1052:
microRNAs is comparable to that elsewhere in the non-coding DNA, implying evolution by neutral drift; however, older microRNAs have a much lower rate of change (often less than one substitution per hundred million years), suggesting that once a microRNA gains a function, it undergoes purifying selection. Individual regions within an miRNA gene face different evolutionary pressures, where regions that are vital for processing and function have higher levels of conservation. At this point, a microRNA is rarely lost from an animal's genome, although newer microRNAs (thus presumably non-functional) are frequently lost. In
704:
1537:, miRNA profiling of the FoxD1-derived cells not only comprehensively defined the transcriptional landscape of miRNAs that are critical for vascular development, but also identified key miRNAs that are likely to modulate the renal phenotype in its absence. These miRNAs include miRs‐10a, 18a, 19b, 24, 30c, 92a, 106a, 130a, 152, 181a, 214, 222, 302a, 370, and 381 that regulate Bcl2L11 (Bim) and miRs‐15b, 18a, 21, 30c, 92a, 106a, 125b‐5p, 145, 214, 222, 296‐5p and 302a that regulate p53-effector genes. Consistent with the profiling results, ectopic
5388:
1211:
57:
1398:. Furthermore, animal studies on specific miRNAs identified distinct roles for miRNAs both during heart development and under pathological conditions, including the regulation of key factors important for cardiogenesis, the hypertrophic growth response and cardiac conductance. Another role for miRNA in cardiovascular diseases is to use their expression levels for diagnosis, prognosis or risk stratification. miRNA's in animal models have also been linked to cholesterol metabolism and regulation.
532:. In this complex, DGCR8 orients the catalytic RNase III domain of Drosha to liberate hairpins from pri-miRNAs by cleaving RNA about eleven nucleotides from the hairpin base (one helical dsRNA turn into the stem). The product resulting has a two-nucleotide overhang at its 3' end; it has 3' hydroxyl and 5' phosphate groups. It is often termed as a pre-miRNA (precursor-miRNA). Sequence motifs downstream of the pre-miRNA that are important for efficient processing have been identified.
1374:, that bind the minor groove of AT-rich DNA stretches in specific regions of DNA. Human neoplasias, including thyroid, prostatic, cervical, colorectal, pancreatic and ovarian carcinomas, show a strong increase of HMGA1a and HMGA1b proteins. Transgenic mice with HMGA1 targeted to lymphoid cells develop aggressive lymphoma, showing that high HMGA1 expression is associated with cancers and that HMGA1 can act as an oncogene. HMGA2 protein specifically targets the promoter of
1417:(MMPs) are upregulated by d-flow, mediating pro-inflammatory and pro-angiogenic signals. These findings were observed in ligated carotid arteries of mice to mimic the effects of d-flow. Within 24 hours, pre-existing immature miR-712 formed mature miR-712 suggesting that miR-712 is flow-sensitive. Coinciding with these results, miR-712 is also upregulated in endothelial cells exposed to naturally occurring d-flow in the greater curvature of the aortic arch.
15738:
583:
468:
1900:
836:
390:
1617:, which results from blockage in the artery of the brain that carries oxygen-rich blood. The obstruction of the blood flow means the brain cannot receive necessary nutrients, such as oxygen and glucose, and remove wastes, such as carbon dioxide. miRNAs plays a role in posttranslational gene silencing by targeting genes in the pathogenesis of cerebral ischemia, such as the inflammatory, angiogenesis, and apoptotic pathway.
840:
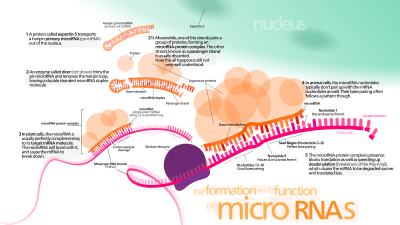
on the degradation of mRNA. 40S and 60S are light and heavy components of the ribosome, 80S is the assembled ribosome bound to mRNA, eIF4F is a translation initiation factor, PABC1 is the Poly-A binding protein, and "cap" is the mRNA cap structure needed for mRNA circularization (which can be the normal m7G-cap or modified A-cap). The initiation of mRNA can proceed in a cap-independent manner, through recruiting 40S to IRES (
771:. It has been demonstrated that given complete complementarity between the miRNA and target mRNA sequence, Ago2 can cleave the mRNA and lead to direct mRNA degradation. In the absence of complementarity, silencing is achieved by preventing translation. The relation of miRNA and its target mRNA can be based on the simple negative regulation of a target mRNA, but it seems that a common scenario is the use of a "coherent
1160:. microRNA maturation can be inhibited at several points by steric-blocking oligos. The miRNA target site of an mRNA transcript can also be blocked by a steric-blocking oligo. For the "in situ" detection of miRNA, LNA or Morpholino probes can be used. The locked conformation of LNA results in enhanced hybridization properties and increases sensitivity and selectivity, making it ideal for detection of short miRNA.
640:. This endoribonuclease interacts with 5' and 3' ends of the hairpin and cuts away the loop joining the 3' and 5' arms, yielding an imperfect miRNA:miRNA* duplex about 22 nucleotides in length. Overall hairpin length and loop size influence the efficiency of Dicer processing. The imperfect nature of the miRNA:miRNA* pairing also affects cleavage. Some of the G-rich pre-miRNAs can potentially adopt the
368:) and "as" (antisense)). However, the mature microRNA found from one arm of the hairpin is usually much more abundant than that found from the other arm, in which case, an asterisk following the name indicates the mature species found at low levels from the opposite arm of a hairpin. For example, miR-124 and miR-124* share a pre-miRNA hairpin, but much more miR-124 is found in the cell.
46:
381:
shows that each has, on average, roughly 400 conserved targets. Likewise, experiments show that a single miRNA species can reduce the stability of hundreds of unique messenger RNAs. Other experiments show that a single miRNA species may repress the production of hundreds of proteins, but that this repression often is relatively mild (much less than 2-fold).
295:
refers to the mature form of the miRNA, while the uncapitalized "mir-" refers to the pre-miRNA and the pri-miRNA. The genes encoding miRNAs are also named using the same three-letter prefix according to the conventions of the organism gene nomenclature. For examples, the official miRNAs gene names in some organisms are "
574:(A to I) transitions. RNA editing can halt nuclear processing (for example, of pri-miR-142, leading to degradation by the ribonuclease Tudor-SN) and alter downstream processes including cytoplasmic miRNA processing and target specificity (e.g., by changing the seed region of miR-376 in the central nervous system).
1413:, a cardiovascular disease of the arterial wall associated with lipid retention and inflammation. Non-laminar blood flow also correlates with development of atherosclerosis as mechanosenors of endothelial cells respond to the shear force of disturbed flow (d-flow). A number of pro-atherogenic genes including
1832:, mice become more insulin-sensitive and remarkably resistant to high fat diet-induced obesity and diabetes. Not only could let-7 inhibition prevent obesity and diabetes, it could also reverse and cure the condition. These experimental findings suggest that let-7 inhibition could represent a new therapy for
700:
may also influence strand choice. The other strand, called the passenger strand due to its lower levels in the steady state, is denoted with an asterisk (*) and is normally degraded. In some cases, both strands of the duplex are viable and become functional miRNA that target different mRNA populations.
1844:
miRNAs also play crucial roles in the regulation of complex enzymatic cascades including the hemostatic blood coagulation system. Large scale studies of functional miRNA targeting have recently uncovered rationale therapeutic targets in the hemostatic system. They have been directly linked to
Calcium
723:
and functions to interact with the 5' end of the guide strand. They bind the mature miRNA and orient it for interaction with a target mRNA. Some argonautes, for example human Ago2, cleave target transcripts directly; argonautes may also recruit additional proteins to achieve translational repression.
699:
Dicer processing of the pre-miRNA is thought to be coupled with unwinding of the duplex. Generally, only one strand is incorporated into the miRISC, selected on the basis of its thermodynamic instability and weaker base-pairing on the 5' end relative to the other strand. The position of the stem-loop
380:
Estimates of the average number of unique messenger RNAs that are targets for repression by a typical miRNA vary, depending on the estimation method, but multiple approaches show that mammalian miRNAs can have many unique targets. For example, an analysis of the miRNAs highly conserved in vertebrates
294:
Under a standard nomenclature system, names are assigned to experimentally confirmed miRNAs before publication. The prefix "miR" is followed by a dash and a number, the latter often indicating order of naming. For example, miR-124 was named and likely discovered prior to miR-456. A capitalized "miR-"
151:
miRNAs are abundant in many mammalian cell types miRNAs appear to target about 60% of the genes of humans and other mammals. Many miRNAs are evolutionarily conserved, which implies that they have important biological functions. For example, 90 families of miRNAs have been conserved since at least the
1456:
TIMP3 also decreases the expression of TNFα (a pro-inflammatory regulator) during turbulent flow. Activity of TNFα in turbulent flow was measured by the expression of TNFα-converting enzyme (TACE) in blood. TNFα decreased if miR-712 was inhibited or TIMP3 overexpressed, suggesting that miR-712
1112:
While researchers focused on miRNA expression in physiological and pathological processes, various technical variables related to microRNA isolation emerged. The stability of stored miRNA samples has been questioned. microRNAs degrade much more easily than mRNAs, partly due to their length, but also
839:
Interaction of microRNA with protein translation process. Several translation repression mechanisms are shown: M1) on the initiation process, preventing assembling of the initiation complex or recruiting the 40S ribosomal subunit; M2) on the ribosome assembly; M3) on the translation process; M7, M8)
818:
enzymes, a modification that may be associated with miRNA degradation. However, uridylation may also protect some miRNAs; the consequences of this modification are incompletely understood. Uridylation of some animal miRNAs has been reported. Both plant and animal miRNAs may be altered by addition of
376:
Plant miRNAs usually have near-perfect pairing with their mRNA targets, which induces gene repression through cleavage of the target transcripts. In contrast, animal miRNAs are able to recognize their target mRNAs by using as few as 6–8 nucleotides (the seed region) at the 5' end of the miRNA, which
275:
RNAs, except their expression patterns were usually inconsistent with a role in regulating the timing of development. This suggested that most might function in other types of regulatory pathways. At this point, researchers started using the term "microRNA" to refer to this class of small regulatory
1256:
Hepatocellular carcinoma cell proliferation may arise from miR-21 interaction with MAP2K3, a tumor repressor gene. Optimal treatment for cancer involves accurately identifying patients for risk-stratified therapy. Those with a rapid response to initial treatment may benefit from truncated treatment
784:
Turnover of mature miRNA is needed for rapid changes in miRNA expression profiles. During miRNA maturation in the cytoplasm, uptake by the
Argonaute protein is thought to stabilize the guide strand, while the opposite (* or "passenger") strand is preferentially destroyed. In what has been called a
344:
Pre-miRNAs, pri-miRNAs and genes that lead to 100% identical mature miRNAs but that are located at different places in the genome are indicated with an additional dash-number suffix. For example, the pre-miRNAs hsa-mir-194-1 and hsa-mir-194-2 lead to an identical mature miRNA (hsa-miR-194) but are
1272:
MicroRNAs have the potential to be used as tools or targets for treatment of different cancers. The specific microRNA, miR-506 has been found to work as a tumor antagonist in several studies. A significant number of cervical cancer samples were found to have downregulated expression of miR-506.
1252:
Furthermore, specific miRNAs may be associated with certain histological subtypes of colorectal cancer. For instance, expression levels of miR-205 and miR-373 have been shown to be increased in mucinous colorectal cancers and mucin-producing
Ulcerative Colitis-associated colon cancers, but not in
1469:
The human homolog of miR-712 was found on the RN45s homolog gene, which maintains similar miRNAs to mice. MiR-205 of humans share similar sequences with miR-712 of mice and is conserved across most vertebrates. MiR-205 and miR-712 also share more than 50% of the cell signaling targets, including
1051:
regions), but also by the duplication and modification of existing microRNAs. microRNAs can also form from inverted duplications of protein-coding sequences, which allows for the creation of a foldback hairpin structure. The rate of evolution (i.e. nucleotide substitution) in recently originated
864:
For partially complementary microRNAs to recognise their targets, nucleotides 2–7 of the miRNA (its 'seed region') must be perfectly complementary. Animal miRNAs inhibit protein translation of the target mRNA (this is present but less common in plants). Partially complementary microRNAs can also
1043:
markers because of their apparently low rate of evolution. microRNAs' origin as a regulatory mechanism developed from previous RNAi machinery that was initially used as a defense against exogenous genetic material such as viruses. Their origin may have permitted the development of morphological
946:
Unlike plant microRNAs, the animal microRNAs target diverse genes. However, genes involved in functions common to all cells, such as gene expression, have relatively fewer microRNA target sites and seem to be under selection to avoid targeting by microRNAs. There is a strong correlation between
1853:
miRNAs are considered to be key regulators of many developmental, homeostatic, and immune processes in plants. Their roles in plant development include shoot apical meristem development, leaf growth, flower formation, seed production, or root expansion. In addition, they play a complex role in
1452:
fibers, providing the structural support and recoil properties of arteries. These fibers play a critical role in regulation of vascular inflammation and permeability, which are important in the development of atherosclerosis. Expressed by endothelial cells, TIMP3 is the only ECM-bound TIMP. A
775:
loop", "mutual negative feedback loop" (also termed double negative loop) and "positive feedback/feed-forward loop". Some miRNAs work as buffers of random gene expression changes arising due to stochastic events in transcription, translation and protein stability. Such regulation is typically
1004:
in a number of diseases.Some researches show that mRNA cargo of exosomes may have a role in implantation, they can savage an adhesion between trophoblast and endometrium or support the adhesion by down regulating or up regulating expression of genes involved in adhesion/invasion.
668:
miRNA biogenesis in plants differs from animal biogenesis mainly in the steps of nuclear processing and export. Instead of being cleaved by two different enzymes, once inside and once outside the nucleus, both cleavages of the plant miRNA are performed by a Dicer homolog, called
1883:
miRNAs can bind to target messenger RNA (mRNA) transcripts of protein-coding genes and negatively control their translation or cause mRNA degradation. It is of key importance to identify the miRNA targets accurately. A comparison of the predictive performance of eighteen
1187:
Just as miRNA is involved in the normal functioning of eukaryotic cells, so has dysregulation of miRNA been associated with disease. A manually curated, publicly available database, miR2Disease, documents known relationships between miRNA dysregulation and human disease.
869:, causing mRNAs to be degraded sooner. While degradation of miRNA-targeted mRNA is well documented, whether or not translational repression is accomplished through mRNA degradation, translational inhibition, or a combination of the two is hotly debated. Recent work on
1058:, the net flux of miRNA genes has been predicted to be between 1.2 and 3.3 genes per million years. This makes them a valuable phylogenetic marker, and they are being looked upon as a possible solution to outstanding phylogenetic problems such as the relationships of
1862:
Viral microRNAs play an important role in the regulation of gene expression of viral and/or host genes to benefit the virus. Hence, miRNAs play a key role in host–virus interactions and pathogenesis of viral diseases. The expression of transcription activators by
1044:
innovation, and by making gene expression more specific and 'fine-tunable', permitted the genesis of complex organs and perhaps, ultimately, complex life. Rapid bursts of morphological innovation are generally associated with a high rate of microRNA accumulation.
695:
The mature miRNA is part of an active RNA-induced silencing complex (RISC) containing Dicer and many associated proteins. RISC is also known as a microRNA ribonucleoprotein complex (miRNP); A RISC with incorporated miRNA is sometimes referred to as a "miRISC."
1230:(BCR) signalling, B-cell migration/adhesion, cell-cell interactions in immune niches and the production and class-switching of immunoglobulins. MiRNAs influence B cell maturation, generation of pre-, marginal zone, follicular, B1, plasma and memory B cells.
1460:
Anti-miR-712 effectively suppresses d-flow-induced miR-712 expression and increases TIMP3 expression. Anti-miR-712 also inhibits vascular hyperpermeability, thereby significantly reducing atherosclerosis lesion development and immune cell infiltration.
724:
The human genome encodes eight argonaute proteins divided by sequence similarities into two families: AGO (with four members present in all mammalian cells and called E1F2C/hAgo in humans), and PIWI (found in the germline and hematopoietic stem cells).
377:
is not enough pairing to induce cleavage of the target mRNAs. Combinatorial regulation is a feature of miRNA regulation in animals. A given miRNA may have hundreds of different mRNA targets, and a given target might be regulated by multiple miRNAs.
1365:
proteins (HMGA1a, HMGA1b and HMGA2) are implicated in cancer, and expression of these proteins is regulated by microRNAs. HMGA expression is almost undetectable in differentiated adult tissues, but is elevated in many cancers. HMGA proteins are
1090:, which are critical to canonical microRNA biogenesis. It is the only animal thus far reported to be missing Drosha. MicroRNAs play a vital role in the regulation of gene expression in all non-ctenophore animals investigated thus far except for
860:
whereas plant miRNAs are usually complementary to coding regions of mRNAs. Perfect or near perfect base pairing with the target RNA promotes cleavage of the RNA. This is the primary mode of plant miRNAs. In animals the match-ups are imperfect.
659:
While the majority of miRNAs are located within the cell, some miRNAs, commonly known as circulating miRNAs or extracellular miRNAs, have also been found in extracellular environment, including various biological fluids and cell culture media.
160:
The first miRNA was discovered in the early 1990s. However, miRNAs were not recognized as a distinct class of biological regulators until the early 2000s. miRNA research revealed different sets of miRNAs expressed in different cell types and
1346:
gene, which reduces protein expression of MGMT. However, for 28% of glioblastomas, the MGMT protein is deficient, but the MGMT promoter is not methylated. In glioblastomas without methylated MGMT promoters, the level of microRNA miR-181d is
1253:
sporadic colonic adenocarcinoma that lack mucinous components. In-vitro studies suggested that miR-205 and miR-373 may functionally induce different features of mucinous-associated neoplastic progression in intestinal epithelial cells.
1393:
heart. This revealed that miRNAs play an essential role during its development. miRNA expression profiling studies demonstrate that expression levels of specific miRNAs change in diseased human hearts, pointing to their involvement in
673:(DL1). DL1 is expressed only in the nucleus of plant cells, which indicates that both reactions take place inside the nucleus. Before plant miRNA:miRNA* duplexes are transported out of the nucleus, its 3' overhangs are methylated by a
1103:
Across all species, in excess of 5000 different miRNAs had been identified by March 2010. Whilst short RNA sequences (50 – hundreds of base pairs) of a broadly comparable function occur in bacteria, bacteria lack true microRNAs.
977:
are thought to be a back channel of communication regulating expression levels between paralogous genes (genes having a similar structure indicating divergence from a common ancestral gene). Given the name "competing endogenous RNAs"
1453:
decrease in TIMP3 expression results in an increase of ECM degradation in the presence of d-flow. Consistent with these findings, inhibition of pre-miR712 increases expression of TIMP3 in cells, even when exposed to turbulent flow.
1140:, slides or chips with probes to hundreds or thousands of miRNA targets, so that relative levels of miRNAs can be determined in different samples. microRNAs can be both discovered and profiled by high-throughput sequencing methods (
1257:
regimens, showing the value of accurate disease response measures. Cell-free circulating miRNAs (cimiRNAs) are highly stable in blood, are overexpressed in cancer and are quantifiable within the diagnostic laboratory. In classical
12731:"Activation of CCAAT/enhancer-binding protein (C/EBP) alpha expression by C/EBP beta during adipogenesis requires a peroxisome proliferator-activated receptor-gamma-associated repression of HDAC1 at the C/ebp alpha gene promoter"
844:) located in 5'UTR region. The actual work of RNA silencing is performed by RISC in which the main catalytic subunit is one of the Argonaute proteins (AGO), and miRNA serves as a template for recognizing specific mRNA sequences.
1175:- and miRNA-data that predict miRNA-targets based on their base sequence. While this is usually done after miRNAs of interest have been detected (e. g. because of high expression levels), ideas for analysis tools that integrate
715:(Ago) protein family are central to RISC function. Argonautes are needed for miRNA-induced silencing and contain two conserved RNA binding domains: a PAZ domain that can bind the single stranded 3' end of the mature miRNA and a
801:, also known as Rat1p. In plants, SDN (small RNA degrading nuclease) family members degrade miRNAs in the opposite (3'-to-5') direction. Similar enzymes are encoded in animal genomes, but their roles have not been described.
762:
Gene silencing may occur either via mRNA degradation or preventing mRNA from being translated. For example, miR16 contains a sequence complementary to the AU-rich element found in the 3'UTR of many unstable mRNAs, such as
12683:
Skårn M, Namløs HM, Noordhuis P, Wang MY, Meza-Zepeda LA, Myklebost O (April 2012). "Adipocyte differentiation of human bone marrow-derived stromal cells is modulated by microRNA-155, microRNA-221, and microRNA-222".
7557:
Lee CT, Risom T, Strauss WM (April 2007). "Evolutionary conservation of microRNA regulatory circuits: an examination of microRNA gene complexity and conserved microRNA-target interactions through metazoan phylogeny".
1665:
of alcohol-dependent mice, suggesting the role of miRNA in orchestrating translational imbalances and the creation of differentially expressed proteins within an area of the brain where complex cognitive behavior and
363:
When two mature microRNAs originate from opposite arms of the same pre-miRNA and are found in roughly similar amounts, they are denoted with a -3p or -5p suffix. (In the past, this distinction was also made with "s"
1637:. miRNA global regulation of many downstream genes deems significant regarding the reorganization or synaptic connections or long term neural adaptations involving the behavioral change from alcohol consumption to
411:), the site-specific modification of RNA sequences to yield products different from those encoded by their DNA. This increases the diversity and scope of miRNA action beyond that implicated from the genome alone.
512:
A single pri-miRNA may contain from one to six miRNA precursors. These hairpin loop structures are composed of about 70 nucleotides each. Each hairpin is flanked by sequences necessary for efficient processing.
9436:
Mencía A, Modamio-Høybjør S, Redshaw N, Morín M, Mayo-Merino F, Olavarrieta L, et al. (May 2009). "Mutations in the seed region of human miR-96 are responsible for nonsyndromic progressive hearing loss".
165:
and multiple roles for miRNAs in plant and animal development and in many other biological processes. Aberrant miRNA expression are implicated in disease states. MiRNA-based therapies are under investigation.
681:(HEN1). The duplex is then transported out of the nucleus to the cytoplasm by a protein called Hasty (HST), an Exportin 5 homolog, where they disassemble and the mature miRNA is incorporated into the RISC.
8132:"Draft genome assemblies and predicted microRNA complements of the intertidal lophotrochozoans Patella vulgata (Mollusca, Patellogastropoda) and Spirobranchus (Pomatoceros) lamarcki (Annelida, Serpulida)"
152:
common ancestor of mammals and fish, and most of these conserved miRNAs have important functions, as shown by studies in which genes for one or more members of a family have been knocked out in mice.
443:
that in turn forms part of a several hundred nucleotide-long miRNA precursor termed a pri-miRNA. When a stem-loop precursor is found in the 3' UTR, a transcript may serve as a pri-miRNA and a mRNA.
