265:
242:
164:
139:
516:
271:
170:
1195:
macrophage activation syndrome, particularly in patients with juvenile rheumatoid arthritis. Onset of symptoms usually occurs after the age of 5. Moreover, some research suggests a correlation between variations in the PRF1 gene and the outcome of allogeneic bone marrow transplantation. However, these associations remain controversial, with studies disproving the connection outweighing those supporting it.
47:
1594:
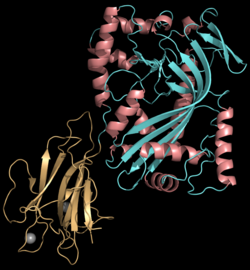
Rosado CJ, Buckle AM, Law RH, Butcher RE, Kan WT, Bird CH, Ung K, Browne KA, Baran K, Bashtannyk-Puhalovich TA, Faux NG, Wong W, Porter CJ, Pike RN, Ellisdon AM, Pearce MC, Bottomley SP, Emsley J, Smith AI, Rossjohn J, Hartland EL, Voskoboinik I, Trapani JA, Bird PI, Dunstone MA, Whisstock JC (2007).
1099:
via membrane phospholipids while phosphatidylcholine binds calcium ions to increase perforin's affinity to the membrane. Perforin oligomerises in a Ca2+ dependent manner to form pores on the target cell. The pore formed allows for the passive diffusion of a family of pro-apoptotic proteases, known as
1194:
Chronic perforinopathies are regarded as a range of immune-related conditions resulting from monoallelic mutations in genes linked to familial hemophagocytic lymphohistiocytosis (FHL). These conditions typically manifest differently from classic FHL and may include conditions like blood cancers and
1182:
Sub-acute perforinopathies encompass a diverse array of symptoms, all stemming from a partial ("sub-total") reduction in cytotoxic lymphocyte (CL) activity due to bi-allelic mutations in one of the four previously mentioned genes. In contrast to the acute form, diagnosing sub-acute perforinopathies
1139:
and is capable of lysing non-specifically a variety of target cells. This protein is one of the main cytolytic proteins of cytolytic granules, and it is known to be a key effector molecule for T-cell- and natural killer-cell-mediated cytolysis. Perforin is thought to act by creating holes in the
1127:
could be induced by adding perforin to washed cells that had already endocytosed granzyme B, even in the absence of perforin during the endocytosis process. In light of these findings, Froelich and colleagues suggested that perforin's action might not occur at the plasma membrane as previously
1063:
Perforin was initially discovered in 1983 and subsequently cloned from an expression library in 1988 using anti-complement C9 antibody cross-reactivity. A sequence comparison showed a notable resemblance between the two proteins in a specific central region, termed the 'membrane attack
1171:(FHL) and meet most or all of the criteria outlined in HLH-2004. Diagnosis is confirmed by reduced NK cell cytotoxicity, as healthy NK cells normally exhibit constitutive, pathogen-independent cytotoxicity, and by identifying mutations in genes such as PRF1, UNC13D, STX11, and STXBP2.
1183:
can be challenging due to their typically milder and more sporadic clinical manifestations, an intermittent disease course, variability in onset age, and their frequent positive response to non-specific immune-suppressive or immune-ablative treatments.
1054:
Perforin (PRF), encoded by the PRF1 gene, is a pore-forming toxic protein housed in the secretory granules of cytotoxic T lymphocytes (CTLs) and natural killer (NK) cells. Together, these cells are known as cytotoxic lymphocytes (CLs).
1809:
278:
177:
1907:
Young JD, Cohn ZA, Podack ER (1986). "The ninth component of complement and the pore-forming protein (perforin 1) from cytotoxic T cells: structural, immunological, and functional similarities".
1652:
Froelich, Christopher J.; Orth, Kim; Turbov, Jane; Seth, Prem; Gottlieb, Roberta; Babior, Bernard; Shah, Girish M.; Bleackley, R. Christopher; Dixit, Vishva M.; Hanna, William (November 1996).
1868:
Young JD, Hengartner H, Podack ER, Cohn ZA (1986). "Purification and characterization of a cytolytic pore-forming protein from granules of cloned lymphocytes with natural killer activity".
1156:
Around the turn of the 21st century, it was recognized that a total loss of perforin activity leads to a severe, fatal autosomal recessive immunoregulatory disorder in infants, known as
1807:(1996). "Target cell apoptosis induced by cytotoxic T cells and natural killer cells involves synergy between the pore-forming protein, perforin, and the serine protease, granzyme B".
1505:(1996). "Target cell apoptosis induced by cytotoxic T cells and natural killer cells involves synergy between the pore-forming protein, perforin, and the serine protease, granzyme B".
1838:
Peitsch MC, Amiguet P, Guy R, et al. (1990). "Localization and molecular modelling of the membrane-inserted domain of the ninth component of human complement and perforin".
1128:
thought, but rather at the endosomal membrane. They proposed that perforin likely facilitates the release of granzymes from endosomes by forming pores in the endosomal membrane.
2513:
1764:
Andrin C, Pinkoski MJ, Burns K, et al. (July 1998). "Interaction between a Ca2+-binding protein calreticulin and perforin, a component of the cytotoxic T-cell granules".
