377:
1500 bps of a transcript. The technique does not depend on restriction enzymes anymore and thereby circumvents bias that is related to the absence or location of the restriction site within the cDNA. Instead, the cDNA is randomly fragmented and the 3'ends are sequenced from the 5' end of the cDNA molecule that carries the poly-A tail. The sequencing length of the tag can be freely chosen. Because of this, the tags can be assembled into contigs and the annotation of the tags can be drastically improved. Therefore, MACE is also use for the analyses of non-model organisms. In addition, the longer contigs can be screened for polymorphisms. As UTRs show a large number of polymorphisms between individuals, the MACE approach can be applied for allele determination, allele specific gene expression profiling and the search for molecular markers for breeding. In addition, the approach allows determining alternative polyadenylation of the transcripts. Because MACE does only require 3’ ends of transcripts, even partly degraded RNA can be analyzed with less degradation dependent bias. The MACE approach uses unique molecular identifiers to allow for identification of PCR bias.
304:. Following this, the linkers, containing internal restriction sites, are digested with the appropriate restriction enzyme and the sticky ends are ligated together into concatamers. Following concatenation, the fragments are ligated into plasmids and are used to transform bacteria to generate many copies of the plasmid containing the inserts. Those may then be sequenced to identify the miRNA present, as well as analysing expression levels of a given miRNA by counting the number of times it is present, similar to SAGE.
20:
141:
attachment site: 1) Sticky ends with the AE cut site to allow for attachment to cleaved cDNA; 2) A recognition site for a restriction endonuclease known as the tagging enzyme (TE), which cuts about 15 nucleotides downstream of its recognition site (within the original cDNA/mRNA sequence); 3) A short primer sequence unique to either adaptor A or B, which will later be used for further amplification via PCR.
312:
LongSAGE was a more robust version of the original SAGE developed in 2002 which had a higher throughput, using 20 μg of mRNA to generate a cDNA library of thousands of tags. Robust LongSage (RL-SAGE) Further improved on the LongSAGE protocol with the ability to generate a library with an insert size
93:
population in a sample of interest in the form of small tags that correspond to fragments of those transcripts. Several variants have been developed since, most notably a more robust version, LongSAGE, RL-SAGE and the most recent SuperSAGE. Many of these have improved the technique with the capture
376:
In the mid 2010s several techniques combined with Next
Generation Sequencing were developed that employ the "tag" principle for "digital gene expression profiling" but without the use of the tagging enzyme. The "MACE" approach, (=Massive Analysis of cDNA Ends) generates tags somewhere in the last
228:
In 1979 teams at
Harvard and Caltech extended the basic idea of making DNA copies of mRNAs in vitro to amplifying a library of such in bacterial plasmids. In 1982–1983, the idea of selecting random or semi-random clones from such a cDNA library for sequencing was explored by Greg Sutcliffe and
140:
The cleaved cDNA downstream from the cleavage site is then discarded, and the remaining immobile cDNA fragments upstream from cleavage sites are divided in half and exposed to one of two adaptor oligonucleotides (A or B) containing several components in the following order upstream from the
345:, expanding the tag-size by at least 6 bp as compared to the predecessor techniques SAGE and LongSAGE. The longer tag-size allows for a more precise allocation of the tag to the corresponding transcript, because each additional base increases the precision of the annotation considerably.
296:, or miRNAs for short, are small (~22nt) segments of RNA which have been found to play a crucial role in gene regulation. One of the most commonly used methods for cloning and identifying miRNAs within a cell or tissue was developed in the Bartel Lab and published in a paper by Lau
300:(2001). Since then, several variant protocols have arisen, but most have the same basic format. The procedure is quite similar to SAGE: The small RNA are isolated, then linkers are added to each, and the RNA is converted to cDNA by
158:
These cDNA tag fragments (with adaptor primers and AE and TE recognition sites attached) are ligated, sandwiching the two tag sequences together, and flanking adaptors A and B at either end. These new constructs, called
233:(EST) and initiated more systematic sequencing of cDNAs as a project (starting with 600 brain cDNAs). The identification of ESTs proceeded rapidly, millions of ESTs now available in public databases (e.g.