2824:
Pasquinelli AE, Reinhart BJ, Slack F, Martindale MQ, Kuroda MI, Maller B, et al. (November 2000). "Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA".
2770:
Reinhart BJ, Slack FJ, Basson M, Pasquinelli AE, Bettinger JC, Rougvie AE, et al. (February 2000). "The 21-nucleotide let-7 RNA regulates developmental timing in
Caenorhabditis elegans".
321:
miRNAs with nearly identical sequences except for one or two nucleotides are annotated with an additional lower case letter. For example, miR-124a is closely related to miR-124b. For example:
10524:
Sgarra R, Rustighi A, Tessari MA, Di
Bernardo J, Altamura S, Fusco A, et al. (September 2004). "Nuclear phosphoproteins HMGA and their relationship with chromatin structure and cancer".
1378:, thus reducing expression of this DNA repair gene. ERCC1 protein expression was deficient in 100% of 47 evaluated colon cancers (though the extent to which HGMA2 was involved is not known).
4145:
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, et al. (February 2005). "Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs".
616:, recognizes a two-nucleotide overhang left by the RNase III enzyme Drosha at the 3' end of the pre-miRNA hairpin. Exportin-5-mediated transport to the cytoplasm is energy-dependent, using
7954:
Allen E, Xie Z, Gustafson AM, Sung GH, Spatafora JW, Carrington JC (December 2004). "Evolution of microRNA genes by inverted duplication of target gene sequences in
Arabidopsis thaliana".
1062:. On the other hand, in multiple cases microRNAs correlate poorly with phylogeny, and it is possible that their phylogenetic concordance largely reflects a limited sampling of microRNAs.
1028:
in both plants and animals, and are thought to be a vital and evolutionarily ancient component of gene regulation. While core components of the microRNA pathway are conserved between
11774:"Long-term potentiation and depression regulatory microRNAs were highlighted in Bisphenol A induced learning and memory impairment by microRNA sequencing and bioinformatics analysis"
1681:) and ultimately reducing its expression. BDNF plays a critical role in the formation and maturation of new neurons and synapses, suggesting a possible implication in synapse growth/
785:"Use it or lose it" strategy, Argonaute may preferentially retain miRNAs with many targets over miRNAs with few or no targets, leading to degradation of the non-targeting molecules.
1261:, plasma miR-21, miR-494, and miR-1973 are promising disease response biomarkers. Circulating miRNAs have the potential to assist clinical decision making and aid interpretation of
11151:
Yang B, Lin H, Xiao J, Lu Y, Luo X, Li B, et al. (April 2007). "The muscle-specific microRNA miR-1 regulates cardiac arrhythmogenic potential by targeting GJA1 and KCNJ2".
939:
It is often impossible to discern these mechanisms using experimental data about stationary reaction rates. Nevertheless, they are differentiated in dynamics and have different
1878:
9095:
Kaur H, Arora A, Wengel J, Maiti S (June 2006). "Thermodynamic, counterion, and hydration effects for the incorporation of locked nucleic acid nucleotides into DNA duplexes".
1928:
1923:
1777:, have been found during the adipogenic programming of both immortalized and primary hMSCs, suggesting that they act as negative regulators of differentiation. Conversely,
1641:
and/or dependence. Up to 35 different miRNAs have been found to be altered in the alcoholic post-mortem brain, all of which target genes that include the regulation of the
5614:
Boeckel JN, Reis SM, Leistner D, Thomé CE, Zeiher AM, Fichtlscherer S, et al. (April 2014). "From heart to toe: heart's contribution on peripheral microRNA levels".
1073:
to the animals. However, the difference in how these microRNAs function and the way they are processed suggests that microRNAs arose independently in plants and animals.
423:(Pol II). The polymerase often binds to a promoter found near the DNA sequence, encoding what will become the hairpin loop of the pre-miRNA. The resulting transcript is
1381:
Single
Nucleotide polymorphisms (SNPs) can alter the binding of miRNAs on 3'UTRs for example the case of hsa-mir181a and hsa-mir181b on the CDON tumor suppressor gene.
1335:
gene. However, up to 15% of MLH1-deficiencies in sporadic colon cancers appeared to be due to over-expression of the microRNA miR-155, which represses MLH1 expression.
1289:. A new strategy for tumor treatment is to inhibit tumor cell proliferation by repairing the defective miRNA pathway in tumors. Cancer is caused by the accumulation of
3053:
Wienholds E, Kloosterman WP, Miska E, Alvarez-Saavedra E, Berezikov E, de Bruijn E, et al. (July 2005). "MicroRNA expression in zebrafish embryonic development".
962:. dsRNAs targeting gene promoters can induce potent transcriptional activation of associated genes. This was demonstrated in human cells using synthetic dsRNAs termed
810:(thale cress), mature plant miRNAs appear to be stabilized by the addition of methyl moieties at the 3' end. The 2'-O-conjugated methyl groups block the addition of
3224:
Poy MN, Eliasson L, Krutzfeldt J, Kuwajima S, Ma X, Macdonald PE, et al. (November 2004). "A pancreatic islet-specific microRNA regulates insulin secretion".
1613:
According to the Center for
Disease Control and Prevention, Stroke is one of the leading causes of death and long-term disability in America. 87% of the cases are
9896:
MicroRNA-21 promotes hepatocellular carcinoma HepG2 cell proliferation through repression of mitogen-activated protein kinase-kinase 3. Guangxian Xu et al., 2013
11237:
van Rooij E, Sutherland LB, Qi X, Richardson JA, Hill J, Olson EN (April 2007). "Control of stress-dependent cardiac growth and gene expression by a microRNA".
1541:
was observed in the cellular derivatives of the FoxD1 derived progenitor lineage and reiterates the importance of renal stromal miRNAs in cellular homeostasis.
1533:. p53 protein levels remained unchanged, suggesting that FoxD1 stromal miRNAs directly repress p53-effector genes. Using a lineage tracing approach followed by
136:
pathway, except miRNAs derive from regions of RNA transcripts that fold back on themselves to form short hairpins, whereas siRNAs derive from longer regions of
1824:. When let-7 was ectopically overexpressed to mimic accelerated aging, mice became insulin-resistant, and thus more prone to high fat diet-induced obesity and
1444:(TIMP3). TIMPs normally regulate activity of matrix metalloproteinases (MMPs) which degrade the extracellular matrix (ECM). Arterial ECM is mainly composed of
1888:
algorithms is available. Large scale studies of functional miRNA targeting suggest that many functional miRNAs can be missed by target prediction algorithms.
12065:
1473:
When tested, d-flow decreased the expression of XRN1 in humans as it did in mice endothelial cells, indicating a potentially common role of XRN1 in humans.
4196:
Selbach M, Schwanhäusser B, Thierfelder N, Fang Z, Khanin R, Rajewsky N (September 2008). "Widespread changes in protein synthesis induced by microRNAs".
9132:"Integrating microRNA and mRNA expression profiles of neuronal progenitors to identify regulatory networks underlying the onset of cortical neurogenesis"
9698:"MicroRNAs and the Diagnosis of Childhood Acute Lymphoblastic Leukemia: Systematic Review, Meta-Analysis and Re-Analysis with Novel Small RNA-Seq Tools"
1359:
of MGMT mRNA). Thus, in 28% of glioblastomas, increased expression of miR-181d and reduced expression of DNA repair enzyme MGMT may be a causal factor.
404:
of other genes. These are usually, though not exclusively, found in a sense orientation, and thus usually are regulated together with their host genes.
13782:
8376:
1553:. Previous studies demonstrate that miRNAs can regulate neuronal differentiation and maturation at various stages. MiRNAs also play important roles in
3338:"Energizing miRNA research: a review of the role of miRNAs in lipid metabolism, with a prediction that miR-103/107 regulates human metabolic pathways"
1047:
New microRNAs are created in multiple ways. Novel microRNAs can originate from the random formation of hairpins in "non-coding" sections of DNA (i.e.
4304:
2946:
Lau NC, Lim LP, Weinstein EG, Bartel DP (October 2001). "An abundant class of tiny RNAs with probable regulatory roles in
Caenorhabditis elegans".
1009:
768:
1370:
of ~100 amino acid residues characterized by a modular sequence organization. These proteins have three highly positively charged regions, termed
1273:
Additionally, miR-506 works to promote apoptosis of cervical cancer cells, through its direct target hedgehog pathway transcription factor, Gli3.
13951:
13836:
11005:
Zhao Y, Samal E, Srivastava D (July 2005). "Serum response factor regulates a muscle-specific microRNA that targets Hand2 during cardiogenesis".
1854:
responses to various abiotic stresses comprising heat stress, low-temperature stress, drought stress, light stress, or gamma radiation exposure.
1311:
cases. However, altered expression of microRNAs, causing DNA repair deficiencies, are frequently associated with cancers and may be an important
815:
5271:
Winter J, Jung S, Keller S, Gregory RI, Diederichs S (March 2009). "Many roads to maturity: microRNA biogenesis pathways and their regulation".
5787:"Structural features of microRNA (miRNA) precursors and their relevance to miRNA biogenesis and small interfering RNA/short hairpin RNA design"
1677:
expression increased in the prefrontal cortex of alcohol-dependent rats, targeting the transcription factor brain-derived neurotrophic factor (
11116:
10610:"High mobility group A2 protein and its derivatives bind a specific region of the promoter of DNA repair gene ERCC1 and modulate its activity"
10477:"O6-Methylguanine DNA methyltransferase protein expression in tumor cells predicts outcome of temozolomide therapy in glioblastoma patients"
3544:
Fasanaro P, Greco S, Ivan M, Capogrossi MC, Martelli F (January 2010). "microRNA: emerging therapeutic targets in acute ischemic diseases".
3144:"bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila"
982:), these microRNAs bind to "microRNA response elements" on genes and pseudogenes and may provide another explanation for the persistence of
8326:"MicroRNAs and essential components of the microRNA processing machinery are not encoded in the genome of the ctenophore Mnemiopsis leidyi"
7109:
1721:
that ultimately result in addictive behaviors. Alternatively, overexpressing miR-382 resulted in attenuated drinking and the inhibition of
144:
may encode over 1900 miRNAs, However, only about 500 human miRNAs represent bona fide miRNAs in the manually curated miRNA gene database
13297:
Wong CE, Zhao YT, Wang XJ, Croft L, Wang ZH, Haerizadeh F, et al. (May 2011). "MicroRNAs in the shoot apical meristem of soybean".
4786:
Gregory RI, Chendrimada TP, Shiekhattar R (2006). "MicroRNA biogenesis: isolation and characterization of the microprocessor complex".
4735:
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J, et al. (September 2003). "The nuclear RNase III Drosha initiates microRNA processing".
10569:"The HMG-I oncogene causes highly penetrant, aggressive lymphoid malignancy in transgenic mice and is overexpressed in human leukemia"
6220:
494:
domains (blue and orange); a double-stranded RNA binding domain (yellow); and a connector/platform domain (gray) containing two bound
14112:
776:
achieved by the virtue of negative feedback loops or incoherent feed-forward loop uncoupling protein output from mRNA transcription.
11194:
Carè A, Catalucci D, Felicetti F, Bonci D, Addario A, Gallo P, et al. (May 2007). "MicroRNA-133 controls cardiac hypertrophy".
7752:
Wheeler BM, Heimberg AM, Moy VN, Sperling EA, Holstein TW, Heber S, et al. (2009). "The deep evolution of metazoan microRNAs".
2880:
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T (October 2001). "Identification of novel genes coding for small expressed RNAs".
652:. Although either strand of the duplex may potentially act as a functional miRNA, only one strand is usually incorporated into the
10047:"miR-506 acts as a tumor suppressor by directly targeting the hedgehog pathway transcription factor Gli3 in human cervical cancer"
15333:
1343:
12374:"Coordinated dysregulation of mRNAs and microRNAs in the rat medial prefrontal cortex following a history of alcohol dependence"
11415:"The atypical mechanosensitive microRNA-712 derived from pre-ribosomal RNA induces endothelial inflammation and atherosclerosis"
1429:
region 2 (ITS2). XRN1 is an exonuclease that degrades the ITS2 region during processing of RN45s. Reduction of XRN1 under d-flow
1301:
cause the accumulation of mutations, which can lead to cancer. Several genes involved in DNA repair are regulated by microRNAs.
15071:
13841:
13775:
4050:
Krek A, Grün D, Poy MN, Wolf R, Rosenberg L, Epstein EJ, et al. (May 2005). "Combinatorial microRNA target predictions".
173:
and including Lee and
Feinbaum. However, additional insight into its mode of action required simultaneously published work by
15023:
15018:
15013:
12640:
Romao JM, Jin W, Dodson MV, Hausman GJ, Moore SS, Guan LL (September 2011). "MicroRNA regulation in mammalian adipogenesis".
10207:
7400:
7025:
5513:
4803:
3688:
13008:"A novel rationale for targeting FXI: Insights from the hemostatic microRNA targetome for emerging anticoagulant strategies"
8087:
Caravas J, Friedrich M (June 2010). "Of mites and millipedes: recent progress in resolving the base of the arthropod tree".
6322:
Chatterjee S, Grosshans H (September 2009). "Active turnover modulates mature microRNA activity in Caenorhabditis elegans".
3632:"Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans"
1171:-expression is therefore often analyzed to check for miRNA-effects in their levels (e.g. in). Databases can be used to pair
11345:
Insull W (January 2009). "The pathology of atherosclerosis: plaque development and plaque responses to medical treatment".
1973:
15008:
15003:
11892:"Genetic ablation of Dicer in adult forebrain neurons results in abnormal tau hyperphosphorylation and neurodegeneration"
10098:
849:
280:
246:
RNA and additional "small temporal RNAs" might regulate the timing of development in diverse animals, including humans.
14987:
12593:"MicroRNA expression profile and functional analysis reveal that miR-382 is a critical novel gene of alcohol addiction"
11833:"Dicer loss in striatal neurons produces behavioral and neuroanatomical phenotypes in the absence of neurodegeneration"
9998:"MiR-506 suppresses proliferation and induces senescence by directly targeting the CDK4/6-FOXM1 axis in ovarian cancer"
9749:"Identification of miR-374a as a prognostic marker for survival in patients with early-stage nonsmall cell lung cancer"
9639:"A comprehensive investigation of histotype-specific microRNA and their variants in Stage I epithelial ovarian cancers"
1534:
456:
12423:"Integration of miRNA and protein profiling reveals coordinated neuroadaptations in the alcohol-dependent mouse brain"
1389:
The global role of miRNA function in the heart has been addressed by conditionally inhibiting miRNA maturation in the
201:, one of which was a ~22-nucleotide RNA that contained sequences partially complementary to multiple sequences in the
15419:
14982:
14977:
14972:
14967:
14962:
14957:
14952:
14947:
14942:
14937:
14932:
14927:
14922:
14917:
14912:
14907:
14902:
14897:
14892:
14887:
14882:
14877:
14872:
14867:
14862:
14857:
14852:
14847:
14842:
14837:
14832:
14827:
14822:
14817:
14812:
14807:
14802:
14797:
14781:
14776:
14771:
14766:
14761:
14756:
14751:
14746:
14741:
14736:
14731:
14726:
14721:
14716:
14711:
14706:
14701:
14696:
14691:
14686:
14681:
14676:
14671:
14666:
14650:
14645:
14640:
14635:
14630:
14625:
14620:
14615:
14610:
14605:
14600:
14595:
14585:
14580:
14575:
14570:
14560:
14555:
14550:
14545:
14540:
14535:
14530:
14514:
14509:
14504:
14499:
14494:
14489:
14484:
14479:
14474:
14469:
14464:
14459:
14454:
14449:
14444:
14439:
14434:
14429:
14424:
14419:
14414:
14409:
14399:
14394:
14389:
14384:
14379:
14374:
14369:
14353:
14343:
14338:
14333:
14328:
14323:
14318:
14313:
14293:
14288:
14283:
14243:
14217:
14212:
14207:
14202:
14187:
14182:
14177:
14172:
14167:
14152:
14142:
14127:
14122:
14072:
14062:
14052:
14047:
14032:
14017:
13768:
12967:"Large-scale identification of functional microRNA targeting reveals cooperative regulation of the hemostatic system"
12024:
8800:"Targeted inhibition of miRNA maturation with morpholinos reveals a role for miR-375 in pancreatic islet development"
1036:, miRNA repertoires in the two kingdoms appear to have emerged independently with different primary modes of action.
870:
516:
The double-stranded RNA (dsRNA) structure of the hairpins in a pri-miRNA is recognized by a nuclear protein known as
9281:"MicroRNA and mRNA integrated analysis (MMIA): a web tool for examining biological functions of microRNA expression"
15706:
15177:
13996:
13991:
13986:
13981:
13976:
13971:
13966:
13961:
13946:
13941:
13931:
13911:
13906:
13901:
13896:
13891:
13876:
13871:
13866:
13861:
13856:
13851:
13846:
10708:
Gibert B, Delloye-Bourgeois C, Gattolliat CH, Meurette O, Le Guernevel S, Fombonne J, et al. (November 2014).
2579:"Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets"
1958:
1241:
a levels may serve as an indicator of poor survival. Either high miR-185 or low miR-133b levels may correlate with
15711:
15105:
13821:
13816:
13806:
1356:
9048:"Revealing details: whole mount microRNA in situ hybridization protocol for zebrafish embryos and adult tissues"
3275:
Chen CZ, Li L, Lodish HF, Bartel DP (January 2004). "MicroRNAs modulate hematopoietic lineage differentiation".
14197:
14022:
8873:
8865:
7514:
Chen K, Rajewsky N (February 2007). "The evolution of gene regulation by transcription factors and microRNAs".
1485:-derived renal progenitor cells in a murine model resulted in a complex renal phenotype including expansion of
13095:"Programming pluripotent precursor cells derived from Xenopus embryos to generate specific tissues and organs"
7341:"A genome-wide microRNA screen identifies the microRNA-183/96/182 cluster as a modulator of circadian rhythms"
15608:
15494:
2674:"A Uniform System for the Annotation of Vertebrate microRNA Genes and the Evolution of the Human microRNAome"
1234:
690:
653:
84:
10475:
Spiegl-Kreinecker S, Pirker C, Filipits M, Lötsch D, Buchroithner J, Pichler J, et al. (January 2010).
8388:
5698:
Rana TM (January 2007). "Illuminating the silence: understanding the structure and function of small RNAs".
2245:
Jonas S, Izaurralde E (July 2015). "Towards a molecular understanding of microRNA-mediated gene silencing".
2202:
Jonas S, Izaurralde E (July 2015). "Towards a molecular understanding of microRNA-mediated gene silencing".
644:
structure as an alternative to the canonical stem-loop structure. For example, human pre-miRNA 92b adopts a
9796:"miR-185 and miR-133b deregulation is associated with overall survival and metastasis in colorectal cancer"
7602:"MicroRNAs and metazoan macroevolution: insights into canalization, complexity, and the Cambrian explosion"
1633:. Chronic alcohol abuse results in persistent changes in brain function mediated in part by alterations in
1561:(contributing to learning and memory). Elimination of miRNA formation in mice by experimental silencing of
1262:
1219:
841:
13725:
12531:"Chronic alcohol-induced microRNA-155 contributes to neuroinflammation in a TLR4-dependent manner in mice"
11933:"Role of miRNAs in Neurodegeneration: From Disease Cause to Tools of Biomarker Discovery and Therapeutics"
11058:"MicroRNA miR-133 represses HERG K+ channel expression contributing to QT prolongation in diabetic hearts"
1845:
homeostasis in the endoplasmic reticulum, which is critical in cell differentiation in early development.
1136:. Variations of this method achieve absolute or relative quantification. miRNAs can also be hybridized to
882:
15721:
15716:
15701:
15326:
13412:"Small RNA deep sequencing identifies microRNAs and other small noncoding RNAs from human herpesvirus 6B"
12770:
Jun-Hao ET, Gupta RR, Shyh-Chang N (March 2016). "Lin28 and let-7 in the Metabolic Physiology of Aging".
10899:"A signature pattern of stress-responsive microRNAs that can evoke cardiac hypertrophy and heart failure"
10710:"Regulation by miR181 family of the dependence receptor CDON tumor suppressive activity in neuroblastoma"
8131:
1913:
1638:
1426:
935:
Transcriptional inhibition through microRNA-mediated chromatin reorganization followed by gene silencing.
764:
11629:"MicroRNAs: Key Regulators in the Central Nervous System and Their Implication in Neurological Diseases"
6928:"Animal MicroRNAs confer robustness to gene expression and have a significant impact on 3'UTR evolution"
4526:"Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes"
407:
The DNA template is not the final word on mature miRNA production: 6% of human miRNAs show RNA editing (
49:
Pre-miRNA instead of Pri-miRNA in the first point of mechanism. Diagram of microRNA (miRNA) action with
15064:
14222:
14157:
14147:
14137:
4379:"Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs"
3912:
3910:
3908:
1565:
has led to pathological outcomes, such as reduced neuronal size, motor abnormalities (when silenced in
1514:
1505:
profiling of the FoxD1-Dicer knockout mouse model revealed ectopic upregulation of pro-apoptotic gene,
608:
Pre-miRNA hairpins are exported from the nucleus in a process involving the nucleocytoplasmic shuttler
11890:
Hébert SS, Papadopoulou AS, Smith P, Galas MC, Planel E, Silahtaroglu AN, et al. (October 2010).
8551:
Shingara J, Keiger K, Shelton J, Laosinchai-Wolf W, Powers P, Conrad R, et al. (September 2005).
6524:"Deep sequencing of tomato short RNAs identifies microRNAs targeting genes involved in fruit ripening"
5222:
Kawahara Y, Megraw M, Kreider E, Iizasa H, Valente L, Hatzigeorgiou AG, et al. (September 2008).
13921:
13916:
13659:"The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14"
10608:
Borrmann L, Schwanbeck R, Heyduk T, Seebeck B, Rogalla P, Bullerdiek J, et al. (December 2003).
7795:
Pashkovskiy PP, Ryazansky SS (June 2013). "Biogenesis, evolution, and functions of plant microRNAs".
4960:"Beyond secondary structure: primary-sequence determinants license pri-miRNA hairpins for processing"
2728:"The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14"
1594:
1199:
A mutation in the seed region of miR-184 causes hereditary keratoconus with anterior polar cataract.
1125:
11982:
Henshall DC, Hamer HM, Pasterkamp RJ, Goldstein DB, Kjems J, Prehn JH, et al. (December 2016).
10897:
van Rooij E, Sutherland LB, Liu N, Williams AH, McAnally J, Gerard RD, et al. (November 2006).
7483:
6771:"miRNA-mediated gene silencing by translational repression followed by mRNA deadenylation and decay"
5387:
3905:
15767:
15290:
15033:
15028:
13831:
13826:
13811:
8038:
Fahlgren N, Jogdeo S, Kasschau KD, Sullivan CM, Chapman EJ, Laubinger S, et al. (April 2010).
6522:
Moxon S, Jing R, Szittya G, Schwach F, Rusholme Pilcher RL, Moulton V, et al. (October 2008).