2147:
Andrin C, Pinkoski MJ, Burns K, et al. (1998). "Interaction between a Ca2+-binding protein calreticulin and perforin, a component of the cytotoxic T-cell granules".
1140:
plasma membrane which triggers an influx of calcium and initiates membrane repair mechanisms. These repair mechanisms bring perforin and granzymes into early endosomes.
1072:
Purifying perforin is challenging due to its tendency to lose activity and stability in solution, and only recently has a recombinant form been successfully produced.
2074:
Goebel WS, Schloemer RH, Brahmi Z (1996). "Target cell-induced perforin mRNA turnover in NK3.3 cells is mediated by multiple elements within the mRNA coding region".
1492:
Osińska, Iwona et al. “Perforin: an important player in immune response.” Central-European journal of immunology vol. 39,1 (2014): 109-15. doi:10.5114/ceji.2014.42135
1160:(FHL), typically appearing before 12 months of age. This condition can only be treated effectively with a bone marrow transplant from a non-related donor.
2410:
Ambach A, Bonnekoh B, Gollnick H (2001). "Perforin granule release from cytotoxic lymphocytes ex vivo is inhibited by ciclosporin but not by methotrexate".
1543:; Masson D; Stanley KK (1986). "Structural/functional similarity between proteins involved in complement- and cytotoxic T-lymphocyte-mediated cytolysis".
784:
1167:
and CTLs to present functional perforin, thereby failing to kill target cells. Infants affected by this condition typically show symptoms of “classic”
765:
100:
2453:
2500:
1310:
1292:
2285:
Badovinac VP, Tvinnereim AR, Harty JT (2000). "Regulation of antigen-specific CD8+ T cell homeostasis by perforin and interferon-gamma".
2221:
Stepp SE, Dufourcq-Lagelouse R, Le Deist F, et al. (1999). "Perforin gene defects in familial hemophagocytic lymphohistiocytosis".
992:
999:
1701:
2654:
1715:
Thiery J, Keefe D, Boulant S, Boucrot E, Walch M, Martinvalet D, Goping IS, Bleackley RC, Kirchhausen T, Lieberman J (2011).
1275:
2252:"Analysis of human V alpha 24+ CD4+ NKT cells activated by alpha-glycosylceramide-pulsed monocyte-derived dendritic cells"
1717:"Perforin pores in the endosomal membrane trigger the release of endocytosed granzyme B into the cytosol of target cells"
1254:
1230:
1168:
1157:
264:
2639:
2493:
241:
2105:"Virus-Specific CD4+ T Cells Eliminate Borna Disease Virus from the Brain via Induction of Cytotoxic CD8+ T Cells"
1940:"Structure of the human perforin gene. A simple gene organization with interesting potential regulatory sequences"
1095:, which works as a chaperone protein to prevent perforin from degrading. Perforin then binds to the target cell's
1279:
1258:
2659:
2522:
829:
1804:
1502:
163:
138:
810:
2486:
80:
1204:
1982:
Shinkai Y, Takio K, Okumura K (1988). "Homology of perforin to the ninth component of complement (C9)".
277:
176:
1447:
1339:"Perforinopathy: A Spectrum of Human Immune Disease Caused by Defective Perforin Delivery or Function"
270:
169:
67:
2472:
950:
903:
88:
17:
2029:
Lichtenheld MG, Olsen KJ, Lu P, et al. (1988). "Structure and function of human perforin".
975:
954:
924:
899:
2149:
1132:
1115:
The initial concept of a plasma membrane pore model was challenged when research revealed that
2447:
152:
2368:"Six novel mutations in the PRF1 gene in children with haemophagocytic lymphohistiocytosis"
2294:
2038:
1993:
1608:
1552:
1163:
The pathogenesis involves a sequence of downstream events resulting from the inability of
108:
8:
1164:
1084:
2298:
2042:
1997:
1612:
1556:
2509:
2435:
2393:
2367:
2349:
2328:
2323:
2209:
2062:
2017:
1970:
1895:
1822:
1741:
1716:
1634:
1576:
1518:
1373:
1338:
112:
2130:
2104:
699:
694:
689:
684:
679:
674:
669:
664:
659:
654:
649:
633:
628:
623:
618:
613:
608:
603:
598:
593:
577:
572:
567:
562:
557:
2664:
2427:
2398:
2354:
2310:
2273:
2238:
2201:
2166:
2135:
2091:
2087:
2054:
2009:
1962:
1926:
1909:
1887:
1883:
1856:
1852:
1826:
1781:
1746:
1683:
1675:
1626:
1568:
1522:
1475:
1467:
1425:
1417:
1378:
1360:
60:
2468:
2439:
2324:"Spectrum of Perforin Gene Mutations in Familial Hemophagocytic Lymphohistiocytosis"
2213:
2121:
1899:
1638:
2591:
2419:
2388:
2380:
2344:
2336:
2302:
2263:
2230:
2191:
2158:
2125:
2117:
2083:
2066:
2046:
2021:
2001:
1984:
1974:
1952:
1918:
1879:
1848:
1818:
1773:
1736:
1728:
1665:
1616:
1580:
1560:
1514:
1459:
1409:
1368:
1350:
1080:
357:
288:
232:
187:
2306:
2234:
92:
1957:
1939:
1096:
332:
116:
2268:
2251:
2196:
2179:
1540:
1315:
National Center for
Biotechnology Information, U.S. National Library of Medicine
1297:
National Center for
Biotechnology Information, U.S. National Library of Medicine
2372:
1870:
433:
2478:
2180:"Role of a STAT binding site in the regulation of the human perforin promoter"
1463:
2648:
1840:
1679:
1670:
1653:
1471:
1421:
1397:
1364:
1355:
1104:, into the target cell. The lytic membrane-inserting part of perforin is the
1088:
544:
2634:
1922:
1621:
1596:
1448:"Delivering the kiss of death: progress on understanding how perforin works"
493:
371:
2530:
2431:
2402:
2358:
2314:
2277:
2242:
2205:
1750:
1630:
1479:
1429:
1382:
1208:
1092:
515:
350:
129:
2170:
2139:
2095:
2058:
2013:
1966:
1930:
1891:
1860:
1830:
1785:
1687:
1572:
1526:
1039:
1034:
2621:
2596:
2384:
1944:
1413:
1120:
874:
855:
2322:
Göransdotter
Ericson K, Fadeel B, Nilsson-Ardnor S, et al. (2001).