264:. However, SAGE sampling is based on sequencing mRNA output, not on hybridization of mRNA output to probes, so transcription levels are measured more quantitatively than by microarray. In addition, the
1050:
Matsumura, H.; Bin Nasir, K. H.; Yoshida, K.; Ito, A.; Kahl, G. N.; Krüger, D. H.; Terauchi, R. (2006). "SuperSAGE array: the direct use of 26-base-pair transcript tags in oligonucleotide arrays".
148:, cDNA are cleaved using TE to remove them from the beads, leaving only a short "tag" of about 11 nucleotides of original cDNA (15 nucleotides minus the 4 corresponding to the AE recognition site).
240:
In 1995, the idea of reducing the tag length from 100 to 800 bp down to tag length of 10 to 22 bp helped reduce the cost of mRNA surveys. In this year, the original SAGE protocol was published by
280:
is more exact in SAGE because it involves directly counting the number of transcripts whereas spot intensities in microarrays fall in non-discrete gradients and are prone to background noise.
364:. Therefore, tag-based gene expression profiling also called "digital gene expression profiling" (DGE) can today provide most accurate transcription profiles that overcome the limitations of
881:
Gowda, M., et al. (2004). "Robust-LongSAGE (RL-SAGE): a substantially improved LongSAGE method for gene discovery and transcriptome analysis." Plant
Physiol 134(3): 890-897.
352:
tags. However, SuperSAGE avoids the bias observed during the less random LongSAGE 20 bp ditag-ligation. By direct sequencing with high-throughput sequencing techniques (
137:
called an anchoring enzyme (AE). The location of the cleavage site and thus the length of the remaining cDNA bound to the bead will vary for each individual cDNA (mRNA).
301:
765:
Adams MD, Kelley JM, Gocayne JD, et al. (June 1991). "Complementary DNA sequencing: expressed sequence tags and human genome project".
714:
Putney SD; Herlihy WC; Schimmel P (1983). "A new troponin T and cDNA clones for 13 different muscle proteins, found by shotgun sequencing".
1149:
166:
The ditags are then cleaved using the original AE, and allowed to link together with other ditags, which will be ligated to create a cDNA
229:
coworkers. and Putney et al. who sequenced 178 clones from a rabbit muscle cDNA library. In 1991 Adams and co-workers coined the term
1095:"Massive analysis of cDNA Ends (MACE) and miRNA expression profiling identifies proatherogenic pathways in chronic kidney disease"
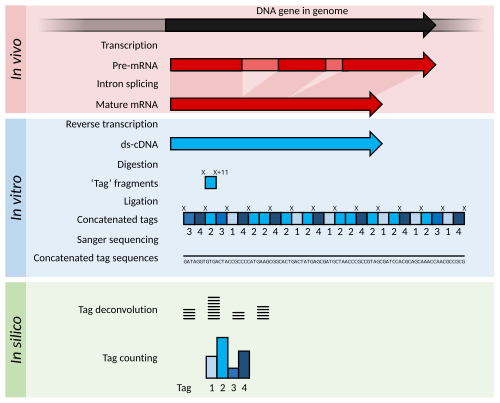
58:(at location 'X' and 'X'+11) to produce 11-nucleotide 'tag' fragments. These tags are concatenated and sequenced using long-read
891:
Matsumura, H.; Reich, S.; Ito, A.; Saitoh, H.; Kamoun, S.; Winter, P.; Kahl, G.; Reuter, M.; Krüger, D.; Terauchi, R. (2003).