3387:"The RNaseIII enzyme Dicer is required for morphogenesis but not patterning of the vertebrate limb"
2422:
Fromm B, Domanska D, Høye E, Ovchinnikov V, Kang W, Aparicio-Puerta E, et al. (January 2020).
1817:
1414:
827:, stabilizes the molecule and plant miRNAs ending with an adenine residue have slower decay rates.
356:) miRNA. Other common prefixes include "v" for viral (miRNA encoded by a viral genome) and "d" for
226:
179:
7290:
Cuman C, Van Sinderen M, Gantier MP, Rainczuk K, Sorby K, Rombauts L, et al. (October 2015).
6714:"Ribosome profiling shows that miR-430 reduces translation before causing mRNA decay in zebrafish"
15777:
8459:
Liu CG, Calin GA, Volinia S, Croce CM (2008). "MicroRNA expression profiling using microarrays".
8275:
Ryan JF, Pang K, Schnitzler CE, Nguyen AD, Moreland RT, Simmons DK, et al. (December 2013).
1582:
1266:
432:
13344:"VIRmiRNA: a comprehensive resource for experimentally validated viral miRNAs and their targets"
11290:"Improved risk stratification in prevention by use of a panel of selected circulating microRNAs"
9480:
Hughes AE, Bradley DT, Campbell M, Lechner J, Dash DP, Simpson DA, et al. (November 2011).
3107:
Jones-Rhoades MW, Bartel DP, Bartel B (2006). "MicroRNAS and their regulatory roles in plants".
2101:"VIRmiRNA: a comprehensive resource for experimentally validated viral miRNAs and their targets"
1729:
upregulation in rat models of alcoholism, demonstrating the possibility of using miRNA-targeted
15741:
15434:
15319:
15115:
12372:
Tapocik JD, Solomon M, Flanigan M, Meinhardt M, Barbier E, Schank JR, et al. (June 2013).
11475:"Loss of Timp3 gene leads to abdominal aortic aneurysm formation in response to angiotensin II"
7478:
7251:"Circulating microRNAs as potential biomarkers for psychiatric and neurodegenerative disorders"
6168:
Meister G, Landthaler M, Peters L, Chen PY, Urlaub H, Lührmann R, et al. (December 2005).
5785:
Krol J, Sobczak K, Wilczynska U, Drath M, Jasinska A, Kaczynska D, et al. (October 2004).
3002:
Lee RC, Ambros V (October 2001). "An extensive class of small RNAs in Caenorhabditis elegans".
1750:
1578:
1574:
1324:
804:
Several miRNA modifications affect miRNA stability. As indicated by work in the model organism
790:
772:
617:
529:
185:
17:
11580:"Renal stromal miRNAs are required for normal nephrogenesis and glomerular mesangial survival"
10817:"Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2"
5959:"The long and short of inverted repeat genes in animals: microRNAs, mirtrons and hairpin RNAs"
1226:". In malignant B cells miRNAs participate in pathways fundamental to B cell development like
1222:. Many other miRNAs also have links with cancer and accordingly are sometimes referred to as "
1163:
High-throughput quantification of miRNAs is error prone, for the larger variance (compared to
15618:
15462:
15207:
15057:
13739:
12813:
Zhu H, Shyh-Chang N, Segrè AV, Shinoda G, Shah SP, Einhorn WS, et al. (September 2011).
12482:"microRNA-206 in rat medial prefrontal cortex regulates BDNF expression and alcohol drinking"
12480:
Tapocik JD, Barbier E, Flanigan M, Solomon M, Pincus A, Pilling A, et al. (March 2014).
9330:"Detection of simultaneous group effects in microRNA expression and related target gene sets"
8942:"Target protectors reveal dampening and balancing of Nodal agonist and antagonist by miR-430"
8842:
8277:"The genome of the ctenophore Mnemiopsis leidyi and its implications for cell type evolution"
4298:
3120:
2361:
1968:
903:
Nine mechanisms of miRNA action are described and assembled in a unified mathematical model:
743:(FMRP), Tudor staphylococcal nuclease-domain-containing protein (Tudor-SN), the putative DNA
472:
129:
118:
13047:
Berardi E, Pues M, Thorrez L, Sampaolesi M (October 2012). "miRNAs in ESC differentiation".
9794:
Akçakaya P, Ekelund S, Kolosenko I, Caramuta S, Ozata DM, Xie H, et al. (August 2011).
9529:
de Pontual L, Yao E, Callier P, Faivre L, Drouin V, Cariou S, et al. (September 2011).
7999:"Comparative analysis of the MIR319a microRNA locus in Arabidopsis and related Brassicaceae"
6119:
Mourelatos Z, Dostie J, Paushkin S, Sharma A, Charroux B, Abel L, et al. (March 2002).
3951:
1269:. They can be performed at each consultation to assess disease response and detect relapse.
15653:
15636:
15467:
15197:
13204:
12875:
12542:
12434:
12095:
11844:
11785:
11533:
11426:
11301:
11288:
Keller T, Boeckel JN, Groß S, Klotsche J, Palapies L, Leistner D, et al. (July 2017).
11246:
11014:
10910:
10766:
10755:"Targeted deletion of Dicer in the heart leads to dilated cardiomyopathy and heart failure"
10375:
10362:
Valeri N, Gasparini P, Fabbri M, Braconi C, Veronese A, Lovat F, et al. (April 2010).
9851:
9591:
9578:
Calin GA, Liu CG, Sevignani C, Ferracin M, Felli N, Dumitru CD, et al. (August 2004).
9341:
8953:
8190:
8143:
7854:
7352:
7054:
6782:
6725:
6378:
Morozova N, Zinovyev A, Nonne N, Pritchard LL, Gorban AN, Harel-Bellan A (September 2012).
6331:
6181:
6073:
5126:
5069:
4744:
4640:
4260:
4205:
4154:
3398:
3284:
3233:
3062:
3011:
2955:
2889:
2833:
2779:
2536:
2471:
Lim LP, Lau NC, Weinstein EG, Abdelhakim A, Yekta S, Rhoades MW, et al. (April 2003).
2301:
1879:
List of RNA structure prediction software § Inter molecular interactions: MicroRNA:UTR
1722:
1710:
1348:
1092:
1054:
963:
806:
13566:
Ambros V, Bartel B, Bartel DP, Burge CB, Carrington JC, Chen X, et al. (March 2003).
10956:
Tatsuguchi M, Seok HY, Callis TE, Thomson JM, Chen JF, Newman M, et al. (June 2007).
10659:"Deficient expression of DNA repair enzymes in early progression to sporadic colon cancer"
10323:"Immunohistochemical analysis reveals high frequency of PMS2 defects in colorectal cancer"
10321:
Truninger K, Menigatti M, Luz J, Russell A, Haider R, Gebbers JO, et al. (May 2005).
6663:
Eulalio A, Huntzinger E, Nishihara T, Rehwinkel J, Fauser M, Izaurralde E (January 2009).
6204:
5649:
Lelandais-Brière C, Sorin C, Declerck M, Benslimane A, Crespi M, Hartmann C (March 2010).
5539:"A potassium ion-dependent RNA structural switch regulates human pre-miRNA 92b maturation"
3712:
Ambros V, Bartel B, Bartel DP, Burge CB, Carrington JC, Chen X, et al. (March 2003).
2910:
2354:
819:
adenine (A) residues to the 3' end of the miRNA. An extra A added to the end of mammalian
348:
Species of origin is designated with a three-letter prefix, e.g., hsa-miR-124 is a human (
197:
miRNA, they found that instead of producing an mRNA encoding a protein, it produced short
8:
15648:
15631:
15489:
15372:
15187:
15136:
13195:
Choudhary A, Kumar A, Kaur H, Kaur N (May 2021). "MiRNA: the taskmaster of plant world".
12864:"Control of glucose homeostasis and insulin sensitivity by the Let-7 family of microRNAs"
10750:
9531:"Germline deletion of the miR-17~92 cluster causes skeletal and growth defects in humans"
8553:"An optimized isolation and labeling platform for accurate microRNA expression profiling"
6967:
He L, Hannon GJ (July 2004). "MicroRNAs: small RNAs with a big role in gene regulation".
6828:"Transcriptional inhibiton of Hoxd4 expression by miRNA-10a in human breast cancer cells"
5352:
1978:
1938:
1865:
1730:
1682:
1573:
neurons). Altered miRNA expression has been found in neurodegenerative diseases (such as
1558:
1316:
1145:
1141:
997:
563:
137:
13208:
12879:
12546:
12438:
12100:
12082:
Feng J, Sun G, Yan J, Noltner K, Li W, Buzin CH, et al. (July 2009). Reif A (ed.).
11848:
11831:
Cuellar TL, Davis TH, Nelson PT, Loeb GB, Harfe BD, Ullian E, et al. (April 2008).
11789:
11537:
11430:
11305:
11250:
11018:
10914:
10770:
10657:
Facista A, Nguyen H, Lewis C, Prasad AR, Ramsey L, Zaitlin B, et al. (April 2012).
10426:
Zhang W, Zhang J, Hoadley K, Kushwaha D, Ramakrishnan V, Li S, et al. (June 2012).
10379:
9855:
9595:
9580:"MicroRNA profiling reveals distinct signatures in B cell chronic lymphocytic leukemias"
9345:
8957:
8194:
8179:"The Ectocarpus genome and the independent evolution of multicellularity in brown algae"
8147:
7858:
7356:
7058:
6786:
6729:
6632:
6607:
6429:"Prediction and identification of Arabidopsis thaliana microRNAs and their mRNA targets"
6335:
6185:
6077:
5130:
5073:
4748:
4644:
4264:
4209:
4158:
3472:
3445:
3402:
3288:
3237:
3066:
3015:
2959:
2893:
2837:
2783:
2689:
2540:
2305:
1717:, a transcription factor that activates a series of transcription events in the nucleus
15542:
15537:
15412:
15217:
15212:
13751:
13640:
13592:
13567:
13538:
13511:
13487:
13460:
13436:
13411:
13368:
13343:
13274:
13247:
13228:
13172:
13145:
13121:
13094:
13072:
12898:
12863:
12839:
12814:
12795:
12665:
12617:
12592:
12565:
12530:
12506:
12481:
12457:
12422:
12398:
12373:
12349:
12324:
12295:
12270:
12246:
12237:
12221:
12171:
12144:
12120:
12083:
12055:
11959:
11932:
11867:
11832:
11808:
11773:
11754:
11706:
11679:
11655:
11628:
11604:
11579:
11557:
11501:
11474:
11447:
11414:
11322:
11289:
11270:
11219:
11176:
11133:
11125:
11084:
11057:
11038:
10982:
10957:
10933:
10898:
10856:
Thum T, Galuppo P, Wolf C, Fiedler J, Kneitz S, van Laake LW, et al. (July 2007).
10815:
Zhao Y, Ransom JF, Li A, Vedantham V, von Drehle M, Muth AN, et al. (April 2007).
10789:
10754:
10685:
10658:
10567:
Xu Y, Sumter TF, Bhattacharya R, Tesfaye A, Fuchs EJ, Wood LJ, et al. (May 2004).
10549:
10501:
10476:
10452:
10427:
10398:
10363:
10298:
10273:
10249:
10224:
10132:
10110:
10076:
10022:
9997:
9978:
9874:
9839:
9776:
9724:
9697:
9678:
9637:
Velle A, Pesenti C, Grassi T, Beltrame L, Martini P, Jaconi M, et al. (May 2023).
9555:
9530:
9506:
9481:
9462:
9413:
9388:
9364:
9329:
9305:
9280:
9256:
9231:
9207:
9182:
9158:
9131:
9072:
9047:
9023:
8998:
8979:
8917:
8892:
8826:
8799:
8775:
8750:
8726:
8701:
8628:
8601:
8577:
8552:
8528:
8503:
8502:
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT, et al. (November 2005).
8484:
8436:
8411:
8352:
8325:
8301:
8276:
8252:
8227:
8177:
Cock JM, Sterck L, Rouzé P, Scornet D, Allen AE, Amoutzias G, et al. (June 2010).
8112:
8064:
8039:
7979:
7931:
7906:
7877:
7842:
7820:
7777:
7726:
7699:
7675:
7650:
7631:
7539:
7446:
7421:
7388:
7375:
7340:
7339:
Zhou L, Miller C, Miraglia LJ, Romero A, Mure LS, Panda S, et al. (January 2021).
7316:
7291:
7226:
7201:
7174:
7149:
7077:
7042:
6992:
6903:
6878:
6854:
6827:
6803:
6770:
6746:
6713:
6689:
6664:
6645:
6548:
6523:
6404:
6379:
6355:
6299:
6274:
6096:
6061:
6037:
6010:
5983:
5958:
5934:
5909:
5723:
5675:
5650:
5473:
5448:
5296:
5248:
5223:
5196:
5171:
5147:
5114:
5090:
5057:
5038:
4984:
4959:
4768:
4712:
4687:
4663:
4628:
4550:
4525:
4501:
4476:
4457:
4403:
4378:
4281:
4248:
4229:
4178:
4119:
4094:
4075:
4032:
3984:
3959:
3889:
3863:"Small RNA discovery in the interaction between barley and the powdery mildew pathogen"
3862:
3838:
3811:
3787:
3762:
3738:
3713:
3607:
3580:
3521:
3496:
3421:
3386:
3362:
3337:
3318:
3257:
3086:
3035:
2979:
2923:
2857:
2803:
2698:
2673:
2649:
2624:
2448:
2423:
2399:
2374:
2373:
Alles J, Fehlmann T, Fischer U, Backes C, Galata V, Minet M, et al. (April 2019).
2332:
2289:
2270:
2227:
2179:
2154:
2125:
2100:
2030:
2005:
1805:
1778:
1654:
1506:
1286:
1025:
897:
740:
444:
12000:
11983:
10874:
10857:
10634:
10609:
9747:
Võsa U, Vooder T, Kolde R, Fischer K, Välk K, Tõnisson N, et al. (October 2011).
9614:
9579:
8677:
8652:
6455:
6428:
6145:
6120:
5885:
5868:
5844:
5827:
4878:"Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex"
4853:
4828:
4629:"Characterization and identification of microRNA core promoters in four model species"
4604:
4579:
4347:
4322:
3935:
3918:
3160:
3143:
2549:
2524:
2497:
2472:
2366:
2076:
2059:
1793:
significantly inhibited adipogenesis and repressed induction of the master regulators
1673:
miRNAs can be either upregulated or downregulated in response to chronic alcohol use.
1425:
Pre-mRNA sequence of miR-712 is generated from the murine ribosomal RN45s gene at the
1156:
oligo or a 2'-O-methyl RNA oligo. A specific miRNA can be silenced by a complementary
889:
848:
The function of miRNAs appears to be in gene regulation. For that purpose, a miRNA is
703:
439:. Animal miRNAs are initially transcribed as part of one arm of an ~80 nucleotide RNA
283:. In this disorder, the miRNAs have a dual role working as both tumor suppressors and
177:'s team, including Wightman and Ha. These groups published back-to-back papers on the
125:
In cells of humans and other animals, miRNAs primarily act by destabilizing the mRNA.
15762:
15626:
15588:
15581:
15532:
15484:
15222:
15131:
13680:
13675:
13658:
13632:
13609:
13597:
13543:
13492:
13441:
13373:
13324:
13319:
13279:
13232:
13220:
13177:
13126:
13064:
13029:
13024:
13007:
12988:
12944:
12903:
12844:
12787:
12752:
12711:
12657:
12622:
12570:
12511:
12462:
12403:
12354:
12340:
12300:
12251:
12176:
12125:
12047:
12005:
11964:
11913:
11872:
11813:
11746:
11711:
11660:
11609:
11549:
11506:
11452:
11362:
11327:
11262:
11211:
11168:
11108:
11089:
11030:
10987:
10938:
10879:
10838:
10794:
10731:
10690:
10639:
10590:
10541:
10506:
10457:
10403:
10344:
10303:
10254:
10203:
10180:
10136:
10124:
10094:
10068:
10027:
9970:
9929:
9879:
9840:"MiR-205 and MiR-373 Are Associated with Aggressive Human Mucinous Colorectal Cancer"
9817:
9768:
9729:
9682:
9670:
9619:
9560:
9511:
9482:"Mutation altering the miR-184 seed region causes familial keratoconus with cataract"
9454:
9418:
9389:"miR2Disease: a manually curated database for microRNA deregulation in human disease"
9369:
9310:
9261:
9212:
9163:
9112:
9077:
9028:
8971:
8922:
8854:
8831:
8780:
8731:
8682:
8633:
8582:
8533:
8476:
8441:
8410:
Tjaden B, Goodwin SS, Opdyke JA, Guillier M, Fu DX, Gottesman S, et al. (2006).
8357:
8306:
8257:
8208:
8159:
8104:
8069:
8020:
7971:
7936:
7882:
7812:
7769:
7765:
7731:
7680:
7623:
7575:
7531:
7496:
7451:
7392:
7380:
7321:
7272:
7231:
7179:
7101:
7082:
7021:
6984:
6949:
6908:
6859:
6808:
6751:
6694:
6637:
6588:
6553:
6501:
6460:
6409:
6347:
6304:
6250:
6209:
6150:
6101:
6042:
5988:
5939:
5890:
5849:
5808:
5767:
5715:
5680:
5631:
5560:
5519:
5509:
5478:
5429:
5376:
5331:
5314:
Ohman M (October 2007). "A-to-I editing challenger or ally to the microRNA process".
5300:
5288:
5253:
5201:
5152:
5095:
5030:
4989:
4940:
4899:
4858:
4809:
4799:
4760:
4717:
4668:
4609:
4555:
4506:
4449:
4444:
4427:
4408:
4352:
4286:
4221:
4170:
4124:
4067:
4024:
3989:
3940:
3894:
3843:
3792:
3743:
3694:
3684:
3653:
3648:
3631:
3612:
3561:
3557:
3526:
3477:
3426:
3367:
3310:
3249:
3206:
3165:
3124:
3078:
3027:
2971:
2915:
2849:
2795:
2749:
2744:
2727:
2703:
2654:
2600:
2554:
2502:
2453:
2404:
2337:
2262:
2219:
2184:
2130:
2081:
2035:
1698:
1686:
1662:
1441:
1246:
1233:
Another role for miRNA in cancers is to use their expression level for prognosis. In
1078:
990:
974:
674:
596:
521:
500:
420:
33:
13742:
that were involved in or affected the transcription of microRNAs from ChIP-seq data.
13644:
13248:"The Critical Role of miRNAs in Regulation of Flowering Time and Flower Development"
12669:
12060:
11223:
10553:
10080:
9466:
8983:
8116:
7983:
7824:
7781:
7635:
7267:
7250:
5042:
4461:
4079:
4036:
3975:
3597:
3090:
3039:
2983:
2927:
2274:
2231:
1196:
A mutation in the seed region of miR-96 causes hereditary progressive hearing loss.
15452:
14590:
14298:
14268:
13670:
13624:
13587:
13579:
13533:
13523:
13482:
13472:
13431:
13423:
13363:
13355:
13314:
13306:
13269:
13259:
13212:
13167:
13157:
13116:
13106:
13076:
13056:
13019:
12978:
12934:
12893:
12883:
12834:
12826:
12799:
12779:
12742:
12701:
12693:
12649:
12612:
12604:
12560:
12550:
12501:
12497:
12493:
12452:
12442:
12393:
12385:
12344:
12336:
12290:
12282:
12241:
12233:
12166:
12156:
12115:
12105:
12039:
11995:
11954:
11944:
11903:
11862:
11852:
11803:
11793:
11758:
11738:
11701:
11691:
11650:
11640:
11599:
11591:
11541:
11496:
11486:
11442:
11434:
11354:
11317:
11309:
11274:
11254:
11203:
11180:
11160:
11100:
11079:
11069:
11042:
11022:
10977:
10969:
10958:"Expression of microRNAs is dynamically regulated during cardiomyocyte hypertrophy"
10928:
10918:
10869:
10858:"MicroRNAs in the human heart: a clue to fetal gene reprogramming in heart failure"
10828:
10784:
10774:
10721:
10680:
10670:
10629:
10621:
10580:
10533:
10496:
10488:
10447:
10439:
10393:
10383:
10334:
10293:
10285:
10244:
10236:
10198:
Lodish H, Berk A, Kaiser CA, Krieger M, Bretscher A, Ploegh H, et al. (2016).
10170:
10114:
10058:
10017:
10009:
9982:
9960:
9919:
9869:
9859:
9807:
9780:
9760:
9719:
9709:
9660:
9650:
9609:
9599:
9550:
9542:
9501:
9493:
9446:
9408:
9400:
9359:
9349:
9300:
9292:
9251:
9243:
9202:
9194:
9153:
9143:
9104:
9067:
9059:
9018:
9010:
8961:
8912:
8904:
8846:
8821:
8811:
8770:
8762:
8721:
8713:
8672:
8664:
8623:
8613:
8572:
8564:
8523:
8515:
8488:
8468:
8431:
8423:
8347:
8337:
8296:
8288:
8247:
8239:
8198:
8151:
8096:
8059:
8051:
8010:
7963:
7926:
7918:
7872:
7862:
7804:
7761:
7721:
7711:
7670:
7662:
7613:
7567:
7523:
7488:
7441:
7433:
7370:
7360:
7311:
7303:
7262:
7221:
7213:
7169:
7161:
7130:
7122:
7093:
7072:
7062:
6996:
6976:
6939:
6898:
6890:
6849:
6839:
6798:
6790:
6741:
6733:
6684:
6676:
6649:
6627:
6619:
6580:
6543:
6535:
6491:
6450:
6440:
6399:
6391:
6359:
6339:
6294:
6286:
6240:
6199:
6189:
6140:
6132:
6091:
6081:
6032:
6022:
5978:
5970:
5929:
5921:
5880:
5839:
5798:
5757:
5727:
5707:
5670:
5662:
5623:
5591:
5580:"Extracellular/Circulating MicroRNAs: Release Mechanisms, Functions and Challenges"
5555:
5550:
5538:
5501:
5468:
5460:
5419:
5368:
5323:
5280:
5243:
5235:
5191:
5183:
5142:
5134:
5085:
5077:
5020:
4979:
4971:
4930:
4889:
4848:
4840:
4791:
4772:
4752:
4707:
4699:
4658:
4648:
4599:
4591:
4545:
4537:
4496:
4488:
4439:
4398:
4390:
4342:
4334:
4276:
4268:
4233:
4213:
4182:
4162:
4114:
4106:
4059:
4016:
3979:
3971:
3930:
3884:
3874:
3833:
3823:
3782:
3774:
3761:
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ (January 2006).
3733:
3725:
3676:
3643:
3602:
3592:
3553:
3516:
3508:
3467:
3457:
3416:
3406:
3357:
3349:
3322:
3300:
3292:
3261:
3241:
3196:
3155:
3116:
3070:
3019:
2963:
2905:
2897:
2861:
2841:
2807:
2787:
2739:
2693:
2685:
2644:
2636:
2590:
2544:
2523:
Lagos-Quintana M, Rauhut R, Yalcin A, Meyer J, Lendeckel W, Tuschl T (April 2002).