2321:
1116:
249:
146:
96:
2423:
2162:
1777:
2540:
2109:
2050:
2005:
1564:
1124:
1109:
1091:, perforin molecules translocate to the target cell with the help of
1076:
729:
316:
303:
215:
202:
104:
31:
1732:
2611:
2606:
2601:
2586:
2570:
2545:
2535:
2340:
1225:
1220:
1123:
independently of perforin. Moreover, experiments demonstrated that
1101:
1023:
1396:
Brennan, A J; Chia, J; Trapani, J A; Voskoboinik, I (April 2010).
841:
796:
714:
710:
1597:"A common fold mediates vertebrate defense and bacterial attack"
84:
2220:
1136:
1108:
domain. This region shares homology with cholesterol-dependent
1007:
751:
524:
1148:
2616:
1395:
1105:
1174:
46:
2553:
2549:
1654:"New Paradigm for Lymphocyte Granule-mediated Cytotoxicity"
1539:
1186:
1867:
1714:
675:
positive regulation of killing of cells of other organism
2366:
Clementi R, zur Stadt U, Savoldi G, et al. (2002).
2365:
2284:
340:
1454:. Lymphocyte activation/Lymphocyte effector functions.
1131:
Perforin has structural and functional similarities to
2409:
1651:
1593:
2250:
Takahashi T, Nieda M, Koezuka Y, et al. (2000).
2249:
2102:
2073:
1702:"Entrez Gene: PRF1 perforin 1 (pore forming protein)"
505:
2146:
1981:
1763:
2178:Yu CR, Ortaldo JR, Curiel RE, et al. (1999).
2028:
1398:"Perforin deficiency and susceptibility to cancer"
1336:
1271:
1269:
1267:
1250:
1248:
1246:
1135:(C9). Like C9, this protein creates transmembrane
1937:
1837:
1446:Pipkin, Matthew E; Lieberman, Judy (2007-06-01).
287:
186:
2646:
2508:
2177:
1906:
1445:
1264:
1243:
1810:Australian and New Zealand Journal of Medicine
1587:
1507:Australian and New Zealand Journal of Medicine
1337:Voskoboinik, Ilia; Trapani, Joseph A. (2013).
1276:GRCm38: Ensembl release 89: ENSMUSG00000037202
2494:
2452:: CS1 maint: DOI inactive as of June 2024 (
2103:Nöske K, Bilzer T, Planz O, Stitz L (1998).
1708:
1757:
1255:GRCh38: Ensembl release 89: ENSG00000180644
1169:familial hemophagocytic lymphohistiocytosis
1158:familial hemophagocytic lymphohistiocytosis
2501:
2487:
1803:
1501:
1495:
2471:at the U.S. National Library of Medicine
2392:
2348:
2267:
2195:
2129:
1956:
1740:
1669:
1620:
1372:
1354:
1067:
1143:
71:, FLH2, HPLH2, P1, PFN1, PFP, perforin 1
27:Mammalian protein found in Homo sapiens
14:
2647:
1694:
2482:
1152:presentations of perforinopathy — FHL
292:
253:
248:
191:
150:
145:
1441:
1439:
1332:
1330:
1328:
1326:
1324:
1064:complex/perforin' (MACPF) domain.
24:
2412:Skin Pharmacol. Appl. Skin Physiol
1938:Lichtenheld MG, Podack ER (1990).