317:, much smaller than previous LongSAGE insert size of 2 μg mRNA and using a lower number of ditag polymerase chain reactions (
133:
The cDNA is bound to
Streptavidin beads via interaction with the biotin attached to the primers, and is then cleaved using a
952:"Robust-LongSAGE (RL-SAGE): A Substantially Improved LongSAGE Method for Gene Discovery and Transcriptome Analysis"
518:"Robust-LongSAGE (RL-SAGE): a substantially improved LongSAGE method for gene discovery and transcriptome analysis"
248:. Although SAGE was originally conceived for use in cancer studies, it has been successfully used to describe the
396:
360:), hundred thousands or millions of tags can be analyzed simultaneously, producing very precise and quantitative
616:"Use of a cDNA library for studies on evolution and developmental expression of the chorion multigene families"
180:, and these sequences can be analysed with computer programs which quantify the recurrence of individual tags.
204:
Statistical methods can be applied to tag and count lists from different samples in order to determine which
145:
82:
872:
Saha, S., et al. (2002). "Using the transcriptome to annotate the genome." Nat
Biotechnol 20(5): 508-512.
276:
experiments are much cheaper to perform, so large-scale studies do not typically use SAGE. Quantifying
1169:
386:
353:
1144:
361:
318:
173:
These concatemers are then transformed into bacteria for amplification through bacterial replication.
47:
245:
189:
The output of SAGE is a list of short sequence tags and the number of times it is observed. Using
134:
67:
27:
819:
Velculescu VE; Zhang L; Vogelstein B; Kinzler KW. (1995). "Serial analysis of gene expression".
411:
230:
115:
43:
904:
828:
774:
723:
668:
8:
950:
Gowda, Malali; Jantasuriyarat, Chatchawan; Dean, Ralph A.; Wang, Guo-Liang (2004-03-01).
565:
Matsumura H; Ito A; Saitoh H; Winter P; Kahl G; Reuter M; Krüger DH; Terauchi R. (2005).
908:
832:
778:
727:
672:
1119:
1094:
1075:
1032:
852:
798:
747:
596:
498:
55:
984:
951:
927:
892:
691:
656:
614:
Sim GK; Kafatos FC; Jones CW; Koehler MD; Efstratiadis A; Maniatis T (December 1979).
542:
517:
1124:
1067:
1024:
989:
971:
932:
844:
790:
739:
696:
637:
632:
615:
600:
588:
583:
566:
547:
490:
455:
241:
209:
190:
176:
The cDNA concatemers can then be isolated and sequenced using modern high-throughput
86:
59:
51:
1079:
1036:
856:
802:
502:
1114:
1106:
1059:
1016:
979:
963:
922:
912:
836:
782:
751:
731:
686:
676:
627:
578:
537:
529:
482:
445:
840:
391:
277:
62:(different shades of blue indicate tags from different genes). The sequences are
406:
357:
273:
261:
160:
152:
473:
Saha S, et al. (2002). "Using the transcriptome to annotate the genome".
193:
a researcher can usually determine, with some confidence, from which original
66:
to find the frequency of each tag. The tag frequency can be used to report on
1163:
975:
459:
249:
177:
119:
90:
63:
39:
917:
786:
681:
450:
433:
1128:
1071:
1028:
993:
936:
893:"Gene expression analysis of plant host-pathogen interactions by SuperSAGE"
818:
592:
551:
494:
334:
31:
1020:
967:
848:
794:
743:
700:
533:
641:
365:
94:
of longer tags, enabling more confident identification of a source gene.
486:
371:
349:
348:
Like in the original SAGE protocol, so-called ditags are formed, using
167:
1110:
654:
515:
1063:
735:
35:
272:, so genes or gene variants which are not known can be discovered.
338:
1007:
Shendure, J. (2008). "The beginning of the end for microarrays?".
613:
19:
401:
234:
1155:
A review of the SAGE technique at the
Science Creative Quarterly
564:
42:
transcripts (red). The mRNA is extracted from the organism, and
1154:
163:, are then PCR amplified using anchor A and B specific primers.
111:
713:
293:
213:
1049:
655:
Sutcliffe JG; Milner RJ; Bloom FE; Lerner RA (August 1982).
949:
342:
322:
314:
265:
217:
205:
198:
194:
170:
with each ditag being separated by the AE recognition site.