2492:
2484:
2443:
2435:
2394:
2386:
2327:
2317:
2309:
2254:
2211:
2174:
2166:
2120:
2112:
2071:
2025:
2017:
1963:
1790:
1786:
1774:
1770:
1602:
1598:
1390:
1258:
1133:
1013:
881:
cultured cells, shows that translational repression is caused by the disruption of
707:
AGO2 (grey) in complex with a microRNA (light blue) and its target mRNA (dark blue)
491:
162:
133:
11561:
10585:
10568:
10537:
10428:"miR-181d: a predictive glioblastoma biomarker that downregulates MGMT expression"
9924:
9907:
9181:
Grimson A, Farh KK, Johnston WK, Garrett-Engele P, Lim LP, Bartel DP (July 2007).
7543:
7292:"Human Blastocyst Secreted microRNA Regulate Endometrial Epithelial Cell Adhesion"
6227:
Jing Q, Huang S, Guth S, Zarubin T, Motoyama A, Chen J, et al. (March 2005).
5025:
5008:
56:
15576:
15547:
15259:
14001:
13729:
13719:: Integrative resource for microRNA and transcription factors analysis in cancer.
12555:
12447:
12323:
Lewohl JM, Nunez YO, Dodd PR, Tiwari GR, Harris RA, Mayfield RD (November 2011).
12110:
11798:
11358:
10339:
10322:
10289:
9864:
9354:
9198:
8850:
8816:
8798:
Kloosterman WP, Lagendijk AK, Ketting RF, Moulton JD, Plasterk RH (August 2007).
8600:
Buermans HP, Ariyurek Y, van Ommen G, den Dunnen JT, 't Hoen PA (December 2010).
8155:
7700:"Vive la différence: biogenesis and evolution of microRNAs in plants and animals"
7217:
7015:
6584:
6480:"MicroRNA-196 inhibits HOXB8 expression in myeloid differentiation of HL60 cells"
5627:
5500:. Current Topics in Microbiology and Immunology. Vol. 320. pp. 99–116.
5327:
5187:
4935:
4918:
4703:
4653:
2672:
Fromm B, Billipp T, Peck LE, Johansen M, Tarver JE, King BL, et al. (2015).
1918:
1702:
1667:
1634:
1614:
1410:
1294:
1227:
1210:
1149:
1087:
955:
893:
795:
517:
428:
111:
13060:
10973:
8891:
Gebert LF, Rebhan MA, Crivelli SE, Denzler R, Stoffel M, Hall J (January 2014).
8602:"New methods for next generation sequencing based microRNA expression profiling"
7307:
7043:"MicroRNA-373 induces expression of genes with complementary promoter sequences"
5505:
3680:
3675:. Advances in Experimental Medicine and Biology. Vol. 889. pp. 23–40.
3353:
15668:
15571:
15388:
15038:
13216:
12868:
Proceedings of the National Academy of Sciences of the United States of America
12830:
12286:
12084:"Evidence for X-chromosomal schizophrenia associated with microRNA alterations"
11837:
Proceedings of the National Academy of Sciences of the United States of America
11473:
Basu R, Fan D, Kandalam V, Lee J, Das SK, Wang X, et al. (December 2012).
11413:
Son DJ, Kumar S, Takabe W, Kim CW, Ni CW, Alberts-Grill N, et al. (2013).
11313:
10903:
Proceedings of the National Academy of Sciences of the United States of America
10833:
10816:
10759:
Proceedings of the National Academy of Sciences of the United States of America
10368:
Proceedings of the National Academy of Sciences of the United States of America
10119:
9908:"Plasma microRNA are disease response biomarkers in classical Hodgkin lymphoma"
9584:
Proceedings of the National Academy of Sciences of the United States of America
9497:
7847:
Proceedings of the National Academy of Sciences of the United States of America
7666:
7345:
Proceedings of the National Academy of Sciences of the United States of America
7165:
7047:
Proceedings of the National Academy of Sciences of the United States of America
6944:
6927:
6245:
6228:
6066:
Proceedings of the National Academy of Sciences of the United States of America
5925:
5666:
5356:
4975:
4894:
4877:
4795:
3861:
Hunt M, Banerjee S, Surana P, Liu M, Fuerst G, Mathioni S, et al. (2019).
3391:
Proceedings of the National Academy of Sciences of the United States of America
2595:
2578:
2170:
2021:
1905:
1658:
1554:
1550:
1395:
983:
959:
736:
720:
198:
96:
72:
13745:
13410:
Tuddenham L, Jung JS, Chane-Woon-Ming B, Dölken L, Pfeffer S (February 2012).
13359:
12783:
12145:"Schizophrenia is associated with an increase in cortical microRNA biogenesis"
12043:
10175:
10158:
8653:"Substrate requirements for let-7 function in the developing zebrafish embryo"
7808:
7492:
7469:
Tanzer A, Stadler PF (May 2004). "Molecular evolution of a microRNA cluster".
6623:
6194:
6169:
5596:
5579:
5447:
Park JE, Heo I, Tian Y, Simanshu DK, Chang H, Jee D, et al. (July 2011).
5408:"Substrate selectivity of exportin 5 and Dicer in the biogenesis of microRNAs"
5372:
3879:
3512:
2116:
1351:
with protein expression of MGMT and the direct target of miR-181d is the MGMT
242:
RNA was found to be conserved in many species, leading to the suggestion that
15756:
15603:
15593:
15474:
15429:
15407:
15243:
15172:
15100:
15095:
14253:
13754:
to enable up-regulation or suppression of endogenous mature microRNA function
13224:
13162:
12016:
11696:
10045:
Wen SY, Lin Y, Yu YQ, Cao SJ, Zhang R, Yang XM, et al. (February 2015).
9387:
Jiang Q, Wang Y, Hao Y, Juan L, Teng M, Zhang X, et al. (January 2009).
9183:"MicroRNA targeting specificity in mammals: determinants beyond seed pairing"
8618:
8342:
8324:
Maxwell EK, Ryan JF, Schnitzler CE, Browne WE, Baxevanis AD (December 2012).
7716:
6826:
Tan Y, Zhang B, Wu T, Skogerbø G, Zhu X, Guo X, et al. (February 2009).
4595:
4492:
3828:
1713:(DRD1), and its overexpression results in the upregulation of DRD1 and delta
1706:
1650:
1590:
1502:
1339:
1176:
1172:
1168:
1164:
1040:
969:
Interactions between microRNAs and complementary sequences on genes and even
866:
853:
678:
621:
539:
directly out of introns, bypassing the Microprocessor complex, are known as "
170:
80:
37:
13628:
12888:
12653:
12143:
Beveridge NJ, Gardiner E, Carroll AP, Tooney PA, Cairns MJ (December 2010).
11857:
11491:
11258:
11104:
10923:
10779:
10492:
10443:
10388:
9714:
9604:
9148:
8966:
8941:
8702:"Zebrafish miR-214 modulates Hedgehog signaling to specify muscle cell fate"
8292:
8015:
7998:
7867:
7365:
7097:
7067:
6844:
6794:
6737:
6496:
6479:
6445:
6086:
5424:
5407:
5359:(June 2004). "miRNAs on the move: miRNA biogenesis and the RNAi machinery".
5007:
Ali PS, Ghoshdastider U, Hoffmann J, Brutschy B, Filipek S (November 2012).
3411:
3296:
3074:
3023:
2967:
2901:
15772:
15686:
15506:
15479:
15424:
15343:
15300:
15110:
13760:
13636:
13601:
13547:
13496:
13445:
13377:
13328:
13283:
13181:
13130:
13068:
13033:
12992:
12965:
Nourse J, Braun J, Lackner K, Hüttelmaier S, Danckwardt S (November 2018).
12948:
12907:
12848:
12791:
12756:
12747:
12730:
12715:
12661:
12626:
12608:
12574:
12515:
12466:
12407:
12358:
12304:
12255:
12180:
12129:
12051:
12009:
11968:
11917:
11876:
11817:
11750:
11715:
11664:
11613:
11578:
Phua YL, Chu JY, Marrone AK, Bodnar AJ, Sims-Lucas S, Ho J (October 2015).
11553:
11510:
11456:
11366:
11331:
11266:
11215:
11172:
11112:
11093:
11074:
11034:
10991:
10942:
10883:
10842:
10798:
10735:
10694:
10643:
10594:
10545:
10510:
10461:
10407:
10348:
10307:
10258:
10184:
10128:
10072:
10031:
9974:
9933:
9883:
9821:
9772:
9733:
9674:
9623:
9564:
9515:
9458:
9422:
9373:
9314:
9265:
9216:
9167:
9116:
9081:
9032:
8975:
8926:
8858:
8835:
8784:
8751:"Sequence-specific inhibition of microRNA- and siRNA-induced RNA silencing"
8735:
8686:
8637:
8586:
8537:
8480:
8445:
8361:
8310:
8261:
8243:
8212:
8163:
8108:
8100:
8073:
8055:
8024:
7975:
7940:
7886:
7841:
Heimberg AM, Sempere LF, Moy VN, Donoghue PC, Peterson KJ (February 2008).
7816:
7773:
7735:
7684:
7627:
7618:
7601:
7579:
7535:
7500:
7455:
7437:
7384:
7325:
7276:
7235:
7183:
7105:
7086:
6988:
6953:
6912:
6863:
6812:
6755:
6698:
6641:
6592:
6557:
6505:
6464:
6413:
6395:
6351:
6308:
6254:
6213:
6154:
6121:"miRNPs: a novel class of ribonucleoproteins containing numerous microRNAs"
6105:
6046:
6027:
5992:
5943:
5894:
5853:
5812:
5803:
5786:
5771:
5719:
5684:
5635:
5564:
5523:
5482:
5433:
5380:
5335:
5292:
5257:
5205:
5156:
5099:
5034:
4993:
4944:
4903:
4862:
4813:
4764:
4721:
4672:
4613:
4559:
4510:
4453:
4412:
4356:
4290:
4247:
Baek D, Villén J, Shin C, Camargo FD, Gygi SP, Bartel DP (September 2008).
4225:
4174:
4128:
4071:
4028:
3993:
3944:
3898:
3847:
3796:
3747:
3698:
3616:
3565:
3530:
3481:
3430:
3371:
3314:
3253:
3210:
3169:
3128:
3082:
3031:
2975:
2919:
2853:
2799:
2707:
2658:
2604:
2572:
2570:
2568:
2558:
2506:
2457:
2408:
2341:
2266:
2223:
2188:
2134:
2085:
2039:
1762:
1754:
1367:
1308:
1218:
The first human disease known to be associated with miRNA deregulation was
1118:
1114:
645:
641:
536:
452:
448:
436:
141:
13684:
13264:
13111:
12697:
11949:
11122:. If this is an intentional citation to a retracted paper, please replace
10726:
10709:
10675:
10240:
9812:
9795:
8700:
Flynt AS, Li N, Thatcher EJ, Solnica-Krezel L, Patton JG (February 2007).
8472:
7571:
7202:"Are circulating microRNAs peripheral biomarkers for Alzheimer's disease?"
6539:
5449:"Dicer recognizes the 5' end of RNA for efficient and accurate processing"
4578:
Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH, et al. (October 2004).
3917:
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB (December 2003).
3657:
2753:
2640:
958:, a mechanism that has been termed "small RNA-induced gene activation" or
209:
mRNA. This complementarity was proposed to inhibit the translation of the
15598:
15444:
15311:
13710:
13477:
13427:
13310:
11908:
11891:
11645:
10707:
10625:
9665:
9404:
9296:
9247:
9014:
8908:
8668:
8519:
8427:
7922:
7249:
van den Berg MM, Krauskopf J, Ramaekers JG, et al. (February 2020).
5974:
5867:
Schwarz DS, Hutvágner G, Du T, Xu Z, Aronin N, Zamore PD (October 2003).
5239:
4323:"Identification of mammalian microRNA host genes and transcription units"
4110:
3778:
3462:
3385:
Harfe BD, McManus MT, Mansfield JH, Hornstein E, Tabin CJ (August 2005).
3052:
2439:
2390:
1758:
1070:
824:
670:
613:
555:
483:
174:
15399:
13583:
13528:
13392:
12389:
11772:
Luo M, Li L, Ding M, Niu Y, Xu X, Shi X, et al. (19 January 2023).
11595:
11545:
11026:
10753:, Tang R, Callis TE, Tatsuguchi M, Deng Z, et al. (February 2008).
9949:"Treating cancer with microRNA replacement therapy: A literature review"
9063:
8766:
8568:
8203:
8178:
8040:"MicroRNA gene evolution in Arabidopsis lyrata and Arabidopsis thaliana"
7119:
has been checked and does not affect the cited material, please replace
6894:
6680:
6343:
5910:"Asymmetry of intronic pre-miRNA structures in functional RISC assembly"
5464:
5284:
5138:
5081:
4919:"Microprocessor activity controls differential miRNA biogenesis in Vivo"
4876:
Han J, Lee Y, Yeom KH, Nam JW, Heo I, Rhee JK, et al. (June 2006).
4844:
4756:
4541:
4394:
4272:
4217:
4166:
3729:
3245:
3201:
3184:
2565:
2488:
2322:
2313:
1082:
appears to lack recognizable microRNAs, as well as the nuclear proteins
15658:
15367:
15362:
15357:
15264:
15238:
12706:
12591:
Li J, Li J, Liu X, Qin S, Guan Y, Liu Y, et al. (September 2013).
12194:
12161:
11438:
11056:
Xiao J, Luo X, Lin H, Zhang Y, Lu Y, Wang N, et al. (April 2007).
10063:
10046:
8862:. If this is an intentional citation to a such a paper, please replace
7017:
RNA and the Regulation of Gene Expression: A Hidden Layer of Complexity
6926:
Stark A, Brennecke J, Bushati N, Russell RB, Cohen SM (December 2005).
6275:"MicroRNA assassins: factors that regulate the disappearance of miRNAs"
6136:
6011:"The RNA-induced silencing complex: a versatile gene-silencing machine"
5762:
5745:
5648:
4338:
3812:"Naming 'junk': human non-protein coding RNA (ncRNA) gene nomenclature"
3760:
3671:
Giza DE, Calin GA (2015). "MicroRNA and Chronic Lymphocytic Leukemia".
3305:
1953:
1828:. In contrast when let-7 was inhibited by injections of let-7-specific
1697:
in alcohol pathophysiology. Downregulation of miR-382 was found in the
1642:
1630:
1486:
1315:
factor. Among 68 sporadic colon cancers with reduced expression of the
1298:
1282:
1242:
1238:
1153:
1137:
996:. Circulating miRNAs are released into body fluids including blood and
970:
732:
591:
480:
263:
76:
12983:
12966:
12939:
12922:
10013:
9996:
Liu G, Sun Y, Ji P, Li X, Cogdell D, Yang D, et al. (July 2014).
9965:
9948:
9764:
9655:
9638:
9328:
Artmann S, Jung K, Bleckmann A, Beissbarth T (2012). Provero P (ed.).
9230:
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (January 2008).
9108:
8130:
Kenny NJ, Namigai EK, Marlétaz F, Hui JH, Shimeld SM (December 2015).
6290:
6229:"Involvement of microRNA in AU-rich element-mediated mRNA instability"
4321:
Rodriguez A, Griffiths-Jones S, Ashurst JL, Bradley A (October 2004).
4320:
2290:"Mammalian microRNAs predominantly act to decrease target mRNA levels"
1202:
Deletion of the miR-17~92 cluster causes skeletal and growth defects.
600:
582:
504:
15522:
15295:
15269:
15151:
15141:
13707:, the experimentally validated microRNA-target interactions database.
13704:
10474:
9229:
7011:
6571:
Mazière P, Enright AJ (June 2007). "Prediction of microRNA targets".
3142:
Brennecke J, Hipfner DR, Stark A, Russell RB, Cohen SM (April 2003).
2845:
2791:
1829:
1746:
1742:
1718:
1690:
1685:
in alcohol abusers. miR-155, important in regulating alcohol-induced
1646:
1626:
1570:
1538:
1498:
1494:
1312:
1157:
1059:
1001:
857:
712:
649:
633:
567:
440:
424:
365:
257:
RNAs were found to be part of a large class of small RNAs present in
202:
145:
88:
11742:
9906:
Jones K, Nourse JP, Keane C, Bhatnagar A, Gandhi MK (January 2014).
9748:
8550:
7527:
6980:
5711:
5536:
2258:
2215:
1144:). The activity of an miRNA can be experimentally inhibited using a
520:(DGCR8 or "Pasha" in invertebrates), named for its association with
467:
79:. Found in plants, animals and some viruses, miRNAs are involved in
15676:
15527:
13735:
13461:"Advances in the techniques for the prediction of microRNA targets"
11207:
11164:
9546:
9450:
8717:
8650:
8599:
7967:
6662:
5496:
Ji X (2008). "The Mechanism of RNase III Action: How Dicer Dices".
4195:
4063:
4020:
2879:
1933:
1899:
1825:
1813:
1694:
1586:
1566:
1549:
MiRNAs are crucial for the healthy development and function of the
1518:
1445:
1328:
1304:
1290:
1097:
923:
835:
744:
648:
structure which is resistant to the Dicer mediated cleavage in the
540:
389:
284:
279:
The first human disease associated with deregulation of miRNAs was
218:
13409:
13342:
Qureshi A, Thakur N, Monga I, Thakur A, Kumar M (1 January 2014).
10202:(8th ed.). New York: W. H. Freeman and Company. p. 203.
8797:
5170:
Berezikov E, Chung WJ, Willis J, Cuppen E, Lai EC (October 2007).
4007:
Rajewsky N (June 2006). "microRNA target predictions in animals".
2823:
2415:
2099:
Qureshi A, Thakur N, Monga I, Thakur A, Kumar M (1 January 2014).
1794:
966:(saRNAs), but has also been demonstrated for endogenous microRNA.
856:(mRNAs). Animal miRNAs are usually complementary to a site in the
447:(Pol III) transcribes some miRNAs, especially those with upstream
15681:
15202:
15146:
14565:
14404:
14348:
14308:
14303:
14278:
14273:
14263:
14258:
14248:
14227:
14192:
14162:
14132:
14117:
14107:
14102:
14097:
14092:
14082:
14077:
14067:
14057:
14042:
14037:
14027:
9946:
9435:
9180:
7843:"MicroRNAs and the advent of vertebrate morphological complexity"
7150:"A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language?"
7148:
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP (August 2011).
7116:
5537:
Mirihana Arachchilage G, Dassanayake AC, Basu S (February 2015).
1943:
1833:
1809:
1782:
1766:
1674:
1449:
1371:
1223:
820:
587:
571:
60:
Examples of miRNA stem-loops, with the mature miRNAs shown in red
13049:
American Journal of Physiology. Heart and Circulatory Physiology
12271:"microRNAs: innovative targets for cerebral ischemia and stroke"
12025:"Heterogeneity and individuality: microRNAs in mental disorders"
11981:
10523:
10364:"Modulation of mismatch repair and genomic stability by miR-155"
9947:
Hosseinahli N, Aghapour M, Duijf PH, Baradaran B (August 2018).
9838:
Eyking A, Reis H, Frank M, Gerken G, Schmid KW, Cario E (2016).
9793:
9130:
Nielsen JA, Lau P, Maric D, Barker JL, Hudson LD (August 2009).
7289:
7041:
Place RF, Li LC, Pookot D, Noonan EJ, Dahiya R (February 2008).
6060:
MacRae IJ, Ma E, Zhou M, Robinson CV, Doudna JA (January 2008).
5006:
4692:
Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms
1820:
family. Let-7 accumulates in human tissues during the course of
15696:
15691:
15641:
15274:
15182:
15049:
14087:
13956:
13936:
13926:
13886:
13881:
13732:: MicroRNA targets database and data analysis and dataviz tool.
13722:
13512:"Predicting effective microRNA target sites in mammalian mRNAs"
12920:
12371:
11236:
10607:
7907:"Origins and evolution of microRNA genes in Drosophila species"
4790:. Methods in Molecular Biology. Vol. 342. pp. 33–47.
2522:
1948:
1801:). This paves the way for possible genetic obesity treatments.
1689:
responses, was found to be upregulated, suggesting the role of
1530:
1457:
and TIMP3 regulate TACE activity in turbulent flow conditions.
1406:
1342:, DNA repair is deficient due to epigenetic methylation of the
1129:
1083:
1066:
1048:
1033:
811:
751:
728:
559:
525:
476:
408:
397:
13615:
Baulcombe D (September 2002). "DNA events. An RNA microcosm".
13459:
Zheng H, Fu R, Wang JT, Liu Q, Chen H, Jiang SW (April 2013).
13144:
Curaba J, Spriggs A, Taylor J, Li Z, Helliwell C (July 2012).
12142:
10896:
9327:
8748:
8037:
7248:
6377:
6118:
4958:
Auyeung VC, Ulitsky I, McGeary SE, Bartel DP (February 2013).
4827:
Han J, Lee Y, Yeom KH, Kim YK, Jin H, Kim VN (December 2004).
4688:"MicroRNA biogenesis: there's more than one way to skin a cat"
3957:
3384:
2769:
1709:
that power motivational habits. miR-382 is the target for the
1625:
The vital role of miRNAs in gene expression is significant to
15499:
15156:
12964:
12479:
11984:"MicroRNAs in epilepsy: pathophysiology and clinical utility"
8699:
8651:
Kloosterman WP, Wienholds E, Ketting RF, Plasterk RH (2004).
5651:"Small RNA diversity in plants and its impact in development"
4785:
4428:"New human and mouse microRNA genes found by homology search"
3223:
1821:
1798:
1562:
1490:
1482:
1375:
1029:
979:
874:
747:
637:
487:
13713:, Web application to search for microRNAs in a plant genome.
13046:
12921:
Teruel-Montoya R, Rosendaal FR, Martínez C (February 2015).
11889:
10361:
10320:
9479:
9279:
Nam S, Li M, Choi K, Balch C, Kim S, Nephew KP (July 2009).
8893:"Miravirsen (SPC3649) can inhibit the biogenesis of miR-122"
7401:"MicroRNAs Play Key Role in Regulation of Circadian Rhythms"
6925:
5115:"Intronic microRNA precursors that bypass Drosha processing"
5058:"Intronic microRNA precursors that bypass Drosha processing"
4095:"Experimental strategies for microRNA target identification"
3958:
Ellwanger DC, Büttner FA, Mewes HW, Stümpflen V (May 2011).
3916:
3763:"miRBase: microRNA sequences, targets and gene nomenclature"
3141:
2421:
554:
As many as 16% of pre-miRNAs may be altered through nuclear
11193:
10955:
10566:
10099:"Scientists Develop a New, Powerful Cancer-Fighting Weapon"
8999:"Design of LNA probes that improve mismatch discrimination"
8890:
8749:
Meister G, Landthaler M, Dorsett Y, Tuschl T (March 2004).
8504:"Real-time quantification of microRNAs by stem-loop RT-PCR"
8409:
8323:
6167:
6062:"In vitro reconstitution of the human RISC-loading complex"
5221:
5009:"Recognition of the let-7g miRNA precursor by human Lin28B"
4957:
4144:
3543:
1726:
1714:
1678:
1526:
1432:
conditions therefore leads to the accumulation of miR-712.