1823:10.1111/j.1445-5994.1995.tb02883.x
1796:
1519:10.1111/j.1445-5994.1995.tb02883.x
1231:Complement membrane attack complex
25:
2676:
2462:
1436:
1321:
1079:protein found in the granules of
1402:Cell Death & Differentiation
514:
456:lumbar subsegment of spinal cord
276:
269:
263:
240:
175:
168:
162:
137:
45:
2523:Antimicrobial cationic peptides
2122:10.1128/JVI.72.5.4387-4395.1998
1658:Journal of Biological Chemistry
1645:
1533:
1198:
1190:presentations of perforinopathy
1178:presentations of perforinopathy
1085:natural killer cells (NK cells)
655:immunological synapse formation
1486:
1389:
1303:
1285:
1081:cytotoxic T lymphocytes (CTLs)
680:defense response to tumor cell
594:integral component of membrane
525:More reference expression data
494:More reference expression data
13:
1:
2307:10.1126/science.290.5495.1354
2235:10.1126/science.286.5446.1957
1452:Current Opinion in Immunology
1236:
1112:from Gram-positive bacteria.
685:immune response to tumor cell
261:
160:
2655:Genes on human chromosome 10
2088:10.1016/0161-5890(95)00155-7
1958:10.4049/jimmunol.143.12.4267
1884:10.1016/0092-8674(86)90007-3
1853:10.1016/0161-5890(90)90001-G
1058:
700:T cell mediated cytotoxicity
7:
2269:10.4049/jimmunol.164.9.4458
2197:10.4049/jimmunol.162.5.2785
1214:
1203:Perforin has been shown to
1075:Perforin is a pore forming
690:protein homooligomerization
472:subcutaneous adipose tissue
414:superficial temporal artery
10:
2681:
573:wide pore channel activity
29:
2579:
2563:
2521:
1464:10.1016/j.coi.2007.04.011
1311:"Mouse PubMed Reference:"
1293:"Human PubMed Reference:"
1038:
1033:
1029:
1022:
1006:
987:
972:
968:
947:
943:
936:
921:
917:
896:
892:
885:
872:
868:
853:
849:
840:
827:
823:
808:
804:
795:
782:
778:
763:
759:
750:
735:
728:
724:
708:
695:defense response to virus
650:cellular defense response
578:identical protein binding
543:
539:
522:
513:
504:
491:
440:
431:
378:
369:
339:
331:
327:
310:
297:
260:
239:
230:
226:
209:
196:
159:
136:
127:
123:
78:
75:
65:
58:
53:
44:
39:
2642:Genetics Home Reference
2473:Medical Subject Headings
1671:10.1074/jbc.271.46.29073
1356:10.3389/fimmu.2013.00441
1000:Chr 10: 61.13 – 61.14 Mb
30:Not to be confused with
2426:(inactive 2024-06-11).
1923:10.1126/science.2425429
1622:10.1126/science.1144706
1343:Frontiers in Immunology
670:transmembrane transport
418:upper lobe of left lung
1133:complement component 9
1068:Structure and Function
993:Chr 10: 70.6 – 70.6 Mb
294:10 B4|10 32.18 cM
2660:Programmed cell death
1144:Clinical Significance
255:Chromosome 10 (mouse)
153:Chromosome 10 (human)
2385:10.1136/jmg.38.9.643
1414:10.1038/cdd.2009.212
599:extracellular region
2510:Pore-forming toxins
2299:2000Sci...290.1354B
2043:1988Natur.335..448L
1998:1988Natur.334..525S
1664:(46): 29073–29079.
1613:2007Sci...317.1548R
1557:1986Natur.322..831T
629:cytoplasmic vesicle
568:calcium ion binding
2329:Am. J. Hum. Genet.
830:ENSMUSG00000037202
643:Biological process
587:Cellular component
551:Molecular function
402:buccal mucosa cell
2630:
2629:
2424:10.1159/000056355
2163:10.1021/bi980595z
1778:10.1021/bi980595z
1607:(5844): 1548–51.
1049:
1048:
1045:
1044:
1018:
1017:
983:
982:
962:
961:
932:
931:
911:
910:
881:
880:
862:
861:
836:
835:
817:
816:
791:
790:
772:
771:
720:
719:
665:apoptotic process
634:cytolytic granule
563:metal ion binding
535:
534:
531:
530:
500:
499:
487:
486:
425:
424:
398:bone marrow cells
323:
322:
222:
221:
16:(Redirected from
2672:
2592:Diphtheria toxin
2503:
2496:
2489:
2480:
2479:
2457:
2451:
2443:
2406:
2396:
2362:
2352:
2318:
2293:(5495): 1354–8.
2281:
2271:
2246:
2229:(5446): 1957–9.
2217:
2199:
2174:
2157:(29): 10386–94.
2143:
2133:
2099:
2070:
2051:10.1038/335448a0
2037:(6189): 448–51.
2025:
2006:10.1038/334525a0
1978:
1960:
1934:
1917:(4760): 184–90.
1903:
1864:
1834:
1790:
1789:
1772:(29): 10386–94.