127:
123:
107:
46:
is used to copy the mRNA into stable double-stranded–cDNA (
341:, to cut 26 bp long sequence tags from each transcript's
333:
SuperSAGE is a derivative of SAGE that uses the type III-
890:
252:
of other diseases and in a wide variety of organisms.
516:
Gowda M; Jantasuriyarat C; Dean RA; Wang GL. (2004).
372:
3'end mRNA sequencing, massive analysis of cDNA ends
260:
The general goal of the technique is similar to the
657:"Common 82-nucleotide sequence unique to brain RNA"
764:
208:are more highly expressed. For example, a normal
1161:
868:
866:
255:
897:Proceedings of the National Academy of Sciences
212:sample can be compared against a corresponding
102:Briefly, SAGE experiments proceed as follows:
884:
863:
151:The cleaved cDNA tags are then repaired with
54:; blue). In SAGE, the ds-cDNA is digested by
434:"Eukaryotic and prokaryotic gene structure"
431:
1118:
983:
926:
916:
690:
680:
631:
582:
541:
449:
1006:
558:
18:
307:
1162:
1092:
814:
812:
758:
509:
472:
432:Shafee, Thomas; Lowe, Rohan (2017).
283:
155:to produce blunt end cDNA fragments.
809:
466:
70:of the gene that the tag came from.
13:
268:sequences do not need to be known
220:tend to be more (or less) active.
75:Serial Analysis of Gene Expression
14:
1181:
1138:
584:10.1111/j.1462-5822.2004.00478.x
288:
26:Within the organisms, genes are
1086:
1043:
1000:
943:
875:
397:Cap analysis of gene expression
122:primers are used to synthesize
707:
648:
607:
425:
1:
1093:Zawada, Adam (January 2014).
419:
256:Comparison to DNA microarrays
89:to produce a snapshot of the
841:10.1126/science.270.5235.484
633:10.1016/0092-8674(79)90241-1
328:
7:
380:
184:
110:of an input sample (e.g. a
97:
16:Molecular biology technique
10:
1186:
387:High-throughput sequencing
354:next-generation sequencing
244:at the Oncology Center of
223:
201:) the tag was extracted.
661:Proc Natl Acad Sci U S A
362:gene expression profiles
246:Johns Hopkins University
135:restriction endonuclease
83:transcriptomic technique
918:10.1073/pnas.2536670100
787:10.1126/science.2047873
682:10.1073/pnas.79.16.4942
438:WikiJournal of Medicine
412:Expressed sequence tags
321:) to obtain a complete
231:expressed sequence tag
71:
1021:10.1038/nmeth0708-585
968:10.1104/pp.103.034496
534:10.1104/pp.103.034496
451:10.15347/wjm/2017.002
197:(and therefore which
116:reverse transcriptase
44:reverse transcriptase
22:
308:LongSAGE and RL-SAGE
114:) is isolated and a
87:molecular biologists
38:) to produce mature
909:2003PNAS..10015718M
903:(26): 15718–15723.
833:1995Sci...270..484V
779:1991Sci...252.1651A
728:1983Natur.302..718P
673:1982PNAS...79.4942S
487:10.1038/nbt0502-508
216:to determine which
56:restriction enzymes
1150:SAGE for Beginners
191:sequence databases
72:
1170:Molecular biology
1111:10.4161/epi.26931
284:Variant protocols
242:Victor Velculescu
60:Sanger sequencing
1177:
1133:
1132:
1122:
1090:
1084:
1083:
1064:10.1038/nmeth882
1047:
1041:
1040:
1004:
998:
997:
987:
956:Plant Physiology
947:
941:
940:
930:
920:
888:
882:
879:
873:
870:
861:
860:
816:
807:
806:
773:(5013): 1651–6.
762:
756:
755:
736:10.1038/302718a0
722:(5910): 718–21.