1409:
microRNA-712 is a potential biomarker (i.e. predictor) for
1362:
1352:
1332:
1320:
798:
716:
636:, the pre-miRNA hairpin is cleaved by the RNase III enzyme
609:
495:
401:
267:
and human cells. The many RNAs of this class resembled the
100:
92:
50:
13699:
13146:"miRNA regulation in the early development of barley seed"
10272:
Jasperson KW, Tuohy TM, Neklason DW, Burt RW (June 2010).
10271:
9695:
9636:
9528:
7840:
7751:
6665:"Deadenylation is a widespread effect of miRNA regulation"
5784:
5169:
2623:
Friedman RC, Farh KK, Burge CB, Bartel DP (January 2009).
2372:
2060:"MicroRNAs: genomics, biogenesis, mechanism, and function"
1107:
360:
miRNA (a fruit fly commonly studied in genetic research).
15561:
15382:
15081:
12812:
12529:
Lippai D, Bala S, Csak T, Kurt-Jones EA, Szabo G (2013).
12325:"Up-regulation of microRNAs in brain of human alcoholics"
11287:
10425:
9905:
9577:
8412:"Target prediction for small, noncoding RNAs in bacteria"
8228:"Evolution and functional diversification of MIRNA genes"
8225:
7422:"Antiquity of microRNAs and their targets in land plants"
4829:"The Drosha-DGCR8 complex in primary microRNA processing"
3106:
2625:"Most mammalian mRNAs are conserved targets of microRNAs"
2470:
2288:
Guo H, Ingolia NT, Weissman JS, Bartel DP (August 2010).
1522:
1510:
1117:. This makes it necessary to cool samples on ice and use
757:
169:
The first miRNA was discovered in 1993 by a group led by
13341:
12682:
12528:
12269:
Ouyang YB, Stary CM, Yang GY, Giffard R (January 2013).
10656:
8996:
8274:
7338:
6426:
5825:
5613:
2671:
2525:"Identification of tissue-specific microRNAs from mouse"
2287:
2155:"MicroRNAs: target recognition and regulatory functions"
2098:
45:
13565:
13397:
Resource for experimental viral miRNA and their targets
11729:
Schratt G (December 2009). "microRNAs at the synapse".
10748:
10197:
9045:
8226:
Cuperus JT, Fahlgren N, Carrington JC (February 2011).
8129:
6879:"RNA and transcriptional modulation of gene expression"
6521:
6170:"Identification of novel argonaute-associated proteins"
5270:
5224:"Frequency and fate of microRNA editing in human brain"
3711:
2622:
1179:- and miRNA-expression information have been proposed.
13194:
12769:
12268:
12022:
11830:
10855:
9746:
8176:
7996:
7651:"Origins and evolution of eukaryotic RNA interference"
7599:
7147:
5869:"Asymmetry in the assembly of the RNAi enzyme complex"
4917:
Conrad T, Marsico A, Gehre M, Orom UA (October 2014).
4916:
2375:"An estimate of the total number of true human miRNAs"
1513:
pathway with increase in p53 effector genes including
103:
molecules by one or more of the following processes:
13510:
Agarwal V, Bell GW, Nam JW, Bartel DP (August 2015).
13143:
12639:
12322:
10814:
9837:
9129:
8997:
You Y, Moreira BG, Behlke MA, Owczarzy R (May 2006).
8501:
7953:
4580:"MicroRNA genes are transcribed by RNA polymerase II"
4246:
4093:
Thomson DW, Bracken CP, Goodall GJ (September 2011).
4092:
3860:
3578:
2945:
1557:(such as dendritogenesis or spine morphogenesis) and
1501:
loss and glomerular aneurysms. High throughput whole
900:
sites, which affects the expression of target genes.
656:(RISC) where the miRNA and its mRNA target interact.
13746:
An animated video of the microRNA biogenesis process
13509:
12222:"MicroRNA in ischemic stroke etiology and pathology"
11577:
9696:
Kyriakidis I, Kyriakidis K, Tsezou A (August 2022).
9094:
8458:
7794:
6768:
6226:
6059:
5866:
5826:
Khvorova A, Reynolds A, Jayasena SD (October 2003).
1999:
1997:
1995:
1993:
1895:
1124:
microRNA expression can be quantified in a two-step
427:
with a specially modified nucleotide at the 5' end,
12815:"The Lin28/let-7 axis regulates glucose metabolism"
12420:
11524:Libby P (2002). "Inflammation in atherosclerosis".
11004:
10225:"MicroRNAs: new players in the DNA damage response"
7040:
6321:
5446:
5412:
Cold Spring Harbor Symposia on Quantitative Biology
1585:) as well as many psychiatric disorders (including
750:, and the RNA recognition motif containing protein
586:The human exportin-5 protein (red) in complex with
13716:
12023:Hommers LG, Domschke K, Deckert J (January 2015).
11931:Roy B, Lee E, Li T, Rampersaud M (February 2022).
11930:
11472:
8939:
6825:
6711:
6427:Wang XJ, Reyes JL, Chua NH, Gaasterland T (2004).
5828:"Functional siRNAs and miRNAs exhibit strand bias"
5609:
5607:
3443:
3274:
1569:neurons), and neurodegeneration (when silenced in
13656:
13296:
12763:
12081:
11412:
10093:
9046:Lagendijk AK, Moulton JD, Bakkers J (June 2012).
7997:Warthmann N, Das S, Lanz C, Weigel D (May 2008).
7697:
7600:Peterson KJ, Dietrich MR, McPeek MA (July 2009).
4626:
4523:
4376:
4316:
4314:
3497:"Therapeutic microRNA strategies in human cancer"
3495:Li C, Feng Y, Coukos G, Zhang L (December 2009).
3494:
3335:
2725:
2424:"MirGeneDB 2.0: the metazoan microRNA complement"
1990:
1307:mutations in DNA repair genes cause only 2–5% of
590:(yellow) and a pre-microRNA (green), showing two-
15754:
13005:
12960:
12958:
9940:
9833:
9831:
9386:
8940:Choi WY, Giraldez AJ, Schier AF (October 2007).
8086:
7836:
7834:
7595:
7593:
7591:
7589:
6380:"Kinetic signatures of microRNA modes of action"
6373:
6371:
6369:
3960:"The sufficient minimal set of miRNA seed types"
3629:
3444:Trang P, Weidhaas JB, Slack FJ (December 2008).
2576:
1869:DNA is believed to be regulated by viral miRNA.
1661:. Altered miRNA levels were found in the medial
1293:from either DNA damage or uncorrected errors in
684:
345:from genes located in different genome regions.
13657:Lee RC, Feinbaum RL, Ambros V (December 1993).
11055:
7747:
7745:
7648:
7556:
5956:
5739:
5737:
5604:
5401:
5399:
5397:
5351:
5347:
5345:
5217:
5215:
4377:Cai X, Hagedorn CH, Cullen BR (December 2004).
4140:
4138:
4049:
2997:
2995:
2993:
2941:
2939:
2937:
2875:
2873:
2871:
2819:
2817:
2765:
2763:
2726:Lee RC, Feinbaum RL, Ambros V (December 1993).
2618:
2616:
2614:
2518:
2516:
2244:
2201:
1741:miRNAs play crucial roles in the regulation of
234:to promote a later developmental transition in
224:In 2000, a second small RNA was characterized:
213:mRNA into the LIN-14 protein. At the time, the
183:gene, which was known to control the timing of
13458:
13092:
12728:
12329:Alcoholism: Clinical and Experimental Research
11573:
11571:
10159:"Significance of multiple mutations in cancer"
9278:
7904:
7900:
7898:
7896:
6712:Bazzini AA, Lee MT, Giraldez AJ (April 2012).
6570:
6272:
5957:Okamura K, Chung WJ, Lai EC (September 2008).
4627:Zhou X, Ruan J, Wang G, Zhang W (March 2007).
4311:
3182:
2721:
2719:
2717:
2577:Lewis BP, Burge CB, Bartel DP (January 2005).
2148:
2146:
2144:
2053:
2051:
2049:
110:Destabilization of the mRNA by shortening its
15327:
15065:
13776:
13452:
13403:
12999:
12955:
12914:
12855:
12806:
12722:
12676:
12633:
12136:
12075:
11722:
11408:
11406:
11404:
11402:
11400:
11398:
11396:
11230:
11187:
11150:
11144:
11049:
10949:
10890:
10849:
10810:
10808:
10742:
9890:
9828:
9787:
9740:
9689:
9571:
9522:
9429:
9321:
9272:
9223:
9174:
9123:
9088:
8990:
8933:
8884:
8544:
8495:
8452:
8403:
8268:
8170:
8080:
7831:
7642:
7586:
7550:
7468:
7462:
7141:
6876:
6870:
6819:
6769:Djuranovic S, Nahvi A, Green R (April 2012).
6477:
6366:
5743:
5112:
5055:
4875:
4826:
4734:
4577:
4303:: CS1 maint: DOI inactive as of April 2024 (
3809:
3102:
3100:
2092:
1464:
919:Co-translational nascent protein degradation;
873:in zebrafish, as well as on bantam-miRNA and
396:As many as 40% of miRNA genes may lie in the
13790:
11771:
11680:"Dynamic Roles of microRNAs in Neurogenesis"
11394:
11392:
11390:
11388:
11386:
11384:
11382:
11380:
11378:
11376:
10962:Journal of Molecular and Cellular Cardiology
8381:Genetic Engineering & Biotechnology News
8317:
7742:
7513:
7419:
6762:
6705:
6517:
6515:
6268:
6266:
6264:
6112:
6053:
5950:
5907:
5901:
5860:
5819:
5778:
5734:
5642:
5489:
5405:
5394:
5342:
5307:
5264:
5212:
5163:
4779:
4679:
4517:
4468:
4419:
4135:
3754:
3705:
3630:Wightman B, Ha I, Ruvkun G (December 1993).
3537:
3488:
3446:"MicroRNAs as potential cancer therapeutics"
3437:
3378:
3336:Wilfred BR, Wang WX, Nelson PT (July 2007).
3329:
3268:
3217:
3176:
3135:
2990:
2934:
2868:
2814:
2760:
2611:
2513:
2464:
1281:Many miRNAs can directly target and inhibit
1182:
594:overhang recognition element (orange). From
107:Cleavage of the mRNA strand into two pieces,
13738:: An open access database for decoding the
13465:International Journal of Molecular Sciences
11633:International Journal of Molecular Sciences
11568:
10044:
8368:
7893:
6008:
4372:
4370:
4368:
4366:
4249:"The impact of microRNAs on protein output"
3623:
3579:Hydbring P, Badalian-Very G (August 2013).
2714:
2141:
2046:
1167:) that comes with methodological problems.
15341:
15334:
15320:
15072:
15058:
13783:
13769:
13568:"A uniform system for microRNA annotation"
13245:
12861:
12590:
12318:
12316:
12314:
12219:
12199:Centers for Disease Control and Prevention
11136:|...|intentional=yes}}
10805:
9995:
8876:|...|intentional=yes}}
7199:
6315:
4573:
4571:
4569:
3919:"Prediction of mammalian microRNA targets"
3714:"A uniform system for microRNA annotation"
3097:
2665:
1797:and CCAAT/enhancer-binding protein alpha (
1765:line hMSC-Tert20. Decreased expression of
1757:, were examined in the immortalized human
1000:and have the potential to be available as
916:Ribosome drop-off (premature termination);
490:molecules (green). Drosha consists of two
27:Small non-coding ribonucleic acid molecule
13674:
13614:
13591:
13537:
13527:
13486:
13476:
13435:
13367:
13318:
13273:
13263:
13171:
13161:
13120:
13110:
13023:
12982:
12938:
12897:
12887:
12838:
12746:
12705:
12616:
12564:
12554:
12505:
12473:
12456:
12446:
12397:
12365:
12348:
12294:
12245:
12170:
12160:
12119:
12109:
12099:
12059:
11999:
11958:
11948:
11907:
11866:
11856:
11807:
11797:
11705:
11695:
11654:
11644:
11626:
11603:
11500:
11490:
11446:
11373:
11321:
11083:
11073:
10981:
10932:
10922:
10873:
10832:
10788:
10778:
10725:
10684:
10674:
10633:
10584:
10500:
10451:
10421:
10419:
10417:
10397:
10387:
10338:
10297:
10248:
10174:
10118:
10062:
10038:
10021:
9989:
9964:
9923:
9873:
9863:
9811:
9723:
9713:
9664:
9654:
9613:
9603:
9554:
9505:
9412:
9363:
9353:
9304:
9255:
9206:
9157:
9147:
9071:
9022:
8965:
8916:
8825:
8815:
8774:
8725:
8676:
8627:
8617:
8576:
8527:
8435:
8351:
8341:
8300:
8251:
8202:
8063:
8014:
7930:
7876:
7866:
7725:
7715:
7674:
7617:
7482:
7445:
7374:
7364:
7315:
7266:
7225:
7195:
7193:
7173:
7076:
7066:
6943:
6902:
6853:
6843:
6802:
6745:
6688:
6631:
6547:
6512:
6495:
6454:
6444:
6403:
6298:
6279:Nature Structural & Molecular Biology
6261:
6244:
6203:
6193:
6144:
6095:
6085:
6036:
6026:
5982:
5933:
5884:
5843:
5802:
5761:
5674:
5595:
5554:
5472:
5423:
5247:
5195:
5146:
5089:
5024:
4983:
4934:
4893:
4852:
4711:
4685:
4662:
4652:
4620:
4603:
4549:
4500:
4443:
4402:
4346:
4280:
4118:
3983:
3934:
3888:
3878:
3837:
3827:
3786:
3737:
3670:
3647:
3606:
3596:
3520:
3471:
3461:
3420:
3410:
3361:
3304:
3200:
3159:
3001:
2909:
2743:
2697:
2648:
2594:
2548:
2496:
2473:"The microRNAs of Caenorhabditis elegans"
2447:
2398:
2331:
2321:
2178:
2124:
2075:
2029:
1323:, most were found to be deficient due to
627:
13006:Nourse J, Danckwardt S (February 2021).
12729:Zuo Y, Qiang L, Farmer SR (March 2006).
12421:Gorini G, Nunez YO, Mayfield RD (2013).
10714:Journal of the National Cancer Institute
10156:
7649:Shabalina SA, Koonin EV (October 2008).
6966:
6608:"Functional aspects of animal microRNAs"
6605:
6004:
6002:
5908:Lin SL, Chang D, Ying SY (August 2005).
5113:Ruby JG, Jan CH, Bartel DP (July 2007).
5056:Ruby JG, Jan CH, Bartel DP (July 2007).
4363:
4006:
3183:Cuellar TL, McManus MT (December 2005).
3121:10.1146/annurev.arplant.57.032905.105218
1276:
1209:
951:gene regulations and mir-92 and mir-19.
834:
702:
581:
466:
388:
55:
44:
12522:
12414:
12311:
11728:
11677:
10222:
7698:Axtell MJ, Westholm JO, Lai EC (2011).
6273:Kai ZS, Pasquinelli AE (January 2010).
4566:
4524:Baskerville S, Bartel DP (March 2005).
1108:Experimental detection and manipulation
1096:, the first known member of the phylum
1076:Focusing on the animals, the genome of
1069:of most eukaryotic organisms, from the
663:
528:, a protein that cuts RNA, to form the
419:miRNA genes are usually transcribed by
14:
15755:
13093:Borchers A, Pieler T (November 2010).
13088:
13086:
12772:Trends in Endocrinology and Metabolism
12586:
12584:
11468:
11466:
11344:
10414:
10274:"Hereditary and familial colon cancer"
9232:"miRBase: tools for microRNA genomics"
8374:
7905:Nozawa M, Miura S, Nei M (July 2010).
7190:
7133:|...|checked=yes}}
4474:
3810:Wright MW, Bruford EA (January 2011).
2152:
2057:
2003:
1804:Another class of miRNAs that regulate
1705:significant in regulating feelings of
910:60S Ribosomal unit joining inhibition;
823:, a liver-enriched miRNA important in
758:Mode of silencing and regulatory loops
15315:
15053:
13764:
13564:miRNA definition and classification:
13399:. Bioinformatics center, CSIR-IMTECH.
12971:Journal of Thrombosis and Haemostasis
12927:Journal of Thrombosis and Haemostasis
11523:
7003:
5999:
5700:Nature Reviews Molecular Cell Biology
5577:
5313:
4425:
1929:List of miRNA target prediction tools
1191:
885:, independent of mRNA deadenylation.
830:
462:
189:larval development by repressing the
13655:, the first miRNA to be discovered:
12862:Frost RJ, Olson EN (December 2011).
7200:Kumar S, Reddy PH (September 2016).
7012:"Small RNA-Mediated Gene Activation"
6877:Hawkins PG, Morris KV (March 2008).
6612:Cellular and Molecular Life Sciences
5697:
5691:
3581:"Clinical applications of microRNAs"
1974:Small nucleolar RNA-derived microRNA
1872:
1440:MiR-712 targets tissue inhibitor of
741:fragile X mental retardation protein
352:) miRNA and oar-miR-124 is a sheep (
13083:
12735:The Journal of Biological Chemistry
12581:
12213:
11479:The Journal of Biological Chemistry
11463:
11062:The Journal of Biological Chemistry
7242:
6015:The Journal of Biological Chemistry
5791:The Journal of Biological Chemistry
5616:International Journal of Cardiology
5000:
2690:10.1146/annurev-genet-120213-092023
1924:List of miRNA gene prediction tools
727:Additional RISC components include
719:domain that structurally resembles
524:. DGCR8 associates with the enzyme
518:DiGeorge Syndrome Critical Region 8
193:gene. When Lee et al. isolated the
24:
13557:
12238:10.1152/physiolgenomics.00158.2010
11627:Cao DD, Li L, Chan WY (May 2016).
9486:American Journal of Human Genetics
7420:Axtell MJ, Bartel DP (June 2005).
7009:
5746:"Why do miRNAs live in the miRNP?"
5744:Schwarz DS, Zamore PD (May 2002).
5495:
4477:"Processing of intronic microRNAs"
1535:Fluorescent-activated cell sorting
1481:Targeted deletion of Dicer in the
457:mammalian wide interspersed repeat
25:
15789:
15458:Micro
13693:
13390:
12642:Experimental Biology and Medicine
12071:from the original on 23 May 2022.
10875:10.1161/CIRCULATIONAHA.107.687947
10229:Journal of Molecular Cell Biology
9800:International Journal of Oncology
7655:Trends in Ecology & Evolution
6009:Pratt AJ, MacRae IJ (July 2009).
5584:Achievements in the Life Sciences
4686:Faller M, Guo F (November 2008).
3342:Molecular Genetics and Metabolism
3185:"MicroRNAs and endocrine biology"
1749:. Studies to determine what role
1745:progenitors differentiating into
1544:
1476:
731:, PACT (protein activator of the
577:
15737:
15736:
15178:Cis-natural antisense transcript
15079:
13503:
13384:
13335:
13290:
13239:
13188:
13137:
13040:
13025:10.1016/j.pharmthera.2020.107676
12341:10.1111/j.1530-0277.2011.01544.x
12262:
12187:
11975:
11924:
11883:
11824:
11765:
11671:
11620:
11517:
11347:The American Journal of Medicine
11338:
11281:
10998:
10701:
10650:
10601:
10560:
10517:
10468:
10355:
10314:
10265:
10216:
10191:
10150:
10087:
9899:
9630:
9473:
9380:
9039:
8791:
8742:
8693:
8644:
8593:
8219:
8123:
8031:
7990:
7947:
7788:
7766:10.1111/j.1525-142X.2008.00302.x
7691:
7507:
7413:
7332:
7283:
7034:
6960:
6919:
5386:
4475:Kim YK, Kim VN (February 2007).
4445:10.1111/j.1432-1033.2004.04389.x
3558:10.1016/j.pharmthera.2009.10.003
1959:Mir-M7 microRNA precursor family
1898:
1384:
1113:because of ubiquitously present
612:. This protein, a member of the
414:
15413:precursor, heterogenous nuclear
15106:Signal recognition particle RNA
13246:Waheed S, Zeng L (March 2020).
13012:Pharmacology & Therapeutics
10157:Loeb KR, Loeb LA (March 2000).
9753:Genes, Chromosomes & Cancer
9643:International Journal of Cancer
8377:"miRNAs' Therapeutic Potential"
8003:Molecular Biology and Evolution
7268:10.1016/j.pneurobio.2019.101732
6656:
6599:
6564:
6471:
6420:
6161:
5571:
5530:
5440:
5361:Current Opinion in Cell Biology
5106:
5049:
4951:
4910:
4869:
4820:
4728:
4240:
4189:
4086:
4043:
4000:
3854:
3803:
3664:
3598:10.12688/f1000research.2-136.v2
3572:
3546:Pharmacology & Therapeutics
3046:
1509:(Bim) and dysregulation of the
1357:three prime untranslated region
888:miRNAs occasionally also cause
566:acting on RNA (ADARs) catalyze
543:." Mirtrons have been found in
289:
130:small interfering RNAs (siRNAs)
15543:Trans-acting small interfering
15507:Enhancer RNAs
15425:Transfer
13299:Journal of Experimental Botany
12498:10.1523/JNEUROSCI.0445-14.2014
12032:Journal of Neural Transmission
9953:Journal of Cellular Physiology
6484:Nucleic Acids Symposium Series
6205:11858/00-001M-0000-0012-E763-B
5556:10.1016/j.chembiol.2014.12.013
3109:Annual Review of Plant Biology
2911:11858/00-001M-0000-0012-F65F-2
2348:
2281:
2238:
2195:
1214:Role of miRNA in a cancer cell
907:Cap-40S initiation inhibition;
217:small RNA was thought to be a
91:to complementary sequences in
75:molecules containing 21 to 23
71:) are small, single-stranded,
13:
1:
15430:Ribosomal
15408:Messenger
12220:Rink C, Khanna S (May 2011).
12001:10.1016/S1474-4422(16)30246-0
10586:10.1158/0008-5472.CAN-04-0044
10538:10.1016/j.febslet.2004.08.013
9925:10.1158/1078-0432.CCR-13-1024
9291:(Web Server issue): W356–62.
8841:(This paper currently has an
6606:Williams AE (February 2008).
5886:10.1016/S0092-8674(03)00759-1
5845:10.1016/S0092-8674(03)00801-8
5026:10.1016/j.febslet.2012.09.034
3976:10.1093/bioinformatics/btr149
3936:10.1016/S0092-8674(03)01018-3
3161:10.1016/S0092-8674(03)00231-9
2550:10.1016/S0960-9822(02)00809-6
2077:10.1016/S0092-8674(04)00045-5
1984:
1839:
1620:
929:mRNA decay (destabilisation);
691:RNA-induced silencing complex
685:RNA-induced silencing complex
654:RNA-induced silencing complex
384:
85:regulation of gene expression
13676:10.1016/0092-8674(93)90529-Y
12556:10.1371/journal.pone.0070945
12448:10.1371/journal.pone.0082565
12378:The Pharmacogenomics Journal
12111:10.1371/journal.pone.0006121
11799:10.1371/journal.pone.0279029
11731:Nature Reviews. Neuroscience
11359:10.1016/j.amjmed.2008.10.013
10340:10.1053/j.gastro.2005.01.056
10290:10.1053/j.gastro.2010.01.054
10223:Hu H, Gatti RA (June 2011).