1761:
1755:
1754:
1744:
1712:
1706:
1705:
1698:
1692:
1691:
1673:
1649:
1643:
1642:
1624:
1591:
1585:
1584:
1565:10.1038/322831a0
1537:
1531:
1530:
1499:
1493:
1490:
1484:
1483:
1443:
1434:
1433:
1393:
1387:
1386:
1376:
1358:
1334:
1319:
1318:
1307:
1301:
1300:
1289:
1283:
1273:
1262:
1252:
1031:
1030:
1002:
995:
978:
966:
965:
957:
941:
940:
937:RefSeq (protein)
927:
915:
914:
906:
890:
889:
866:
865:
847:
846:
821:
820:
802:
801:
776:
775:
757:
756:
726:
725:
541:
540:
527:
518:
511:
510:
496:
436:
434:Top expressed in
429:
428:
374:
372:Top expressed in
367:
366:
346:
345:
329:
328:
319:
306:
295:
280:
273:
267:
256:
244:
228:
227:
218:
205:
194:
179:
172:
166:
155:
141:
125:
124:
119:
117:PRF1 - orthologs
70:
63:
49:
37:
36:
21:
2680:
2679:
2675:
2674:
2673:
2671:
2670:
2669:
2645:
2644:
2631:
2626:
2580:Other, nonhuman
2575:
2559:
2517:
2507:
2465:
2460:
2445:
2444:
1992:(6182): 525–7.
1951:(12): 4267–74.
1799:
1797:Further reading
1794:
1793:
1762:
1758:
1733:10.1038/ni.2050
1713:
1709:
1700:
1699:
1695:
1650:
1646:
1592:
1588:
1551:(6082): 831–4.
1538:
1534:
1500:
1496:
1491:
1487:
1444:
1437:
1394:
1390:
1335:
1322:
1309:
1308:
1304:
1291:
1290:
1286:
1274:
1265:
1253:
1244:
1239:
1217:
1201:
1192:
1180:
1154:
1146:
1097:plasma membrane
1070:
1061:
1040:View/Edit Mouse
1035:View/Edit Human
998:
991:
988:Location (UCSC)
974:
953:
949:
923:
902:
898:
811:ENSG00000180644
704:
638:
609:plasma membrane
582:
558:protein binding
523:
492:
483:
478:
474:
470:
466:
462:
458:
454:
450:
446:
432:
421:
416:
412:
408:
404:
400:
396:
392:
388:
384:
370:
314:
301:
293:
283:
282:
281:
274:
254:
231:Gene location (
213:
200:
192:
182:
181:
180:
173:
151:
128:Gene location (
79:
66:
59:
35:
28:
23:
22:
15:
12:
11:
5:
2678:
2668:
2667:
2662:
2657:
2628:
2627:
2625:
2624:
2619:
2614:
2609:
2604:
2599:
2594:
2589:
2583:
2581:
2577:
2576:
2574:
2573:
2567:
2565:
2561:
2560:
2558:
2557:
2543:
2538:
2533:
2527:
2525:
2519:
2518:
2506:
2505:
2498:
2491:
2483:
2477:
2476:
2464:
2463:External links
2461:
2459:
2458:
2407:
2373:J. Med. Genet.
2363:
2341:10.1086/318796
2319:
2282:
2262:(9): 4458–64.
2247:
2218:
2190:(5): 2785–90.
2175:
2144:
2116:(5): 4387–95.
2100:
2082:(4–5): 341–9.
2071:
2026:
1979:
1935:
1904:
1865:
1847:(7): 589–602.
1835:
1800:
1798:
1795:
1792:
1791:
1756:
1707:
1693:
1644:
1586:
1532:
1494:
1485:
1458:(3): 301–308.
1435:
1408:(4): 607–615.
1388:
1320:
1302:
1284:
1263:
1241:
1240:
1238:
1235:
1234:
1233:
1228:
1223:
1216:
1213:
1200:
1197:
1191:
1185:
1179:
1173:
1153:
1147:
1145:
1142:
1119:could undergo
1069:
1066:
1060:
1057:
1047:
1046:
1043:
1042:
1037:
1027:
1026:
1020:
1019:
1016:
1015:
1013:
1011:
1004:
1003:
996:
989:
985:
984:
981:
980:
970:
969:
963:
960:
959:
945:
944:
938:
934:
933:
930:
929:
919:
918:
912:
909:
908:
894:
893:
887:
883:
882:
879:
878:
870:
869:
863:
860:
859:
851:
850:
844:
838:
837:
834:
833:
825:
824:
818:
815:
814:
806:
805:
799:
793:
792:
789:
788:
780:
779:
773:
770:
769:
761:
760:
754:
748:
747:
742:
737:
733:
732:
722:
721:
718:
717:
706:
705:
703:
702:
697:
692:
687:
682:
677:
672:
667:
662:
657:
652:
646:
644:
640:
639:
637:
636:
631:
626:
621:
616:
614:endosome lumen
611:
606:
601:
596:
590:
588:
584:
583:
581:
580:
575:
570:
565:
560:
554:
552:
548:
547:
537:
536:
533:
532:
529:
528:
520:
519:
508:
502:
501:
498:
497:
489:
488:
485:
484:
482:
481:
477:
473:
469:
465:
461:
457:
453:
449:
445:
441:
438:
437:
426:
423:
422:
420:
419:
415:
411:
407:
403:
399:
395:
391:
387:
383:
379:
376:
375:
363:
362:
354:
343:
337:
336:
333:RNA expression
325:
324:
321:
320:
312:
308:
307:
299:
296:
291:
285:
284:
275:
268:
262:
258:
257:
252:
246:
245:
237:
236:
224:
223:
220:
219:
211:
207:
206:
198:
195:
190:
184:
183:
174:
167:
161:
157:
156:
149:
143:
142:
134:
133:
121:
120:
77:
73:
72:
64:
56:
55:
51:
50:
42:
41:
26:
9:
6:
4:
3:
2:
2677:
2666:
2663:
2661:
2658:
2656:
2653:
2652:
2650:
2643:
2641:
2637:
2636:
2623:
2620:
2618:
2615:
2613:
2610:
2608:
2605:
2603:
2600:
2598:
2595:
2593:
2590:
2588:
2585:
2584:
2582:
2578:
2572:
2569:
2568:
2566:
2562:
2555:
2551:
2547:
2544:
2542:
2539:
2537:
2534:
2532:
2529:
2528:
2526:
2524:
2520:
2515:
2511:
2504:
2499:
2497:
2492:
2490:
2485:
2484:
2481:
2474:
2470:
2467:
2466:
2455:
2449:
2441:
2437:
2433:
2429:
2425:
2421:
2418:(5): 249–60.
