:
1012:
1011:
1001:
977:
968:
967:
941:
935:
934:
924:
907:(5): 1281–1289.
892:
881:
880:
870:
838:
829:
828:
818:
786:
771:
770:
760:
728:
705:
704:
686:
680:
679:
651:
500:Natural TCR IELs
289:Induced TCR IELs
60:epithelial cells
1750:
1749:
1745:
1744:
1743:
1741:
1740:
1739:
1720:
1719:
1718:
1717:
1672:
1668:
1615:
1611:
1562:
1558:
1521:
1517:
1472:
1468:
1423:
1412:
1367:
1360:
1323:
1319:
1280:
1276:
1237:
1233:
1194:
1190:
1183:
1161:
1157:
1150:
1118:
1111:
1096:
1074:
1067:
1022:
1015:
978:
971:
964:
942:
938:
893:
884:
839:
832:
787:
774:
729:
708:
701:
687:
683:
662:(11): 815–819.
652:
643:
638:
628:
620:
608:
600:Crohn´s disease
592:
584:
544:
511:
502:
464:metabolites of
410:
402:coeliac disease
367:
365:TCRαβCD8αβ IELs
336:
291:
283:
244:
233:
229:
211:
191:immune response
183:
143:
100:
68:small intestine
33:mucosal linings
12:
11:
5:
1748:
1738:
1737:
1732:
1716:
1715:
1666:
1609:
1556:
1535:(4): 527–538.
1515:
1486:(2): 244–256.
1466:
1410:
1381:(2): 339–347.
1358:
1337:(6): 907–919.
1317:
1290:(6): 405–417.
1274:
1247:(4): 308–317.
1231:
1204:(6): 504–513.
1188:
1181:
1155:
1148:
1109:
1094:
1065:
1013:
992:(2): 149–159.
969:
962:
936:
882:
853:(6): 513–521.
830:
801:(8): 514–525.
772:
743:(4): 264–275.
706:
699:
681:
640:
639:
637:
634:
627:
624:
619:
616:
607:
604:
591:
588:
583:
580:
543:
540:
510:
507:
501:
498:
409:
406:
366:
363:
335:
332:
290:
287:
282:
281:Classification
279:
271:Lynch syndrome
243:
240:
231:
227:
216:natural killer
210:
207:
182:
179:
142:
139:
99:
96:
84:celiac disease
62:(IEC) and the
35:, such as the
9:
6:
4:
3:
2:
1747:
1736:
1733:
1731:
1728:
1727:
1725:
1711:
1707:
1702:
1697:
1693:
1689:
1685:
1681:
1677:
1670:
1662:
1658:
1653:
1648:
1644:
1640:
1636:
1632:
1628:
1624:
1620:
1613:
1605:
1601:
1596:
1591:
1587:
1583:
1579:
1575:
1571:
1569:
1560:
1552:
1548:
1543:
1538:
1534:
1530:
1526:
1519:
1511:
1507:
1502:
1497:
1493:
1489:
1485:
1481:
1477:
1470:
1462:
1458:
1453:
1448:
1444:
1440:
1436:
1432:
1428:
1421:
1419:
1417:
1415:
1406:
1402:
1397:
1392:
1388:
1384:
1380:
1376:
1372:
1365:
1363:
1354:
1350:
1345:
1340:
1336:
1332:
1328:
1321:
1313:
1309:
1305:
1301:
1297:
1293:
1289:
1285:
1278:
1270:
1266:
1262:
1258:
1254:
1250:
1246:
1242:
1235:
1227:
1223:
1219:
1215:
1211:
1207:
1203:
1199:
1192:
1184:
1182:9780124158474
1178:
1174:
1170:
1166:
1159:
1151:
1149:9780122267659
1145:
1141:
1137:
1133:
1129:
1124:
1116:
1114:
1105:
1101:
1097:
1095:9780120224586
1091:
1087:
1083:
1079:
1072:
1070:
1061:
1057:
1052:
1047:
1043:
1039:
1035:
1031:
1027:
1020:
1018:
1009:
1005:
1000:
995:
991:
987:
983:
976:
974:
965:
963:9780470015902
959:
955:
951:
947:
940:
932:
928:
923:
918:
914:
910:
906:
902:
898:
891:
889:
887:
878:
874:
869:
864:
860:
856:
852:
848:
844:
837:
835:
826:
822:
817:
812:
808:
804:
800:
796:
792:
785:
783:
781:
779:
777:
768:
764:
759:
754:
750:
746:
742:
738:
734:
727:
725:
723:
721:
719:
717:
715:
713:
711:
702:
696:
692:
685:
677:
673:
669:
665:
661:
657:
650:
648:
646:
641:
633:
632:
623:
615:
613:
603:
601:
597:
590:ILC-like IELs
587:
579:
577:
572:
570:
566:
562:
558:
554:
550:
539:
537:
533:
529:
525:
521:
517:
506:
497:
495:
490:
488:
484:
480:
475:
473:
472:
467:
463:
459:
455:
451:
450:α4β7-integrin
447:
446:Retionic acid
443:
439:
435:
430:
425:
423:
419:
414:
405:
403:
398:
396:
392:
388:
384:
380:
376:
372:
362:
360:
359:retinoic acid
356:
352:
348:
343:
341:
334:TCRαβCD4 IELs
331:
329:
325:
321:
317:
313:
309:
305:
301:
297:
286:
278:
276:
272:
268:
264:
260:
255:
253:
249:
239:
237:
225:
221:
217:
206:
204:
198:
196:
192:
188:
187:naive T cells
178:
175:
171:
166:
164:
160:
156:
152:
148:
138:
136:
132:
128:
124:
120:
115:
113:
109:
105:
91:
87:
85:
79:
77:
73:
69:
65:
61:
56:
54:
50:
46:
42:
38:
34:
30:
27:found in the
26:
22:
18:
1683:
1679:
1669:
1626:
1622:
1612:
1577:
1573:
1567:
1559:
1532:
1528:
1518:
1483:
1479:
1469:
1437:(1): 11–17.
1434:
1430:
1378:
1374:
1334:
1330:
1320:
1287:
1283:
1277:
1244:
1240:
1234:
1201:
1197:
1191:
1164:
1158:
1127:
1077:
1033:
1029:
989:
985:
945:
939:
904:
900:
850:
846:
798:
794:
740:
736:
690:
684:
659:
655:
629:
621:
609:
593:
585:
573:
545:
512:
503:
491:
476:
469:
426:
415:
411:
399:
368:
344:
337:
292:
284:
256:
248:celiac sprue
245:
212:
199:
184:
167:
144:
116:
101:
80:
57:
20:
16:
15:
1730:Lymphocytes
1431:BMB Reports
404:in humans.
395:CD8 T-cells
371:integrin β7
340:CD4 T-cells
238:responses.
181:Development
155:CD8 T-cells
72:enterocytes
25:lymphocytes
1724:Categories
1036:(1): 1–6.
636:References
542:TCRγδ IELs
520:microbiota
509:TCRαβ IELs
479:granzyme B
466:tryptophan
375:granzyme B
189:during an
127:granzyme B
29:epithelial
1132:1774–1780
524:vitamin D
242:Pathology
224:cytokines
147:homodimer
98:Phenotype
49:cytokines
1710:28942917
1661:27256884
1604:28775213
1551:15485630
1529:Immunity
1510:25148025
1480:Immunity
1461:26592937
1405:32678936
1353:22749351
1331:Immunity
1312:25600795
1304:19851131
1269:49430640
1261:29943210
1226:13329863
1218:27900603
1060:20850477
1008:18275828
986:Immunity
931:29674648
877:20890736
825:29717233
767:29221933
676:16787773
626:See also
596:NK cells
582:TCR IELs
275:lymphoma
263:prostate
259:cervical
230:1- and T
209:Function
203:antigens
131:perforin
55:(GALT).