711:
705:
704:
694:
684:
652:
646:
645:
635:
611:
605:
604:
586:
562:
556:
555:
545:
513:
507:
506:
470:
464:
463:
453:
429:
278:gene expressions
24:Summary of SAGE.
1185:
1184:
1180:
1179:
1178:
1176:
1175:
1174:
1160:
1159:
1141:
1136:
1091:
1087:
1048:
1044:
1005:
1001:
948:
944:
889:
885:
880:
876:
871:
864:
827:(5235): 484–7.
817:
810:
763:
759:
712:
708:
653:
649:
612:
608:
563:
559:
514:
510:
471:
467:
430:
426:
422:
407:DNA microarrays
392:Transcriptomics
383:
374:
331:
310:
291:
286:
258:
226:
187:
100:
17:
12:
11:
5:
1183:
1173:
1172:
1158:
1157:
1152:
1147:
1140:
1139:External links
1137:
1135:
1134:
1105:(1): 161–172.
1085:
1052:Nature Methods
1042:
1009:Nature Methods
999:
962:(3): 890–897.
942:
883:
874:
862:
808:
757:
706:
667:(16): 4942–6.
647:
626:(4): 1303–16.
606:
571:Cell Microbiol
557:
508:
475:Nat Biotechnol
465:
423:
421:
418:
417:
416:
415:
414:
409:
404:
399:
389:
382:
379:
373:
370:
358:pyrosequencing
330:
327:
313:of 50 ng
309:
306:
290:
287:
285:
282:
262:DNA microarray
257:
254:
225:
222:
186:
183:
182:
181:
178:DNA sequencers
174:
171:
164:
156:
153:DNA polymerase
149:
144:After adaptor
142:
138:
131:
99:
96:
15:
9:
6:
4:
3:
2:
1182:
1171:
1168:
1167:
1165:
1156:
1153:
1151:
1148:
1146:
1143:
1142:
1130:
1126:
1121:
1116:
1112:
1108:
1104:
1100:
1096:
1089:
1081:
1077:
1073:
1069:
1065:
1061:
1058:(6): 469–74.
1057:
1053:
1046:
1038:
1034:
1030:
1026:
1022:
1018:
1014:
1010:
1003:
995:
991:
986:
981:
977:
973:
969:
965:
961:
957:
953:
946:
938:
934:
929:
924:
919:
914:
910:
906:
902:
898:
894:
887:
878:
869:
867:
858:
854:
850:
846:
842:
838:
834:
830:
826:
822:
815:
813:
804:
800:
796:
792:
788:
784:
780:
776:
772:
768:
761:
753:
749:
745:
741:
737:
733:
729:
725:
721:
717:
710:
702:
698:
693:
688:
683:
678:
674:
670:
666:
662:
658:
651:
643:
639:
634:
629:
625:
621:
617:
610:
602:
598:
594:
590:
585:
580:
576:
572:
568:
561:
553:
549:
544:
539:
535:
531:
527:
523:
522:Plant Physiol
519:
512:
504:
500:
496:
492:
488:
484:
481:(5): 508–12.
480:
476:
469:
461:
457:
452:
447:
443:
439:
435:
428:
424:
413:
410:
408:
405:
403:
400:
398:
395:
394:
393:
390:
388:
385:
384:
378:
369:
367:
363:
359:
355:
351:
346:
344:
340:
336:
326:
324:
320:
316:
305:
303:
299:
295:
289:miRNA cloning
281:
279:
275:
271:
267:
263:
253:
251:
250:transcriptome
247:
243:
238:
236:
232:
221:
219:
215:
211:
207:
202:
200:
196:
192:
179:
175:
172:
169:
165:
162:
157:
154:
150:
147:
143:
139:
136:
132:
129:
125:
121:
117:
113:
109:
105:
104:
103:
95:
92:
91:messenger RNA
88:
84:
80:
76:
69:
68:transcription
65:
61:
57:
53:
49:
45:
41:
37:
33:
29:
25:
21:
1102:
1098:
1088:
1055:
1051:
1045:
1015:(7): 585–7.