2417:
2413:
2408:
2404:
2400:
2395:
2390:
2386:
2382:
2378:
2375:
2374:
2369:
2364:
2360:
2356:
2351:
2346:
2342:
2338:
2334:
2331:
2330:
2325:
2320:
2316:
2312:
2308:
2304:
2300:
2296:
2292:
2288:
2283:
2279:
2275:
2270:
2265:
2261:
2257:
2253:
2248:
2244:
2240:
2236:
2232:
2228:
2224:
2219:
2215:
2211:
2207:
2203:
2198:
2193:
2189:
2185:
2181:
2176:
2172:
2168:
2164:
2160:
2156:
2152:
2151:
2145:
2141:
2137:
2132:
2127:
2123:
2119:
2115:
2112:
2111:
2106:
2101:
2097:
2093:
2089:
2085:
2081:
2077:
2072:
2068:
2064:
2060:
2056:
2052:
2048:
2044:
2040:
2036:
2032:
2027:
2023:
2019:
2015:
2011:
2007:
2003:
1999:
1995:
1991:
1987:
1986:
1980:
1976:
1972:
1968:
1964:
1959:
1954:
1950:
1947:
1946:
1941:
1936:
1932:
1928:
1924:
1920:
1916:
1912:
1911:
1905:
1901:
1897:
1893:
1889:
1885:
1881:
1878:(6): 849–59.
1877:
1873:
1872:
1866:
1862:
1858:
1854:
1850:
1846:
1843:
1842:
1841:Mol. Immunol.
1836:
1832:
1828:
1824:
1820:
1816:
1812:
1811:
1806:
1802:
1801:
1787:
1783:
1779:
1775:
1771:
1767:
1760:
1752:
1748:
1743:
1738:
1734:
1730:
1726:
1722:
1718:
1711:
1703:
1697:
1689:
1685:
1681:
1677:
1672:
1667:
1663:
1659:
1655:
1648:
1640:
1636:
1632:
1628:
1623:
1618:
1614:
1610:
1606:
1602:
1598:
1590:
1582:
1578:
1574:
1570:
1566:
1562:
1558:
1554:
1550:
1546:
1542:
1536:
1528:
1524:
1520:
1516:
1512:
1508:
1504:
1498:
1489:
1481:
1477:
1473:
1469:
1465:
1461:
1457:
1453:
1449:
1442:
1440:
1431:
1427:
1423:
1419:
1415:
1411:
1407:
1403:
1399:
1392:
1384:
1380:
1375:
1370:
1366:
1362:
1357:
1352:
1348:
1344:
1340:
1333:
1331:
1329:
1327:
1325:
1316:
1312:
1306:
1298:
1294:
1288:
1281:
1277:
1272:
1270:
1268:
1260:
1256:
1251:
1249:
1247:
1242:
1232:
1229:
1227:
1224:
1222:
1219:
1218:
1212:
1210:
1206:
1196:
1189:
1184:
1177:
1172:
1170:
1166:
1161:
1159:
1151:
1141:
1138:
1134:
1129:
1126:
1122:
1118:
1113:
1111:
1107:
1103:
1098:
1094:
1090:
1089:degranulation
1086:
1082:
1078:
1073:
1065:
1056:
1053:
1041:
1036:
1032:
1028:
1025:
1021:
1014:
1012:
1009:
1005:
1001:
997:
994:
990:
986:
979:
977:
971:
967:
964:
958:
956:
952:
946:
942:
939:
935:
928:
926:
920:
916:
913:
907:
905:
901:
895:
891:
888:
886:RefSeq (mRNA)
884:
877:
876:
871:
867:
864:
858:
857:
852:
848:
845:
843:
839:
832:
831:
826:
822:
819:
813:
812:
807:
803:
800:
798:
794:
787:
786:
781:
777:
774:
768:
767:
762:
758:
755:
753:
749:
746:
743:
741:
738:
734:
731:
727:
723:
716:
712:
707:
701:
698:
696:
693:
691:
688:
686:
683:
681:
678:
676:
673:
671:
668:
666:
663:
661:
658:
656:
653:
651:
648:
647:
645:
642:
641:
635:
632:
630:
627:
625:
622:
620:
617:
615:
612:
610:
607:
605:
602:
600:
597:
595:
592:
591:
589:
586:
585:
579:
576:
574:
571:
569:
566:
564:
561:
559:
556:
555:
553:
550:
549:
546:
545:Gene ontology
542:
538:
526:
521:
517:
512:
509:
507:
503:
495:
490:
479:
475:
471:
467:
463:
459:
455:
451:
447:
443:
442:
439:
435:
430:
427:
417:
413:
409:
406:apex of heart
405:
401:
397:
393:
389:
385:
381:
380:
377:
373:
368:
365:
364:
361:
359:
355:
353:
352:
348:
347:
344:
342:
338:
334:
330:
326:
318:
313:
309:
305:
300:
290:
286:
279:
272:
266:
259:
251:
247:
243:
238:
234:
229:
225:
217:
212:
208:
204:
199:
189:
185:
178:
171:
165:
158:
154:
148:
144:
140:
135:
131:
126:
122:
118:
114:
110:
106:
102:
98:
94:
90:
86:
82:
74:
69:
62:
57:
52:
48:
43:
38:
33:
19:
2633:
2632:
2564:Other, human
2531:Cathelicidin
2448:cite journal
2415:
2411:
2379:(9): 643–6.