1701:5670000
1652:4968079
1631:Bibcode
1623:Science
1595:5687812
1574:Science
1501:4287410
1452:4914207
1396:7891415
1104:7741030
1051:2967663
922:6178824
868:3224371
816:6063796
758:8056148
576:colitis
516:colitis
45:T cells
1708:
1698:
1659:
1649:
1602:
1592:
1549:
1508:
1498:
1459:
1449:
1403:
1393:
1351:
1310:
1302:
1267:
1259:
1224:
1216:
1179:
1146:
1102:
1092:
1058:
1048:
1006:
960:
929:
919:
875:
865:
823:
813:
765:
755:
697:
674:
612:MHC II
462:indole
418:thymus
326:, and
236:B cell
195:thymus
174:thymus
23:) are
1308:S2CID
1265:S2CID
1222:S2CID
606:iCD8α
565:IL-10
561:IL-13
557:IFN-γ
553:TNF-α
549:TGF-β
536:IL-15
487:TGF-β
483:IL-10
442:IL-15
438:IL-27
434:IFN-y
429:Runx3
391:IFN-γ
387:TNF-α
379:CD103
355:IL-27
351:IFN-γ
347:TGF-β
324:LFA-1
300:MHCII
159:MHC I
141:CD8αα
119:CD103
76:CD103
1706:PMID
1680:Cell
1657:PMID
1600:PMID
1547:PMID
1506:PMID
1457:PMID
1401:PMID
1349:PMID
1300:PMID
1257:PMID
1214:PMID
1177:ISBN
1144:ISBN
1100:PMID
1090:ISBN
1056:PMID
1004:PMID
958:ISBN
927:PMID
873:PMID
821:PMID
763:PMID
695:ISBN
672:PMID
567:and
563:and
528:NOD2
522:and
485:and
454:CCR9
444:and
389:and
383:CD69
381:and
357:and
328:Thy1
320:CD28
308:CD69
304:CD44
296:MHCI
261:and
135:CD25
129:and
123:CD69
104:CD3+
39:and
1696:PMC
1688:doi
1684:171
1647:PMC
1639:doi
1627:352
1590:PMC
1582:doi
1578:357
1537:doi
1496:PMC
1488:doi
1447:PMC
1439:doi
1391:PMC
1383:doi
1379:109
1339:doi
1292:doi
1249:doi
1206:doi
1169:doi
1136:doi
1082:doi
1046:PMC
1038:doi
1034:134
994:doi
950:doi
946:eLS
917:PMC
909:doi
863:PMC
855:doi
811:PMC
803:doi
753:PMC
745:doi
664:doi
494:IBD
316:CD5
312:CD2
163:TCR
151:CD8
112:TCR
108:CD8
21:IEL
1726::
1704:.
1694:.
1682:.
1678:.
1655:.
1645:.
1637:.
1625:.
1621:.
1598:.
1588:.
1576:.
1572:.
1545:.
1533:21
1531:.
1527:.
1504:.
1494:.
1484:41
1482:.
1478:.
1455:.
1445:.
1435:49
1433:.
1429:.
1413:^
1399:.
1389:.
1377:.
1373:.
1361:^
1347:.
1335:36
1333:.
1329:.
1306:.
1298:.
1288:16
1286:.
1263:.
1255:.
1245:13
1243:.
1220:.
1212:.
1202:11
1200:.
1175:.
1142:.
1134:.
1112:^
1098:.
1088:.
1068:^
1054:.
1044:.
1032:.
1028:.
1016:^
1002:.
990:28
988:.
984:.
972:^
956:.
925:.
915:.
905:11
903:.
899:.
885:^
871:.
861:.
851:12
849:.
845:.
833:^
819:.
809:.
799:18
797:.
793:.
775:^
761:.
751:.
741:39
739:.
735:.
709:^
670:.
660:38
658:.
644:^
559:,
555:,
551:,
526:.
496:.
440:,
436:,
397:.
377:,
373:,
353:,
349:,
322:,
318:,
314:,
306:,
277:.
197:.
125:,
1712:.
1690::
1663:.
1641::
1633::
1606:.
1584::
1566:"
1553:.
1539::
1512:.
1490::
1463:.
1441::
1407:.
1385::
1355:.
1341::
1314:.
1294::
1271:.
1251::
1228:.
1208::
1185:.
1171::
1152:.
1138::
1106:.
1084::
1062:.
1040::
1010:.
996::
966:.
952::
933:.
911::
879:.
857::
827:.
805::
769:.
747::
703:.
678:.
666::
232:h
228:h
86:.
19:(
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.