1012:
1008:
1002:
959:
955:
945:
900:
896:
886:
877:
824:
820:
770:
766:
760:
719:
715:
709:
664:
660:
650:
623:
619:
609:
574:
570:
560:
528:(3): 890–7.
525:
521:
511:
478:
474:
468:
441:
437:
427:
375:
347:
335:endonuclease
332:
311:
297:
292:
269:
259:
239:
227:
203:
188:
120:biotinylated
101:
78:
74:
73:
64:deconvoluted
23:
1099:Epigenetics
577:(1): 11–8.
567:"SuperSAGE"
366:microarrays
350:blunt-ended
337:EcoP15I of
28:transcribed
420:References
274:Microarray
168:concatemer
36:eukaryotes
976:1532-2548
601:221579149
460:2002-4436
329:SuperSAGE
325:library.
294:MicroRNAs
1164:Category
1129:24184689
1080:19160070
1072:16721381
1037:29682662
1029:18587314
994:15020752
937:14676315
857:16281846
803:13436211
593:15617519
552:15020752
503:12709815
495:11981567
381:See also
339:phage P1
270:a priori
185:Analysis
146:ligation
98:Overview
85:used by
1145:SAGEnet
1120:3928179
905:Bibcode
849:7570003
829:Bibcode
821:Science
795:2047873
775:Bibcode
767:Science
752:4364361
744:6687628
724:Bibcode
701:6956902
669:Bibcode
402:RNA-Seq
356:, i.e.
235:GenBank
224:History
81:) is a
32:spliced
1127:
1117:
1078:
1070:
1035:
1027:
992:
985:389912
982:
974:
935:
928:307634
925:
855:
847:
801:
793:
750:
742:
716:Nature
699:
692:346801
689:
642:519770
640:
599:
591:
550:
543:389912
540:
501:
493:
458:
302:RT-PCR
298:et al.
210:tissue
161:ditags
112:tumour
1076:S2CID
1033:S2CID
853:S2CID
799:S2CID
748:S2CID
597:S2CID
499:S2CID
444:(1).
218:genes
214:tumor
206:genes
126:from
1125:PMID
1068:PMID
1025:PMID
990:PMID
972:ISSN
933:PMID
845:PMID
791:PMID
740:PMID
697:PMID
638:PMID
620:Cell
589:PMID
548:PMID
491:PMID
456:ISSN
343:cDNA
323:cDNA
315:mRNA
266:mRNA
199:gene
195:mRNA
128:mRNA
124:cDNA
118:and
108:mRNA
106:The
79:SAGE
52:cDNA
40:mRNA
34:(in
30:and
1115:PMC
1107:doi
1060:doi
1017:doi
980:PMC
964:doi
960:134
923:PMC
913:doi
901:100
837:doi
825:270
783:doi
771:252
732:doi
720:302
687:PMC
677:doi
628:doi
579:doi
538:PMC
530:doi
526:134
483:doi
446:doi
319:PCR
237:).
1166::
1123:.
1113:.
1101:.
1097:.
1074:.
1066:.
1054:.
1031:.
1023:.
1011:.
988:.
978:.
970:.
958:.
954:.
931:.
921:.
911:.
899:.
895:.
865:^
851:.
843:.
835:.
823:.
811:^
797:.
789:.
781:.
769:.
746:.
738:.
730:.
718:.
695:.
685:.
675:.
665:79
663:.
659:.
636:.
624:18
622:.
618:.
595:.
587:.
573:.
569:.
546:.
536:.
524:.
520:.
497:.
489:.
479:20
477:.
454:.
440:.
436:.
368:.
48:ds
1131:.
1109::
1103:9
1082:.
1062::
1056:3
1039:.
1019::
1013:5
996:.
966::
939:.
915::
907::
859:.
839::
831::
805:.
785::
777::
754:.
734::
726::
703:.
679::
671::
644:.
630::
603:.
581::
575:7
554:.
532::
505:.
485::
462:.
448::
442:4
130:.
77:(
50:-
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.