2376:
2371:
2335:(3): 590–7.
2332:
2327:
2290:
2286:
2259:
2255:
2226:
2222:
2187:
2183:
2154:
2150:Biochemistry
2148:
2113:
2108:
2079:
2076:Mol. Immunol
2075:
2034:
2030:
1989:
1983:
1948:
1943:
1914:
1908:
1875:
1869:
1844:
1839:
1817:(6): 793–9.
1814:
1808:
1769:
1766:Biochemistry
1765:
1759:
1727:(8): 770–7.
1724:
1721:Nat. Immunol
1720:
1710:
1696:
1661:
1657:
1647:
1604:
1600:
1589:
1548:
1544:
1535:
1513:(6): 793–9.
1510:
1506:
1497:
1488:
1455:
1451:
1405:
1401:
1391:
1346:
1342:
1314:
1305:
1296:
1287:
1209:calreticulin
1202:
1199:Interactions
1193:
1187:
1181:
1175:
1162:
1155:
1149:
1130:
1114:
1093:calreticulin
1074:
1071:
1062:
1051:
1050:
973:
951:NP_001076585
948:
922:
904:NM_001083116
897:
873:
854:
828:
809:
783:
764:
744:
739:
356:
349:
76:External IDs
2622:Pneumolysin
2597:Dermaseptin
1945:J. Immunol.
1121:endocytosis
382:granulocyte
315:61,140,459
302:61,133,612
214:70,602,759
201:70,597,348
54:Identifiers
2649:Categories
2635:Perforin-1
2256:J. Immunol
2184:J. Immunol
1805:Trapani JA
1503:Trapani JA
1282:, May 2017
1261:, May 2017
1237:References
1117:granzyme B
1110:cytolysins
1052:Perforin-1
480:right lung
468:lymph node
410:lymph node
360:(ortholog)
97:HomoloGene
2541:Dermcidin
2110:J. Virol.
1680:0021-9258
1541:Tschopp J
1472:0952-7915
1422:1350-9047
1365:1664-3224
1221:Granzymes
1176:Sub-acute
1125:apoptosis
1102:granzymes
1077:cytolytic
1059:Discovery
976:NP_035203
955:NP_005032
925:NM_011073
900:NM_005041
730:Orthologs
660:cytolysis
105:GeneCards
32:Porphyrin
2665:Proteins
2612:Melittin
2607:Magainin
2602:Latarcin
2587:Cecropin
2571:Perforin
2546:Histatin
2536:Defensin
2469:perforin
2440:30142804
2432:11586066
2403:11565555
2359:11179007
2315:11082062
2278:10779745
2243:10583959
2214:26096007
2206:10072525
1900:30182487
1751:21685908
1639:20372720
1631:17717151
1480:17433871
1430:20075937
1383:24376445
1278:–
1257:–
1226:Defensin
1215:See also
1205:interact
1165:NK cells
1024:Wikidata
709:Sources:
619:membrane
604:endosome
444:gastrula
18:Perforin
2394:1734943
2350:1274472
2295:Bibcode
2287:Science
2223:Science
2171:9671507
2140:9557729
2096:8676885
2067:4359028
2059:3419519
2039:Bibcode
2022:4348928
2014:3261391
1994:Bibcode
1975:8326644
1967:2480391
1931:2425429
1910:Science
1892:2420467
1861:2395434
1831:8770355
1786:9671507
1742:3140544
1688:8910561
1609:Bibcode
1601:Science
1581:4330219
1573:2427956
1553:Bibcode
1527:8770355
1374:3860100
1349:: 441.