9865:10.1371/journal.pone.0156871
9355:10.1371/journal.pone.0038365
9199:10.1016/j.molcel.2007.06.017
8851:10.1371/journal.pbio.3001631
8817:10.1371/journal.pbio.0050203
8156:10.1016/j.margen.2015.07.004
7911:Genome Biology and Evolution
7471:Journal of Molecular Biology
7218:10.1016/j.bbadis.2016.06.001
7020:. Horizon Scientific Press.
6585:10.1016/j.drudis.2007.04.002
6478:Kawasaki H, Taira K (2004).
5628:10.1016/j.ijcard.2014.01.082
5406:Lund E, Dahlberg JE (2006).
5328:10.1016/j.biochi.2007.06.002
5188:10.1016/j.molcel.2007.09.028
4936:10.1016/j.celrep.2014.09.007
4704:10.1016/j.bbagrm.2008.08.005
4654:10.1371/journal.pcbi.0030037
3649:10.1016/0092-8674(93)90530-4
3189:The Journal of Endocrinology
2745:10.1016/0092-8674(93)90529-Y
1435:
1401:
1263:positron emission tomography
1220:chronic lymphocytic leukemia
1019:
1012:seems to play a key role in
842:Internal Ribosome Entry Site
794:is mediated by the 5'-to-3'
675:RNA methyltransferaseprotein
479:protein in complex with the
281:chronic lymphocytic leukemia
7:
13061:10.1152/ajpheart.00338.2012
12486:The Journal of Neuroscience
10974:10.1016/j.yjmcc.2007.04.004
9399:(Database issue): D98–104.
8375:Dimond PF (15 March 2010).
7754:Evolution & Development
7308:10.1016/j.ebiom.2015.09.003
5506:10.1007/978-3-540-75157-1_5
3681:10.1007/978-3-319-23730-5_2
3354:10.1016/j.ymgme.2007.03.011
1914:Anti-miRNA oligonucleotides
1891:
1427:internal transcribed spacer
779:
10:
15794:
15609:Multicopy single-stranded
15453:Interferential
15265:Reverse transcribing virus
13217:10.1007/s11756-021-00720-1
12831:10.1016/j.cell.2011.08.033
12686:Stem Cells and Development
12287:10.2174/138945013804806424
11314:10.1038/s41598-017-04040-w
10834:10.1016/j.cell.2007.03.030
10120:10.1016/j.cell.2022.04.030
9498:10.1016/j.ajhg.2011.09.014
9242:(Database issue): D154–8.
7667:10.1016/j.tree.2008.06.005
7166:10.1016/j.cell.2011.07.014
6945:10.1016/j.cell.2005.11.023
6246:10.1016/j.cell.2004.12.038
5926:10.1016/j.gene.2005.04.036
5667:10.2174/138920210790217918
4976:10.1016/j.cell.2013.01.031
4895:10.1016/j.cell.2006.03.043
4633:PLOS Computational Biology
4275:(inactive 26 April 2024).
3773:(Database issue): D140–4.
2596:10.1016/j.cell.2004.12.035
2171:10.1016/j.cell.2009.01.002
2153:Bartel DP (January 2009).
2058:Bartel DP (January 2004).
2022:10.1016/j.cell.2018.03.006
1876:
1857:
1736:
1655:nervous system development
1465:Human homolog microRNA-205
788:Decay of mature miRNAs in
688:
371:
155:
121:of the mRNA into proteins.
31:
15732:
15667:
15617:
15560:
15523:Guide
15515:
15443:
15398:
15381:
15350:
15283:
15252:
15231:
15165:
15124:
15088:
14996:
14790:
14659:
14523:
14362:
14236:
14010:
13799:
13752:miRNA modulation reagents
12923:"MicroRNAs in hemostasis"
12784:10.1016/j.tem.2015.12.006
12044:10.1007/s00702-014-1338-4
11684:Frontiers in Neuroscience
7809:10.1134/S0006297913060084
7493:10.1016/j.jmb.2004.03.065
6624:10.1007/s00018-007-7355-9
6195:10.1016/j.cub.2005.10.048
5597:10.1016/j.als.2016.11.007
5373:10.1016/j.ceb.2004.04.003
5172:"Mammalian mirtron genes"
4426:Weber MJ (January 2005).
3880:10.1186/s12864-019-5947-z
3513:10.1208/s12248-009-9145-9
2678:Annual Review of Genetics
1848:
1608:
1420:
1415:matrix metalloproteinases
1205:
1183:Human and animal diseases
1126:polymerase chain reaction
1065:microRNAs feature in the
989:miRNAs are also found as
852:to a part of one or more
83:and post-transcriptional
15485:Small nuclear
13163:10.1186/1471-2229-12-120
11896:Human Molecular Genetics
11697:10.3389/fnins.2012.00071
10002:The Journal of Pathology
9912:Clinical Cancer Research
8619:10.1186/1471-2164-11-716
8343:10.1186/1471-2164-13-714
7797:Biochemistry. Biokhimiia
7717:10.1186/gb-2011-12-4-221
6969:Nature Reviews. Genetics
4796:10.1385/1-59745-123-1:33
4596:10.1038/sj.emboj.7600385
4493:10.1038/sj.emboj.7601512
3829:10.1186/1479-7364-5-2-90
2247:Nature Reviews. Genetics
2204:Nature Reviews. Genetics
2004:Bartel DP (March 2018).
954:dsRNA can also activate
32:Not to be confused with
15599:Genomic
15232:Cis-regulatory elements
15203:Repeat-associated siRNA
13629:10.1126/science.1077906
13360:10.1093/database/bau103
12889:10.1073/pnas.1118922109
12654:10.1258/ebm.2011.011101
12597:EMBO Molecular Medicine
11858:10.1073/pnas.0801689105
11678:Lang MF, Shi Y (2012).
11492:10.1074/jbc.M112.425652
11259:10.1126/science.1139089
11105:10.1074/jbc.A111.700015
10924:10.1073/pnas.0608791103
10780:10.1073/pnas.0710228105
10389:10.1073/pnas.1002472107
10176:10.1093/carcin/21.3.379
9715:10.3390/cancers14163976
9605:10.1073/pnas.0404432101
9149:10.1186/1471-2202-10-98
8967:10.1126/science.1147535
8293:10.1126/science.1242592
7868:10.1073/pnas.0712259105
7516:Nature Reviews Genetics
7366:10.1073/pnas.2020454118
7098:10.1073/pnas.1803343115
7068:10.1073/pnas.0707594105
6845:10.1186/1471-2199-10-12
6795:10.1126/science.1215691
6738:10.1126/science.1215704
6446:10.1186/gb-2004-5-9-r65
6125:Genes & Development
6087:10.1073/pnas.0710869105
5750:Genes & Development
5543:Chemistry & Biology
5425:10.1101/sqb.2006.71.050
4833:Genes & Development
3412:10.1073/pnas.0504834102
3297:10.1126/science.1091903
3075:10.1126/science.1114519
3024:10.1126/science.1065329
2968:10.1126/science.1065062
2902:10.1126/science.1064921
2477:Genes & Development
2117:10.1093/database/bau103
1267:computerized tomography
459:(MWIR) promoter units.
339:aacauucaUUgcugucggugGgu
330:aacauucaACgcugucggugAgu
134:RNA interference (RNAi)
15702:Artificial chromosomes
15490:Small nucleolar
15116:Transfer-messenger RNA
12748:10.1074/jbc.M510682200
12609:10.1002/emmm.201201900
12226:Physiological Genomics
11075:10.1074/jbc.C700015200
10614:Nucleic Acids Research
10200:Molecular Cell Biology
9393:Nucleic Acids Research
9285:Nucleic Acids Research
9236:Nucleic Acids Research
9003:Nucleic Acids Research
8897:Nucleic Acids Research
8657:Nucleic Acids Research
8508:Nucleic Acids Research
8416:Nucleic Acids Research
8387:(6): 1. Archived from
8244:10.1105/tpc.110.082784
8101:10.1002/bies.201000005
8056:10.1105/tpc.110.073999
7619:10.1002/bies.200900033
7438:10.1105/tpc.105.032185
7014:. In Morris KV (ed.).
6396:10.1261/rna.032284.112
6028:10.1074/jbc.R900012200
5804:10.1074/jbc.M404931200
5228:Nucleic Acids Research
4099:Nucleic Acids Research
3767:Nucleic Acids Research
2428:Nucleic Acids Research
2379:Nucleic Acids Research
1751:pluripotent stem cells
1325:epigenetic methylation
1215:
913:Elongation inhibition;
883:translation initiation
845:
791:Caenorhabditis elegans
708:
628:Cytoplasmic processing
618:guanosine triphosphate
605:
530:Microprocessor complex
509:
435:(a poly(A) tail), and
393:
61:
53:
15495:Small Cajal Body RNAs
15208:Small interfering RNA
13740:transcription factors
13613:review of small RNA:
13320:10536/DRO/DU:30106047
13265:10.3390/genes11030319
13112:10.3390/ijms232314755
12698:10.1089/scd.2010.0503
11988:The Lancet. Neurology
11950:10.3390/genes13030425
11584:Physiological Reports
11419:Nature Communications
11128:|...}}
11099:(Retracted, see
10676:10.1186/2041-9414-3-3
10493:10.1093/neuonc/nop003
10444:10.1093/neuonc/nos089
10097:(13 September 2022).
9813:10.3892/ijo.2011.1043
8874:expression of concern
8868:|...}}
8866:expression of concern
8843:expression of concern
8473:10.1038/nprot.2008.14
8016:10.1093/molbev/msn029
7572:10.1089/dna.2006.0545
7125:|...}}
6832:BMC Molecular Biology
6540:10.1101/gr.080127.108
6497:10.1093/nass/48.1.211
2641:10.1101/gr.082701.108
2362:Manchester University
2358:miRNAs in the miRBase
1969:Small interfering RNA
1836:and type 2 diabetes.
1701:, a structure in the
1493:cells, smooth muscle
1277:DNA repair and cancer
1245:and poor survival in
1213:
1039:microRNAs are useful
964:small activating RNAs
838:
706:
585:
470:
392:
230:RNA, which represses
59:
48:
15548:Subgenomic messenger
15463:Small interfering
15435:Transfer-messenger
15198:Piwi-interacting RNA
13728:4 March 2014 at the
13700:The miRBase database
13478:10.3390/ijms14048179
13428:10.1128/JVI.05911-11
12275:Current Drug Targets
12149:Molecular Psychiatry
11646:10.3390/ijms17060842
7560:DNA and Cell Biology
7206:Biochim Biophys Acta
6573:Drug Discovery Today
5975:10.4161/cc.7.18.6734
3463:10.1038/onc.2009.353
2006:"Metazoan MicroRNAs"
1711:dopamine receptor D1
1583:Huntington's disease
1555:synaptic development
1442:metalloproteinases 3
1349:inversely correlated
1128:process of modified
1093:Trichoplax adhaerens
1055:Arabidopsis thaliana
1049:introns or intergene
890:histone modification
807:Arabidopsis thaliana
739:), the SMN complex,
664:Biogenesis in plants
564:adenosine deaminases
535:Pre-miRNAs that are
498:ion (spheres). From
128:miRNAs resemble the
15137:Small nucleolar RNA
13584:10.1261/rna.2183803
13529:10.7554/eLife.05005
13416:Journal of Virology
13209:2021Biolg..76.1551C
12880:2011PNAS..10821075F
12547:2013PLoSO...870945L
12439:2013PLoSO...882565G
12390:10.1038/tpj.2012.17
12101:2009PLoSO...4.6121F
11849:2008PNAS..105.5614C
11790:2023PLoSO..1879029L
11596:10.14814/phy2.12537
11546:10.1038/nature01323
11538:2002Natur.420..868L
11431:2013NatCo...4.3000S
11353:(1 Suppl): S3–S14.
11306:2017NatSR...7.4511K
11251:2007Sci...316..575V
11027:10.1038/nature03817
11019:2005Natur.436..214Z
10915:2006PNAS..10318255V
10771:2008PNAS..105.2111C
10727:10.1093/jnci/dju318
10380:2010PNAS..107.6982V
10241:10.1093/jmcb/mjq042
9856:2016PLoSO..1156871E
9596:2004PNAS..10111755C
9590:(32): 11755–11760.
9346:2012PLoSO...738365A
9064:10.1242/bio.2012810
8958:2007Sci...318..271C
8767:10.1261/rna.5235104
8569:10.1261/rna.2610405
8204:10.1038/nature09016
8195:2010Natur.465..617C
8148:2015MarGn..24..139K
7859:2008PNAS..105.2946H
7357:2021PNAS..11820454Z
7059:2008PNAS..105.1608P
6895:10.4161/cc.7.5.5522
6787:2012Sci...336..237D
6730:2012Sci...336..233B
6681:10.1261/rna.1399509
6344:10.1038/nature08349
6336:2009Natur.461..546C
6186:2005CBio...15.2149M
6078:2008PNAS..105..512M
5465:10.1038/nature10198
5285:10.1038/ncb0309-228
5273:Nature Cell Biology
5139:10.1038/nature05983
5131:2007Natur.448...83R
5082:10.1038/nature05983
5074:2007Natur.448...83R
4845:10.1101/gad.1262504
4757:10.1038/nature01957
4749:2003Natur.425..415L
4645:2007PLSCB...3...37Z
4542:10.1261/rna.7240905
4395:10.1261/rna.7135204
4273:10.1038/nature07242
4265:2008Natur.455...64B
4218:10.1038/nature07228
4210:2008Natur.455...58S
4167:10.1038/nature03315
4159:2005Natur.433..769L
3730:10.1261/rna.2183803
3403:2005PNAS..10210898H
3289:2004Sci...303...83C
3246:10.1038/nature03076
3238:2004Natur.432..226P
3202:10.1677/joe.1.06426
3067:2005Sci...309..310W
3016:2001Sci...294..862L
2960:2001Sci...294..858L
2894:2001Sci...294..853L
2838:2000Natur.408...86P
2784:2000Natur.403..901R
2541:2002CBio...12..735L
2489:10.1101/gad.1074403
2314:10.1038/nature09267
2306:2010Natur.466..835G
1979:C19MC miRNA cluster
1939:MicroRNA Biosensors
1866:human herpesvirus-6
1683:synaptic plasticity
1579:Parkinson's disease
1575:Alzheimer's disease
1559:synaptic plasticity
1489:progenitors, fewer
1317:DNA mismatch repair
1146:locked nucleic acid
1142:microRNA sequencing
1008:Moreover, miRNA as
998:cerebrospinal fluid
620:(GTP) bound to the
138:double-stranded RNA
15577:Chloroplast
15420:modified Messenger
15383:Ribonucleic acids
15218:Trans-acting siRNA
15213:Small temporal RNA
15188:Long noncoding RNA
13794:precursor families
13311:10.1093/jxb/erq437
12162:10.1038/mp.2009.84
11909:10.1093/hmg/ddq311
11439:10.1038/ncomms4000
11294:Scientific Reports
10626:10.1093/nar/gkg884
10064:10.1038/onc.2014.9
9405:10.1093/nar/gkn714
9297:10.1093/nar/gkp294
9248:10.1093/nar/gkm952
9015:10.1093/nar/gkl175
8909:10.1093/nar/gkt852
8669:10.1093/nar/gkh968
8520:10.1093/nar/gni178
8428:10.1093/nar/gkl356
7923:10.1093/gbe/evq009
7351:(1): e2020454118.
6137:10.1101/gad.974702
5763:10.1101/gad.992502
5240:10.1093/nar/gkn479
4788:MicroRNA Protocols
4339:10.1101/gr.2722704
4111:10.1093/nar/gkr330
3779:10.1093/nar/gkj112
3456:(Suppl 2): S52–7.
2440:10.1093/nar/gkz885
2391:10.1093/nar/gkz097
1806:insulin resistance
1779:ectopic expression
1670:likely originate.
1287:cell proliferation
1216:
1192:Inherited diseases
994:circulating miRNAs
941:kinetic signatures
846:
831:Cellular functions
709:
614:karyopherin family
606:
510:
463:Nuclear processing
445:RNA polymerase III
394:
249:A year later, the
62:
54:
15750:
15749:
15627:Xeno
15589:Complementary
15562:Deoxyribonucleic
15556:
15555:
15533:Small hairpin
15309:
15308:
15223:Short hairpin RNA
15132:Small nuclear RNA
15089:Protein synthesis
15047:
15046:
13736:ChIPBase database
13150:BMC Plant Biology
12984:10.1111/jth.14290
12977:(11): 2233–2245.
12940:10.1111/jth.12788
12155:(12): 1176–1189.
11994:(13): 1368–1376.
11902:(20): 3959–3969.
11843:(14): 5614–5619.
10209:978-1-4641-8339-3
10113:: 1888–1904.e24.
10095:Peking University
10014:10.1002/path.4348
9966:10.1002/jcp.26514
9765:10.1002/gcc.20902
9656:10.1002/ijc.34408
9109:10.1021/bi060307w
8287:(6164): 1242592.
7407:. 6 January 2021.
7302:(10): 1528–1535.
7027:978-1-904455-25-7
6291:10.1038/nsmb.1762
6021:(27): 17897–901.
5578:Sohel MH (2016).
5515:978-3-540-75156-4
4805:978-1-59745-123-9
3690:978-3-319-23729-9
3397:(31): 10898–903.
2434:(D1): D132–D141.
2300:(7308): 835–840.
1873:Target prediction
1699:nucleus accumbens
1693:and inflammatory
1687:neuroinflammation
1663:prefrontal cortex
1603:anxiety disorders
1285:genes to control
1247:colorectal cancer
1121:-free equipment.
1079:Mnemiopsis leidyi
975:sequence homology
922:Sequestration in
816:uridyltransferase
558:. Most commonly,
522:DiGeorge Syndrome
473:crystal structure
421:RNA polymerase II
312:Rattus norvegicus
34:Mitochondrial DNA
16:(Redirected from
15785:
15740:
15739:
15717:Yeast
15538:Small temporal
15468:Piwi-interacting
15396:
15395:
15392:
15373:Deoxynucleotides
15336:
15329:
15322:
15313:
15312:
15074:
15067:
15060:
15051:
15050:
13785:
13778:
13771:
13762:
13761:
13688:
13678:
13648:
13623:(5589): 2002–3.
13605:
13595:
13552:
13551:
13541:
13531:
13507:
13501:
13500:
13490:
13480:
13456:
13450:
13449:
13439:
13407:
13401:
13400:
13388:
13382:
13381:
13371:
13339:
13333:
13332:
13322:
13305:(8): 2495–2506.
13294:
13288:
13287:
13277:
13267:
13243:
13237:
13236:
13203:(5): 1551–1567.
13192:
13186:
13185:
13175:
13165:
13141:
13135:
13134:
13124:
13114:
13090:
13081:
13080:
13055:(8): H931–H939.
13044:
13038:
13037:
13027:
13003:
12997:
12996:
12986:
12962:
12953:
12952:
12942:
12918:
12912:
12911:
12901:
12891:
12874:(52): 21075–80.
12859:
12853:
12852:
12842:
12810:
12804:
12803:
12767:
12761:
12760:
12750:
12726:
12720:
12719:
12709:
12680:
12674:
12673:
12637:
12631:
12630:
12620:
12588:
12579:
12578:
12568:
12558:
12526:
12520:
12519:
12509:
12477:
12471:
12470:
12460:
12450:
12418:
12412:
12411:
12401:
12369:
12363:
12362:
12352:
12320:
12309:
12308:
12298:
12266:
12260:
12259:
12249:
12217:
12211:
12210:
12208:
12206:
12191:
12185:
12184:
12174:
12164:
12140:
12134:
12133:
12123:
12113:
12103:
12079:
12073:
12072:
12070:
12063:
12029:
12020:
12014:
12013:
12003:
11979:
11973:
11972:
11962:
11952:
11928:
11922:
11921:
11911:
11887:
11881:
11880:
11870:
11860:
11828:
11822:
11821:
11811:
11801:
11769:
11763:
11762:
11726:
11720:
11719:
11709:
11699:
11675:
11669:
11668:
11658:
11648:
11624:
11618:
11617:
11607:
11575:
11566:
11565:
11532:(6917): 868–74.
11521:
11515:
11514:
11504:
11494:
11485:(53): 44083–96.
11470:
11461:
11460:
11450:
11410:
11371:
11370:
11342:
11336:
11335:
11325:
11285:
11279:
11278:
11234:
11228:
11227:
11191:
11185:
11184:
11148:
11142:
11141:
11139:
11137:
11129:
11118:Retraction Watch
11097:
11087:
11077:
11053:
11047:
11046:
11013:(7048): 214–20.
11002:
10996:
10995:
10985:
10953:
10947:
10946:
10936:
10926:
10909:(48): 18255–60.
10894:
10888:
10887:
10877:
10853:
10847:
10846:
10836:
10812:
10803:
10802:
10792:
10782:
10746:
10740:
10739:
10729:
10705:
10699:
10698:
10688:
10678:
10663:Genome Integrity
10654:
10648:
10647:
10637:
10605:
10599:
10598:
10588:
10564:
10558:
10557:
10521:
10515:
10514:
10504:
10472:
10466:
10465:
10455:
10423:
10412:
10411:
10401:
10391:
10359:
10353:
10352:
10342:
10327:Gastroenterology
10318:
10312:
10311:
10301:
10278:Gastroenterology
10269:
10263:
10262:
10252:
10220:
10214:
10213:
10195:
10189:
10188:
10178:
10154:
10148:
10147:
10145:
10143:
10122:
10091:
10085:
10084:
10066:
10042:
10036:
10035:
10025:
9993:
9987:
9986:
9968:
9959:(8): 5574–5588.
9944:
9938:
9937:
9927:
9903:
9897:
9894:
9888:
9887:
9877:
9867:
9835:
9826:
9825:
9815:
9791:
9785:
9784:
9744:
9738:
9737:
9727:
9717:
9693:
9687:
9686:
9668:
9658:
9649:(9): 1989–2001.
9634:
9628:
9627:
9617:
9607:
9575:
9569:
9568:
9558:
9526:
9520:
9519:
9509:
9477:
9471:
9470:
9433:
9427:
9426:
9416:
9384:
9378:
9377:
9367:
9357:
9325:
9319:
9318:
9308:
9276:
9270:
9269:
9259:
9227:
9221:
9220:
9210:
9178:
9172:
9171:
9161:
9151:
9136:BMC Neuroscience
9127:
9121:
9120:
9092:
9086:
9085:
9075:
9043:
9037:
9036:
9026:
8994:
8988:
8987:
8969:
8937:
8931:
8930:
8920:
8888:
8882:
8881:
8879:
8877:
8869:
8839:
8829:
8819:
8795:
8789:
8788:
8778:
8746:
8740:
8739:
8729:
8697:
8691:
8690:
8680:
8648:
8642:
8641:
8631:
8621:
8597:
8591:
8590:
8580:
8548:
8542:
8541:
8531:
8499:
8493:
8492:
8461:Nature Protocols
8456:
8450:
8449:
8439:
8407:
8401:
8400:
8398:
8396:
8372:
8366:
8365:
8355:
8345:
8321:
8315:
8314:
8304:
8272:
8266:
8265:
8255:
8223:
8217:
8216:
8206:
8189:(7298): 617–21.