1280:Ensembl
1259:Ensembl
1188:Chronic
1137:tubules
1087:. Upon
842:UniProt
797:Ensembl
736:Species
715:QuickGO
624:cytosol
448:decidua
394:decidua
335:pattern
193:10q22.1
61:Aliases
2475:(MeSH)
2438:
2430:
2401:
2391:
2357:
2347:
2313:
2276:
2241:
2212:
2204:
2169:
2138:
2131:109669
2128:
2094:
2065:
2057:
2031:Nature
2020:
2012:
1985:Nature
1973:
1965:
1929:
1898:
1890:
1859:
1829:
1784:
1749:
1739:
1686:
1678:
1637:
1629:
1579:
1571:
1545:Nature
1525:
1478:
1470:
1428:
1420:
1381:
1371:
1363:
1010:search
1008:PubMed
875:P10820
856:P14222
752:Entrez
506:BioGPS
476:thymus
464:spleen
460:embryo
390:spleen
85:170280
2617:Nisin
2514:TC 1C
2436:S2CID
2210:S2CID
2063:S2CID
2018:S2CID
1971:S2CID
1896:S2CID
1635:S2CID
1577:S2CID
1207:with
1150:Acute
1106:MACPF
785:18646
745:Mouse
740:Human
711:Amigo
452:blood
386:blood
358:Mouse
351:Human
298:Start
233:Mouse
197:Start
130:Human
93:97551
2554:HTN3
2550:HTN1
2454:link
2428:PMID
2399:PMID
2355:PMID
2311:PMID
2274:PMID
2239:PMID
2202:PMID
2167:PMID
2136:PMID
2092:PMID
2055:PMID
2010:PMID
1963:PMID
1927:PMID
1888:PMID
1871:Cell
1857:PMID
1827:PMID
1782:PMID
1747:PMID
1684:PMID
1676:ISSN
1627:PMID
1569:PMID
1523:PMID
1476:PMID
1468:ISSN
1426:PMID
1418:ISSN
1379:PMID
1361:ISSN
1100:the
1083:and
766:5551
341:Bgee
289:Band
250:Chr.
188:Band
147:Chr.
109:PRF1
101:3698
81:OMIM
68:PRF1
40:PRF1
2640:NLM
2638:at
2420:doi
2389:PMC
2381:doi
2345:PMC
2337:doi
2303:doi
2291:290
2264:doi
2260:164
2231:doi
2227:286
2192:doi
2188:162
2159:doi
2126:PMC
2118:doi
2084:doi
2047:doi
2035:335
2002:doi
1990:334
1953:doi
1949:143
1919:doi
1915:233
1880:doi
1849:doi
1819:doi
1774:doi
1737:PMC
1729:doi
1666:doi
1662:271
1617:doi
1605:317
1561:doi
1549:322
1515:doi
1460:doi
1410:doi
1369:PMC
1351:doi
311:End
210:End
113:OMA
89:MGI
2651::
2552:,
2450:}}
2446:{{
2434:.
2416:14
2414:.
2397:.
2387:.
2377:38
2370:.
2353:.
2343:.
2333:68
2326:.
2309:.
2301:.
2289:.
2272:.
2258:.
2254:.
2237:.
2225:.
2208:.
2200:.
2186:.
2182:.
2165:.
2155:37
2153:.
2134:.
2124:.
2114:72
2107:.
2090:.
2080:33
2078:.
2061:.
2053:.
2045:.
2033:.
2016:.
2008:.
2000:.
1988:.
1969:.
1961:.
1942:.
1925:.
1913:.
1894:.
1886:.
1876:44
1874:.
1855:.
1845:27
1825:.
1815:25
1813:.
1780:.
1770:37
1768:.
1745:.
1735:.
1725:12
1723:.
1719:.
1682:.
1674:.
1660:.
1656:.
1633:.
1625:.
1615:.
1603:.
1599:.
1575:.
1567:.
1559:.
1547:.
1521:.
1511:25
1509:.
1474:.
1466:.
1456:19
1450:.
1438:^
1424:.
1416:.
1406:17
1404:.
1400:.
1377:.
1367:.
1359:.
1345:.
1341:.
1323:^
1313:.
1295:.
1266:^
1245:^
1211:.
713:/
317:bp
304:bp
216:bp
203:bp
111:;
107::
103:;
99::
95:;
91::
87:;
83::
2556:)
2548:(
2516:)
2512:(
2502:e
2495:t
2488:v
2456:)
2442:.
2422::
2405:.
2383::
2361:.
2339::
2317:.
2305::
2297::
2280:.
2266::
2245:.
2233::
2216:.
2194::
2173:.
2161::
2142:.
2120::
2098:.
2086::
2069:.
2049::
2041::
2024:.
2004::
1996::
1977:.
1955::
1933:.
1921::
1902:.
1882::
1863:.
1851::
1833:.
1821::
1788:.
1776::
1753:.
1731::
1704:.
1690:.
1668::
1641:.
1619::
1611::
1583:.
1563::
1555::
1529:.
1517::
1482:.
1462::
1432:.
1412::
1385:.
1353::
1347:4
1317:.
1299:.
235:)
132:)
115::
34:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.