8174:
8168:
8167:
8127:
8121:
8120:
8084:
8078:
8077:
8067:
8035:
8029:
8028:
8018:
7994:
7988:
7987:
7951:
7945:
7944:
7934:
7902:
7891:
7890:
7880:
7870:
7838:
7829:
7828:
7792:
7786:
7785:
7749:
7740:
7739:
7729:
7719:
7695:
7689:
7688:
7678:
7646:
7640:
7639:
7621:
7597:
7584:
7583:
7554:
7548:
7547:
7511:
7505:
7504:
7486:
7466:
7460:
7459:
7449:
7417:
7411:
7408:
7396:
7378:
7368:
7336:
7330:
7329:
7319:
7287:
7281:
7280:
7270:
7246:
7240:
7239:
7229:
7197:
7188:
7187:
7177:
7145:
7139:
7138:
7136:
7134:
7126:
7111:Retraction Watch
7090:
7080:
7070:
7038:
7032:
7031:
7007:
7001:
7000:
6964:
6958:
6957:
6947:
6923:
6917:
6916:
6906:
6874:
6868:
6867:
6857:
6847:
6823:
6817:
6816:
6806:
6781:(6078): 237–40.
6766:
6760:
6759:
6749:
6709:
6703:
6702:
6692:
6660:
6654:
6653:
6635:
6603:
6597:
6596:
6579:(11–12): 452–8.
6568:
6562:
6561:
6551:
6519:
6510:
6509:
6499:
6475:
6469:
6468:
6458:
6448:
6424:
6418:
6417:
6407:
6375:
6364:
6363:
6319:
6313:
6312:
6302:
6270:
6259:
6258:
6248:
6224:
6218:
6217:
6207:
6197:
6165:
6159:
6158:
6148:
6116:
6110:
6109:
6099:
6089:
6057:
6051:
6050:
6040:
6030:
6006:
5997:
5996:
5986:
5954:
5948:
5947:
5937:
5905:
5899:
5898:
5888:
5864:
5858:
5857:
5847:
5823:
5817:
5816:
5806:
5782:
5776:
5775:
5765:
5741:
5732:
5731:
5695:
5689:
5688:
5678:
5655:Current Genomics
5646:
5640:
5639:
5611:
5602:
5601:
5599:
5575:
5569:
5568:
5558:
5534:
5528:
5527:
5498:RNA Interference
5493:
5487:
5486:
5476:
5444:
5438:
5437:
5427:
5403:
5392:
5391:
5390:
5384:
5349:
5340:
5339:
5311:
5305:
5304:
5268:
5262:
5261:
5251:
5219:
5210:
5209:
5199:
5167:
5161:
5160:
5150:
5110:
5104:
5103:
5093:
5053:
5047:
5046:
5028:
5004:
4998:
4997:
4987:
4955:
4949:
4948:
4938:
4914:
4908:
4907:
4897:
4873:
4867:
4866:
4856:
4824:
4818:
4817:
4783:
4777:
4776:
4732:
4726:
4725:
4715:
4683:
4677:
4676:
4666:
4656:
4624:
4618:
4617:
4607:
4584:The EMBO Journal
4575:
4564:
4563:
4553:
4521:
4515:
4514:
4504:
4481:The EMBO Journal
4472:
4466:
4465:
4447:
4432:The FEBS Journal
4423:
4417:
4416:
4406:
4374:
4361:
4360:
4350:
4333:(10A): 1902–10.
4318:
4309:
4308:
4302:
4294:
4284:
4244:
4238:
4237:
4193:
4187:
4186:
4153:(7027): 769–73.
4142:
4133:
4132:
4122:
4090:
4084:
4083:
4047:
4041:
4040:
4004:
3998:
3997:
3987:
3955:
3949:
3948:
3938:
3914:
3903:
3902:
3892:
3882:
3858:
3852:
3851:
3841:
3831:
3807:
3801:
3800:
3790:
3758:
3752:
3751:
3741:
3709:
3703:
3702:
3673:MicroRNA: Cancer
3668:
3662:
3661:
3651:
3627:
3621:
3620:
3610:
3600:
3576:
3570:
3569:
3541:
3535:
3534:
3524:
3501:The AAPS Journal
3492:
3486:
3485:
3475:
3465:
3441:
3435:
3434:
3424:
3414:
3382:
3376:
3375:
3365:
3333:
3327:
3326:
3308:
3272:
3266:
3265:
3232:(7014): 226–30.
3221:
3215:
3214:
3204:
3180:
3174:
3173:
3163:
3139:
3133:
3132:
3104:
3095:
3094:
3050:
3044:
3043:
2999:
2988:
2987:
2954:(5543): 858–62.
2943:
2932:
2931:
2913:
2877:
2866:
2865:
2846:10.1038/35040556
2821:
2812:
2811:
2792:10.1038/35002607
2767:
2758:
2757:
2747:
2723:
2712:
2711:
2701:
2669:
2663:
2662:
2652:
2620:
2609:
2608:
2598:
2574:
2563:
2562:
2552:
2520:
2511:
2510:
2500:
2468:
2462:
2461:
2451:
2419:
2413:
2412:
2402:
2385:(7): 3353–3364.
2370:
2364:
2352:
2346:
2345:
2335:
2325:
2285:
2279:
2278:
2242:
2236:
2235:
2199:
2193:
2192:
2182:
2150:
2139:
2138:
2128:
2096:
2090:
2089:
2079:
2055:
2044:
2043:
2033:
2001:
1964:RNA interference
1908:
1903:
1902:
1615:ischemic strokes
1599:bipolar disorder
1595:major depression
1396:cardiomyopathies
1259:Hodgkin lymphoma
1134:quantitative PCR
1024:miRNAs are well
1014:circadian rhythm
814:(U) residues by
603:
507:
492:ribonuclease III
340:
336:
331:
327:
95:molecules, then
21:
15793:
15792:
15788:
15787:
15786:
15784:
15783:
15782:
15768:Gene expression
15753:
15752:
15751:
15746:
15728:
15669:Cloning vectors
15663:
15649:Locked
15613:
15563:
15552:
15511:
15439:
15386:
15385:
15377:
15346:
15340:
15310:
15305:
15279:
15260:Retrotransposon
15248:
15227:
15166:Gene regulation
15161:
15120:
15084:
15078:
15048:
15043:
14992:
14786:
14655:
14519:
14358:
14232:
14006:
13795:
13789:
13758:
13730:Wayback Machine
13696:
13691:
13560:
13558:Further reading
13555:
13508:
13504:
13457:
13453:
13408:
13404:
13389:
13385:
13340:
13336:
13295:
13291:
13244:
13240:
13193:
13189:
13142:
13138:
13091:
13084:
13045:
13041:
13004:
13000:
12963:
12956:
12919:
12915:
12860:
12856:
12811:
12807:
12768:
12764:
12727:
12723:
12681:
12677:
12648:(9): 997–1004.
12638:
12634:
12589:
12582:
12527:
12523:
12478:
12474:
12419:
12415:
12370:
12366:
12335:(11): 1928–37.
12321:
12312:
12267:
12263:
12232:(10): 521–528.
12218:
12214:
12204:
12202:
12201:. 15 March 2019
12193:
12192:
12188:
12141:
12137:
12080:
12076:
12068:
12027:
12021:
12017:
11980:
11976:
11929:
11925:
11888:
11884:
11829:
11825:
11784:(1): e0279029.
11770:
11766:
11743:10.1038/nrn2763
11737:(12): 842–849.
11727:
11723:
11676:
11672:
11625:
11621:
11576:
11569:
11522:
11518:
11471:
11464:
11411:
11374:
11343:
11339:
11286:
11282:
11245:(5824): 575–9.
11235:
11231:
11196:Nature Medicine
11192:
11188:
11153:Nature Medicine
11149:
11145:
11131:
11123:
11121:
11098:
11068:(17): 12363–7.
11054:
11050:
11003:
10999:
10954:
10950:
10895:
10891:
10854:
10850:
10813:
10806:
10747:
10743:
10706:
10702:
10655:
10651:
10620:(23): 6841–51.
10606:
10602:
10573:Cancer Research
10565:
10561:
10522:
10518:
10473:
10469:
10424:
10415:
10360:
10356:
10319:
10315:
10270:
10266:
10221:
10217:
10210:
10196:
10192:
10155:
10151:
10141:
10139:
10092:
10088:
10043:
10039:
9994:
9990:
9945:
9941:
9904:
9900:
9895:
9891:
9850:(6): e0156871.
9836:
9829:
9792:
9788:
9745:
9741:
9694:
9690:
9635:
9631:
9576:
9572:
9541:(10): 1026–30.
9535:Nature Genetics
9527:
9523:
9478:
9474:
9439:Nature Genetics
9434:
9430:
9385:
9381:
9326:
9322:
9277:
9273:
9228:
9224:
9179:
9175:
9128:
9124:
9103:(23): 7347–55.
9093:
9089:
9044:
9040:
8995:
8991:
8952:(5848): 271–4.
8938:
8934:
8889:
8885:
8871:
8863:
8861:
8840:
8796:
8792:
8747:
8743:
8706:Nature Genetics
8698:
8694:
8663:(21): 6284–91.
8649:
8645:
8598:
8594:
8549:
8545:
8500:
8496:
8457:
8453:
8422:(9): 2791–802.
8408:
8404:
8394:
8392:
8391:on 19 July 2010
8373:
8369:
8322:
8318:
8273:
8269:
8224:
8220:
8175:
8171:
8136:Marine Genomics
8128:
8124:
8085:
8081:
8036:
8032:
7995:
7991:
7962:(12): 1282–90.
7956:Nature Genetics
7952:
7948:
7903:
7894:
7839:
7832:
7793:
7789:
7750:
7743:
7696:
7692:
7647:
7643:
7598:
7587:
7555:
7551:
7528:10.1038/nrg1990
7512:
7508:
7484:10.1.1.194.1598
7467:
7463:
7418:
7414:
7399:
7337:
7333:
7288:
7284:
7247:
7243:
7198:
7191:
7146:
7142:
7128:
7120:
7114:
7092:(Erratum:
7091:
7039:
7035:
7028:
7008:
7004:
6981:10.1038/nrg1379
6965:
6961:
6924:
6920:
6875:
6871:
6824:
6820:
6767:
6763:
6724:(6078): 233–7.
6710:
6706:
6661:
6657:
6604:
6600:
6569:
6565:
6528:Genome Research
6520:
6513:
6476:
6472:
6425:
6421:
6376:
6367:
6330:(7263): 546–9.
6320:
6316:
6271:
6262:
6225:
6221:
6180:(23): 2149–55.
6174:Current Biology
6166:
6162:
6117:
6113:
6058:
6054:
6007:
6000:
5955:
5951:
5906:
5902:
5865:
5861:
5824:
5820:
5797:(40): 42230–9.
5783:
5779:
5742:
5735:
5712:10.1038/nrm2085
5696:
5692:
5647:
5643:
5612:
5605:
5576:
5572:
5535:
5531:
5516:
5494:
5490:
5459:(7355): 201–5.
5445:
5441:
5404:
5395:
5385:
5350:
5343:
5312:
5308:
5269:
5265:
5234:(16): 5270–80.
5220:
5213:
5168:
5164:
5125:(7149): 83–86.
5111:
5107:
5068:(7149): 83–86.
5054:
5050:
5019:(22): 3986–90.
5005:
5001:
4956:
4952:
4915:
4911:
4874:
4870:
4839:(24): 3016–27.
4825:
4821:
4806:
4784:
4780:
4743:(6956): 415–9.
4733:
4729:
4684:
4680:
4625:
4621:
4590:(20): 4051–60.
4576:
4567:
4522:
4518:
4473:
4469:
4424:
4420:
4389:(12): 1957–66.
4375:
4364:
4327:Genome Research
4319:
4312:
4296:
4295:
4259:(7209): 64–71.
4245:
4241:
4204:(7209): 58–63.
4194:
4190:
4143:
4136:
4105:(16): 6845–53.
4091:
4087:
4052:Nature Genetics
4048:
4044:
4009:Nature Genetics
4005:
4001:
3970:(10): 1346–50.
3956:
3952:
3915:
3906:
3859:
3855:
3808:
3804:
3759:
3755:
3710:
3706:
3691:
3669:
3665:
3628:
3624:
3577:
3573:
3542:
3538:
3493:
3489:
3442:
3438:
3383:
3379:
3334:
3330:
3273:
3269:
3222:
3218:
3181:
3177:
3140:
3136:
3105:
3098:
3061:(5732): 310–1.
3051:
3047:
3010:(5543): 862–4.
3000:
2991:
2944:
2935:
2888:(5543): 853–8.
2878:
2869:
2822:
2815:
2778:(6772): 901–6.
2768:
2761:
2724:
2715:
2670:
2666:
2629:Genome Research
2621:
2612:
2575:
2566:
2529:Current Biology
2521:
2514:
2483:(8): 991–1008.
2469:
2465:
2420:
2416:
2371:
2367:
2353:
2349:
2286:
2282:
2259:10.1038/nrg3965
2243:
2239:
2216:10.1038/nrg3965
2200:
2196:
2151:
2142:
2097:
2093:
2056:
2047:
2002:
1991:
1987:
1919:Gene expression
1904:
1897:
1894:
1881:
1875:
1860:
1851:
1842:
1739:
1733:in treatments.
1731:pharmaceuticals
1703:basal forebrain
1668:decision making
1635:gene expression
1629:, specifically
1623:
1611:
1547:
1479:
1467:
1438:
1423:
1411:atherosclerosis
1404:
1387:
1295:DNA replication
1279:
1228:B-cell receptor
1208:
1194:
1185:
1110:
1022:
956:gene expression
894:DNA methylation
833:
796:exoribonuclease
782:
760:
711:Members of the
693:
687:
666:
630:
595:
580:
551:, and mammals.
499:
465:
417:
387:
374:
338:
334:
329:
325:
292:
199:non-coding RNAs
158:
41:
28:
23:
22:
15:
12:
11:
5:
15791:
15781:
15780:
15778:Non-coding RNA
15775:
15770:
15765:
15748:
15747:
15745:
15744:
15733:
15730:
15729:
15727:
15726:
15725:
15724:
15719:
15714:
15709:
15699:
15694:
15689:
15684:
15679:
15673:
15671:
15665:
15664:
15662:
15661:
15656:
15654:Peptide
15651:
15646:
15645:
15644:
15639:
15634:
15632:Glycol
15623:
15621:
15615:
15614:
15612:
15611:
15606:
15601:
15596:
15591:
15586:
15585:
15584:
15579:
15568:
15566:
15558:
15557:
15554:
15553:
15551:
15550:
15545:
15540:
15535:
15530:
15525:
15519:
15517:
15513:
15512:
15510:
15509:
15504:
15503:
15502:
15497:
15492:
15487:
15477:
15472:
15471:
15470:
15465:
15460:
15449:
15447:
15441:
15440:
15438:
15437:
15432:
15427:
15422:
15417:
15416:
15415:
15404:
15402:
15393:
15379:
15378:
15376:
15375:
15370:
15365:
15360:
15354:
15352:
15348:
15347:
15344:nucleic acids
15339:
15338:
15331:
15324:
15316:
15307:
15306:
15304:
15303:
15298:
15293:
15291:Telomerase RNA
15287:
15285:
15281:
15280:
15278:
15277:
15272:
15267:
15262:
15256:
15254:
15250:
15249:
15247:
15246:
15241:
15235:
15233:
15229:
15228:
15226:
15225:
15220:
15215:
15210:
15205:
15200:
15195:
15190:
15185:
15180:
15175:
15169:
15167:
15163:
15162:
15160:
15159:
15154:
15149:
15144:
15139:
15134:
15128:
15126:
15125:RNA processing
15122:
15121:
15119:
15118:
15113:
15108:
15103:
15098:
15092:
15090:
15086:
15085:
15077:
15076:
15069:
15062:
15054:
15045:
15044:
15042:
15041:
15036:
15031:
15026:
15021:
15016:
15011:
15006:
15000:
14998:
14994:
14993:
14991:
14990:
14985:
14980:
14975:
14970:
14965:
14960:
14955:
14950:
14945:
14940:
14935:
14930:
14925:
14920:
14915:
14910:
14905:
14900:
14895:
14890:
14885:
14880:
14875:
14870:
14865:
14860:
14855:
14850:
14845:
14840:
14835:
14830:
14825:
14820:
14815:
14810:
14805:
14800:
14794:
14792:
14788:
14787:
14785:
14784:
14779:
14774:
14769:
14764:
14759:
14754:
14749:
14744:
14739:
14734:
14729:
14724:
14719:
14714:
14709:
14704:
14699:
14694:
14689:
14684:
14679:
14674:
14669:
14663:
14661:
14657:
14656:
14654:
14653:
14648:
14643:
14638:
14633:
14628:
14623:
14618:
14613:
14608:
14603:
14598:
14593:
14588:
14583:
14578:
14573:
14568:
14563:
14558:
14553:
14548:
14543:
14538:
14533:
14527:
14525:
14521:
14520:
14518:
14517:
14512:
14507:
14502:
14497:
14492:
14487:
14482:
14477:
14472:
14467:
14462:
14457:
14452:
14447:
14442:
14437:
14432:
14427:
14422:
14417:
14412:
14407:
14402:
14397:
14392:
14387:
14382:
14377:
14372:
14366:
14364:
14360:
14359:
14357:
14356:
14351:
14346:
14341:
14336:
14331:
14326:
14321:
14316:
14311:
14306:
14301:
14296:
14291:
14286:
14281:
14276:
14271:
14266:
14261:
14256:
14251:
14246:
14240:
14238:
14234:
14233:
14231:
14230:
14225:
14220:
14215:
14210:
14205:
14200:
14195:
14190:
14185:
14180:
14175:
14170:
14165:
14160:
14155:
14150:
14145:
14140:
14135:
14130:
14125:
14120:
14115:
14110:
14105:
14100:
14095:
14090:
14085:
14080:
14075:
14070:
14065:
14060:
14055:
14050:
14045:
14040:
14035:
14030:
14025:
14020:
14014:
14012:
14008:
14007:
14005:
14004:
13999:
13994:
13989:
13984:
13979:
13974:
13969:
13964:
13959:
13954:
13949:
13944:
13939:
13934:
13929:
13924:
13919:
13914:
13909:
13904:
13899:
13894:
13889:
13884:
13879:
13874:
13869:
13864:
13859:
13854:
13849:
13844:
13839:
13834:
13829:
13824:
13819:
13814:
13809:
13803:
13801:
13797:
13796:
13788:
13787:
13780:
13773:
13765:
13756:
13755:
13749:
13743:
13733:
13720:
13714:
13708:
13702:
13695:
13694:External links
13692:
13690:
13689:
13649:
13606:
13561:
13559:
13556:
13554:
13553:
13502:
13471:(4): 8179–87.
13451:
13422:(3): 1638–49.
13402:
13383:
13334:
13289:
13238:
13187:
13136:
13105:(3): 413–426.
13082:
13039:
12998:
12954:
12933:(2): 170–181.
12913:
12854:
12805:
12778:(3): 132–141.
12762:
12741:(12): 7960–7.
12721:
12675:
12632:
12603:(9): 1402–14.
12580:
12521:
12492:(13): 4581–8.
12472:
12433:(12): e82565.
12413:
12364:
12310:
12261:
12212:
12195:"Stroke Facts"
12186:
12135:
12074:
12015:
11974:
11923:
11882:
11823:
11764:
11721:
11670:
11619:
11590:(10): e12537.
11567:
11516:
11462:
11372:
11337:
11280:
11229:
11208:10.1038/nm1582
11186:
11165:10.1038/nm1569
11143:
11048:
10997:
10968:(6): 1137–41.
10948:
10889:
10848:
10804:
10741:
10700:
10649:
10600:
10579:(10): 3371–5.
10559:
10516:
10481:Neuro-Oncology
10467:
10432:Neuro-Oncology
10413:
10374:(15): 6982–7.
10354:
10333:(5): 1160–71.
10313:
10284:(6): 2044–58.
10264:
10235:(3): 151–158.
10215:
10208:
10190:
10169:(3): 379–385.
10163:Carcinogenesis
10149:
10086:
10037:
9988:
9939:
9898:
9889:
9827:
9786:
9759:(10): 812–22.
9739:
9688:
9629:
9570:
9547:10.1038/ng.915
9521:
9472:
9451:10.1038/ng.355
9428:
9379:
9320:
9271:
9222:
9187:Molecular Cell
9173:
9122:
9087:
9038:
8989:
8932:
8883:
8790:
8741:
8718:10.1038/ng1953
8692:
8643:
8592:
8563:(9): 1461–70.
8543:
8494:
8451:
8402:
8367:
8316:
8267:
8232:The Plant Cell
8218:
8169:
8122:
8079:
8050:(4): 1074–89.
8044:The Plant Cell
8030:
8009:(5): 892–902.
7989:
7968:10.1038/ng1478
7946:
7892:
7853:(8): 2946–50.
7830:
7787:
7741:
7704:Genome Biology
7690:
7661:(10): 578–87.
7641:
7585:
7549:
7506:
7461:
7432:(6): 1658–73.
7426:The Plant Cell
7412:
7410:
7409:
7331:
7282:
7255:Prog Neurobiol
7241:
7212:(9): 1617–27.
7189:
7140:
7053:(5): 1608–13.
7033:
7026:
7010:Li LC (2008).
7002:
6975:(7): 522–531.
6959:
6938:(6): 1133–46.
6918:
6869:
6818:
6761:
6704:
6655:
6598:
6563:
6534:(10): 1602–9.
6511:
6470:
6433:Genome Biology
6419:
6390:(9): 1635–55.
6365:
6314:
6260:
6219:
6160:
6111:
6052:
5998:
5969:(18): 2840–5.
5949:
5900:
5879:(2): 199–208.
5859:
5818:
5777:
5756:(9): 1025–31.
5733:
5690:
5641:
5603:
5590:(2): 175–186.
5570:
5529:
5514:
5488:
5439:
5393:
5341:
5322:(10): 1171–6.
5306:
5263:
5211:
5182:(2): 328–336.
5176:Molecular Cell
5162:
5105:
5048:
4999:
4950:
4909:
4888:(5): 887–901.
4868:
4819:
4804:
4778:
4727:
4678:
4619:
4565:
4516:
4467:
4418:
4362:
4310:
4239:
4188:
4134:
4085:
4064:10.1038/ng1536
4058:(5): 495–500.
4042:
4021:10.1038/ng1798
3999:
3964:Bioinformatics
3950:
3904:
3853:
3816:Human Genomics
3802:
3753:
3704:
3689:
3663:
3622:
3571:
3536:
3487:
3436:
3377:
3328:
3283:(5654): 83–6.
3267:
3216:
3175:
3134:
3096:
3045:
2989:
2933:
2867:
2832:(6808): 86–9.
2813:
2759:
2713:
2664:
2610:
2564:
2512:
2463:
2414:
2365:
2347:
2280:
2253:(7): 421–433.
2237:
2210:(7): 421–433.
2194:
2140:
2091:
2070:(2): 281–297.
2045:
1988:
1986:
1983:
1982:
1981:
1976:
1971:
1966:
1961:
1956:
1951:
1946:
1941:
1936:
1931:
1926:
1921:
1916:
1910:
1909:
1906:Biology portal
1893:
1890:
1874:
1871:
1859:
1856:
1850:
1847:
1841:
1838:
1781:of the miRNAs
1738:
1735:
1659:cell signaling
1622:
1619:
1610:
1607:
1551:nervous system
1546:
1545:Nervous system
1543:
1497:, progressive
1478:
1477:Kidney disease
1475:
1466:
1463:
1437:
1434:
1422:
1419:
1403:
1400:
1386:
1383:
1331:island of the
1278:
1275:
1265:combined with
1207:
1204:
1193:
1190:
1184:
1181:
1109:
1106:
1021:
1018:
1010:miR-183/96/182
984:non-coding DNA
937:
936:
933:
932:mRNA cleavage;
930:
927:
920:
917:
914:
911:
908:
854:messenger RNAs
832:
829:
781:
778:
759:
756:
737:protein kinase
721:ribonuclease-H
689:Main article:
686:
683:
665:
662:
629:
626:
579:
578:Nuclear export
576:
464:
461:
431:with multiple
429:polyadenylated
416:
413:
386:
383:
373:
370:
342:
341:
332:
291:
288:
221:idiosyncrasy.
157:
154:
123:
122:
115:
108:
73:non-coding RNA
36:(m(t)DNA); or
26:
9:
6:
4:
3:
2:
15790:
15779:
15776:
15774:
15771:
15769:
15766:
15764:
15761:
15760:
15758:
15743:
15735:
15734:
15731:
15723:
15720:
15718:
15715:
15713:
15710:
15708:
15705:
15704:
15703:
15700:
15698:
15695:
15693:
15690:
15688:
15685:
15683:
15680:
15678:
15675:
15674:
15672:
15670:
15666:
15660:
15657:
15655:
15652:
15650:
15647:
15643:
15640:
15638:
15637:Threose
15635:
15633:
15630:
15629:
15628:
15625:
15624:
15622:
15620:
15616:
15610:
15607:
15605:
15602:
15600:
15597:
15595:
15594:Deoxyribozyme
15592:
15590:
15587:
15583:
15582:Mitochondrial
15580:
15578:
15575:
15574:
15573:
15570:
15569:
15567:
15565:
15559:
15549:
15546:
15544:
15541:
15539:
15536:
15534:
15531:
15529:
15526:
15524:
15521:
15520:
15518:
15514:
15508:
15505:
15501:
15498:
15496:
15493:
15491:
15488:
15486:
15483:
15482:
15481:
15478:
15476:
15473:
15469:
15466:
15464:
15461:
15459:
15456:
15455:
15454:
15451:
15450:
15448:
15446:
15442:
15436:
15433:
15431:
15428:
15426:
15423:
15421:
15418:
15414:
15411:
15410:
15409:
15406:
15405:
15403:
15401:
15400:Translational
15397:
15394:
15390:
15384:
15380:
15374:
15371:
15369:
15366:
15364:
15361:
15359:
15356:
15355:
15353:
15349:
15345:
15337:
15332:
15330:
15325:
15323:
15318:
15317:
15314:
15302:
15299:
15297:
15294:
15292:
15289:
15288:
15286:
15282:
15276:
15273:
15271:
15268:
15266:
15263:
15261:
15258:
15257:
15255:
15251:
15245:
15244:SECIS element
15242:
15240:
15237:
15236:
15234:
15230:
15224:
15221:
15219:
15216:
15214:
15211:
15209:
15206:
15204:
15201:
15199:
15196:
15194:
15191:
15189:
15186:
15184:
15181:
15179:
15176:
15174:
15173:Antisense RNA
15171:
15170:
15168:
15164:
15158:
15155:
15153:
15150:
15148:
15145:
15143:
15140:
15138:
15135:
15133:
15130:
15129:
15127:
15123:
15117:
15114:
15112:
15109:
15107:
15104:
15102:
15101:Ribosomal RNA
15099:
15097:
15096:Messenger RNA
15094:
15093:
15091:
15087:
15083:
15075:
15070:
15068:
15063:
15061:
15056:
15055:
15052:
15040:
15037:
15035:
15032:
15030:
15027:
15025:
15022:
15020:
15017:
15015:
15012:
15010:
15007:
15005:
15002:
15001:
14999:
14995:
14989:
14986:
14984:
14981:
14979:
14976:
14974:
14971:
14969:
14966:
14964:
14961:
14959:
14956:
14954:
14951:
14949:
14946:
14944:
14941:
14939:
14936:
14934:
14931:
14929:
14926:
14924:
14921:
14919:
14916:
14914:
14911:
14909:
14906:
14904:
14901:
14899:
14896:
14894:
14891:
14889:
14886:
14884:
14881:
14879:
14876:
14874:
14871:
14869:
14866:
14864:
14861:
14859:
14856:
14854:
14851:
14849:
14846:
14844:
14841:
14839:
14836:
14834:
14831:
14829:
14826:
14824:
14821:
14819:
14816:
14814:
14811:
14809:
14806:
14804:
14801:
14799:
14796:
14795:
14793:
14789:
14783:
14780:
14778:
14775:
14773:
14770:
14768:
14765:
14763:
14760:
14758:
14755:
14753:
14750:
14748:
14745:
14743:
14740:
14738:
14735:
14733:
14730:
14728:
14725:
14723:
14720:
14718:
14715:
14713:
14710:
14708:
14705:
14703:
14700:
14698:
14695:
14693:
14690:
14688:
14685:
14683:
14680:
14678:
14675:
14673:
14670:
14668:
14665:
14664:
14662:
14658:
14652:
14649:
14647:
14644:
14642:
14639:
14637:
14634:
14632:
14629:
14627:
14624:
14622:
14619:
14617:
14614:
14612:
14609:
14607:
14604:
14602:
14599:
14597:
14594:
14592:
14589:
14587:
14584:
14582:
14579:
14577:
14574:
14572:
14569:
14567:
14564:
14562:
14559:
14557:
14554:
14552:
14549:
14547:
14544:
14542:
14539:
14537:
14534:
14532:
14529:
14528:
14526:
14522:
14516:
14513:
14511:
14508:
14506:
14503:
14501:
14498:
14496:
14493:
14491:
14488:
14486:
14483:
14481:
14478:
14476:
14473:
14471:
14468:
14466:
14463:
14461:
14458:
14456:
14453:
14451:
14448:
14446:
14443:
14441:
14438:
14436:
14433:
14431:
14428:
14426:
14423:
14421:
14418:
14416:
14413:
14411:
14408:
14406:
14403:
14401:
14398:
14396:
14393:
14391:
14388:
14386:
14383:
14381:
14378:
14376:
14373:
14371:
14368:
14367:
14365:
14361:
14355:
14352:
14350:
14347:
14345:
14342:
14340:
14337:
14335:
14332:
14330:
14327:
14325:
14322:
14320:
14317:
14315:
14312:
14310:
14307:
14305:
14302:
14300:
14297:
14295:
14292:
14290:
14287:
14285:
14282:
14280:
14277:
14275:
14272:
14270:
14267:
14265:
14262:
14260:
14257:
14255:
14252:
14250:
14247:
14245:
14242:
14241:
14239:
14235:
14229:
14226:
14224:
14221:
14219:
14216:
14214:
14211:
14209:
14206:
14204:
14201:
14199:
14196:
14194:
14191:
14189:
14186:
14184:
14181:
14179:
14176:
14174:
14171:
14169:
14166:
14164:
14161:
14159:
14156:
14154:
14151:
14149:
14146:
14144:
14141:
14139:
14136:
14134:
14131:
14129:
14126:
14124:
14121:
14119:
14116:
14114:
14111:
14109:
14106:
14104:
14101:
14099:
14096:
14094:
14091:
14089:
14086:
14084:
14081:
14079:
14076:
14074:
14071:
14069:
14066:
14064:
14061:
14059:
14056:
14054:
14051:
14049:
14046:
14044:
14041:
14039:
14036:
14034:
14031:
14029:
14026:
14024:
14021:
14019:
14016:
14015:
14013:
14009:
14003:
14000:
13998:
13995:
13993:
13990:
13988:
13985:
13983:
13980:
13978:
13975:
13973:
13970:
13968:
13965:
13963:
13960:
13958:
13955:
13953:
13950:
13948:
13945:
13943:
13940:
13938:
13935:
13933:
13930:
13928:
13925:
13923:
13920:
13918:
13915:
13913:
13910:
13908:
13905:
13903:
13900:
13898:
13895:
13893:
13890:
13888:
13885:
13883:
13880:
13878:
13875:
13873:
13870:
13868:
13865:
13863:
13860:
13858:
13855:
13853:
13850:
13848:
13845:
13843:
13840:
13838:
13835:
13833:
13830:
13828:
13825:
13823:
13820:
13818:
13815:
13813:
13810:
13808:
13805:
13804:
13802:
13798:
13793:
13786:
13781:
13779:
13774:
13772:
13767:
13766:
13763:
13759:
13753:
13750:
13747:
13744:
13741:
13737:
13734:
13731:
13727:
13724:
13721:
13718:
13715:
13712:
13709:
13706:
13703:
13701:
13698:
13697:
13686:
13682:
13677:
13672:
13669:(5): 843–54.
13668:
13664:
13660:
13654:
13651:Discovery of
13650:
13646:
13642:
13638:
13634:
13630:
13626:
13622:
13618:
13612:
13611:
13607:
13603:
13599:
13594:
13589:
13585:
13581:
13577:
13573:
13569:
13563:
13562:
13549:
13545:
13540:
13535:
13530:
13525:
13521:
13517:
13513:
13506:
13498:
13494:
13489:
13484:
13479:
13474:
13470:
13466:
13462:
13455:
13447:
13443:
13438:
13433:
13429:
13425:
13421:
13417:
13413:
13406:
13398:
13394:
13387:
13379:
13375:
13370:
13365:
13361:
13357:
13353:
13349:
13345:
13338:
13330:
13326:
13321:
13316:
13312:
13308:
13304:
13300:
13293:
13285:
13281:
13276:
13271:
13266:
13261:
13257:
13253:
13249:
13242:
13234:
13230:
13226:
13222:
13218:
13214:
13210:
13206:
13202:
13198:
13191:
13183:
13179:
13174:
13169:
13164:
13159:
13155:
13151:
13147:
13140:
13132:
13128:
13123:
13118:
13113:
13108:
13104:
13100:
13096:
13089:
13087:
13078:
13074:
13070:
13066:
13062:
13058:
13054:
13050:
13043:
13035:
13031:
13026:
13021:
13017:
13013:
13009:
13002:
12994:
12990:
12985:
12980:
12976:
12972:
12968:
12961:
12959:
12950:
12946:
12941:
12936:
12932:
12928:
12924:
12917:
12909:
12905:
12900:
12895:
12890:
12885:
12881:
12877:
12873:
12869:
12865:
12858:
12850:
12846:
12841:
12836:
12832:
12828:
12824:
12820:
12816:
12809:
12801:
12797:
12793:
12789:
12785:
12781:
12777:
12773:
12766:
12758:
12754:
12749:
12744:
12740:
12736:
12732:
12725:
12717:
12713:
12708:
12703:
12699:
12695:
12692:(6): 873–83.
12691:
12687:
12679:
12671:
12667:
12663:
12659:
12655:
12651:
12647:
12643:
12636:
12628:
12624:
12619:
12614:
12610:
12606:
12602:
12598:
12594:
12587:
12585:
12576:
12572:
12567:
12562:
12557:
12552:
12548:
12544:
12541:(8): e70945.
12540:
12536:
12532:
12525:
12517:
12513:
12508:
12503:
12499:
12495:
12491:
12487:
12483:
12476:
12468:
12464:
12459:
12454:
12449:
12444:
12440:
12436:
12432:
12428:
12424:
12417:
12409:
12405:
12400:
12395:
12391:
12387:
12384:(3): 286–96.
12383:
12379:
12375:
12368:
12360:
12356:
12351:
12346:
12342:
12338:
12334:
12330:
12326:
12319:
12317:
12315:
12306:
12302:
12297:
12292:
12288:
12284:
12281:(1): 90–101.
12280:
12276:
12272:
12265:
12257:
12253:
12248:
12243:
12239:
12235:
12231:
12227:
12223:
12216:
12200:
12196:
12190:
12182:
12178:
12173:
12168:
12163:
12158:
12154:
12150:
12146:
12139:
12131:
12127:
12122:
12117:
12112:
12107:
12102:
12097:
12093:
12089:
12085:
12078:
12067:
12062:
12057:
12053:
12049:
12045:
12041:
12037:
12033:
12026:
12019:
12011:
12007:
12002:
11997:
11993:
11989:
11985:
11978:
11970:
11966:
11961:
11956:
11951:
11946:
11942:
11938:
11934:
11927:
11919:
11915:
11910:
11905:
11901:
11897:
11893:
11886:
11878:
11874:
11869:
11864:
11859:
11854:
11850:
11846:
11842:
11838:
11834:
11827:
11819:
11815:
11810:
11805:
11800:
11795:
11791:
11787:
11783:
11779:
11775:
11768:
11760:
11756:
11752:
11748:
11744:
11740:
11736:
11732:
11725:
11717:
11713:
11708:
11703:
11698:
11693:
11689:
11685:
11681:
11674:
11666:
11662:
11657:
11652:
11647:
11642:
11638:
11634:
11630:
11623:
11615:
11611:
11606:
11601:
11597:
11593:
11589:
11585:
11581:
11574:
11572:
11563:
11559:
11555:
11551:
11547:
11543:
11539:
11535:
11531:
11527:
11520:
11512:
11508:
11503:
11498:
11493:
11488:
11484:
11480:
11476:
11469:
11467:
11458:
11454:
11449:
11444:
11440:
11436:
11432:
11428:
11424:
11420:
11416:
11409:
11407:
11405:
11403:
11401:
11399:
11397:
11395:
11393:
11391:
11389:
11387:
11385:
11383:
11381:
11379:
11377:
11368:
11364:
11360:
11356:
11352:
11348:
11341:
11333:
11329:
11324:
11319:
11315:
11311:
11307:
11303:
11299:
11295:
11291:
11284:
11276:
11272:
11268:
11264:
11260:
11256:
11252:
11248:
11244:
11240:
11233:
11225:
11221:
11217:
11213:
11209:
11205:
11201:
11197:
11190:
11182:
11178:
11174:
11170:
11166:
11162:
11159:(4): 486–91.
11158:
11154:
11147:
11135:
11127:
11120:
11119:
11114:
11110:
11106:
11102:
11095:
11091:
11086:
11081:
11076:
11071:
11067:
11063:
11059:
11052:
11044:
11040:
11036:
11032:
11028:
11024:
11020:
11016:
11012:
11008:
11001:
10993:
10989:
10984:
10979:
10975:
10971:
10967:
10963:
10959:
10952:
10944:
10940:
10935:
10930:
10925:
10920:
10916:
10912:
10908:
10904:
10900:
10893:
10885:
10881:
10876:
10871:
10868:(3): 258–67.
10867:
10863:
10859:
10852:
10844:
10840:
10835:
10830:
10827:(2): 303–17.
10826:
10822:
10818:
10811:
10809:
10800:
10796:
10791:
10786:
10781:
10776:
10772:
10768:
10765:(6): 2111–6.
10764:
10760:
10756:
10752:
10745:
10737:
10733:
10728:
10723:
10719:
10715:
10711:
10704:
10696:
10692:
10687:
10682:
10677:
10672:
10668:
10664:
10660:
10653:
10645:
10641:
10636:
10631:
10627:
10623:
10619:
10615:
10611:
10604:
10596:
10592:
10587:
10582:
10578:
10574:
10570:
10563:
10555:
10551:
10547:
10543:
10539:
10535:
10531:
10527:
10520:
10512:
10508:
10503:
10498:
10494:
10490:
10486:
10482:
10478:
10471:
10463:
10459:
10454:
10449:
10445:
10441:
10437:
10433:
10429:
10422:
10420:
10418:
10409:
10405:
10400:
10395:
10390:
10385:
10381:
10377:
10373:
10369:
10365:
10358:
10350:
10346:
10341:
10336:
10332:
10328:
10324:
10317:
10309:
10305:
10300:
10295:
10291:
10287:
10283:
10279:
10275:
10268:
10260:
10256:
10251:
10246:
10242:
10238:
10234:
10230:
10226:
10219:
10211:
10205:
10201:
10194:
10186:
10182:
10177:
10172:
10168:
10164:
10160:
10153:
10138:
10134:
10130:
10126:
10121:
10116:
10112:
10111:SciTech Daily
10108:
10104:
10100:
10096:
10090:
10082:
10078:
10074:
10070:
10065:
10060:
10057:(6): 717–25.
10056:
10052:
10048:
10041:
10033:
10029:
10024:
10019:
10015:
10011:
10008:(3): 308–18.
10007:
10003:
9999:
9992:
9984:
9980:
9976:
9972:
9967:
9962:
9958:
9954:
9950:
9943:
9935:
9931:
9926:
9921:
9918:(1): 253–64.
9917:
9913:
9909:
9902:
9893:
9885:
9881:
9876:
9871:
9866:
9861:
9857:
9853:
9849:
9845:
9841:
9834:
9832:
9823:
9819:
9814:
9809:
9805:
9801:
9797:
9790:
9782:
9778:
9774:
9770:
9766:
9762:
9758:
9754:
9750:
9743:
9735:
9731:
9726:
9721:
9716:
9711:
9707:
9703:
9699:
9692:
9684:
9680:
9676:
9672:
9667:
9666:11577/3478062
9662:
9657:
9652:
9648:
9644:
9640:
9633:
9625:
9621:
9616:
9611:
9606:
9601:
9597:
9593:
9589:
9585:
9581:
9574:
9566:
9562:
9557:
9552:
9548:
9544:
9540:
9536:
9532:
9525:
9517:
9513:
9508:
9503:
9499:
9495:
9492:(5): 628–33.
9491:
9487:
9483:
9476:
9468:
9464:
9460:
9456:
9452:
9448:
9445:(5): 609–13.
9444:
9440:
9432:
9424:
9420:
9415:
9410:
9406:
9402:
9398:
9394:
9390:
9383:
9375:
9371:
9366:
9361:
9356:
9351:
9347:
9343:
9340:(6): e38365.
9339:
9335:
9331:
9324:
9316:
9312:
9307:
9302:
9298:
9294:
9290:
9286:
9282:
9275:
9267:
9263:
9258:
9253:
9249:
9245:
9241:
9237:
9233:
9226:
9218:
9214:
9209:
9204:
9200:
9196:
9193:(1): 91–105.
9192:
9188:
9184:
9177:
9169:
9165:
9160:
9155:
9150:
9145:
9141:
9137:
9133:
9126:
9118:
9114:
9110:
9106:
9102:
9098:
9091:
9083:
9079:
9074:
9069:
9065:
9061:
9057:
9053:
9049:
9042:
9034:
9030:
9025:
9020:
9016:
9012:
9008:
9004:
9000:
8993:
8985:
8981:
8977:
8973:
8968:
8963:
8959:
8955:
8951:
8947:
8943:
8936:
8928:
8924:
8919:
8914:
8910:
8906:
8903:(1): 609–21.
8902:
8898:
8894:
8887:
8875:
8867:
8860:
8856:
8852:
8848:
8844:
8837:
8833:
8828:
8823:
8818:
8813:
8809:
8805:
8801:
8794:
8786:
8782:
8777:
8772:
8768:
8764:
8761:(3): 544–50.
8760:
8756:
8752:
8745:
8737:
8733:
8728:
8723:
8719:
8715:
8712:(2): 259–63.
8711:
8707:
8703:
8696:
8688:
8684:
8679:
8674:
8670:
8666:
8662:
8658:
8654:
8647:
8639:
8635:
8630:
8625:
8620:
8615:
8611:
8607:
8603:
8596:
8588:
8584:
8579:
8574:
8570:
8566:
8562:
8558:
8554:
8547:
8539:
8535:
8530:
8525:
8521:
8517:
8513:
8509:
8505:
8498:
8490:
8486:
8482:
8478:
8474:
8470:
8467:(4): 563–78.
8466:
8462:
8455:
8447:
8443:
8438:
8433:
8429:
8425:
8421:
8417:
8413:
8406:
8390:
8386:
8382:
8378:
8371:
8363:
8359:
8354:
8349:
8344:
8339:
8335:
8331:
8327:
8320:
8312:
8308:
8303:
8298:
8294:
8290:
8286:
8282:
8278:
8271:
8263:
8259:
8254:
8249:
8245:
8241:
8238:(2): 431–42.
8237:
8233:
8229:
8222:
8214:
8210:
8205:
8200:
8196:
8192:
8188:
8184:
8180:
8173:
8165:
8161:
8157:
8153:
8149:
8145:
8142:(2): 139–46.
8141:
8137:
8133:
8126:
8118:
8114:
8110:
8106:
8102:
8098:
8095:(6): 488–95.
8094:
8090:
8083:
8075:
8071:
8066:
8061:
8057:
8053:
8049:
8045:
8041:
8034:
8026:
8022:
8017:
8012:
8008:
8004:
8000:
7993:
7985:
7981:
7977:
7973:
7969:
7965:
7961:
7957:
7950:
7942:
7938:
7933:
7928:
7924:
7920:
7916:
7912:
7908:
7901:
7899:
7897:
7888:
7884:
7879:
7874:
7869:
7864:
7860:
7856:
7852:
7848:
7844:
7837:
7835:
7826:
7822:
7818:
7814:
7810:
7806:
7803:(6): 627–37.
7802:
7798:
7791:
7783:
7779:
7775:
7771:
7767:
7763:
7759:
7755:
7748:
7746:
7737:
7733:
7728:
7723:
7718:
7713:
7709:
7705:
7701:
7694:
7686:
7682:
7677:
7672:
7668:
7664:
7660:
7656:
7652:
7645:
7637:
7633:
7629:
7625:
7620:
7615:
7612:(7): 736–47.
7611:
7607:
7603:
7596:
7594:
7592:
7590:
7581:
7577:
7573:
7569:
7566:(4): 209–18.
7565:
7561:
7553:
7545:
7541:
7537:
7533:
7529:
7525:
7522:(2): 93–103.
7521:
7517:
7510:
7502:
7498:
7494:
7490:
7485:
7480:
7477:(2): 327–35.
7476:
7472:
7465:
7457:
7453:
7448:
7443:
7439:
7435:
7431:
7427:
7423:
7416:
7406:
7402:
7398:
7397:
7394:
7390:
7386:
7382:
7377:
7372:
7367:
7362:
7358:
7354:
7350:
7346:
7342:
7335:
7327:
7323:
7318:
7313:
7309:
7305:
7301:
7297:
7293:
7286:
7278:
7274:
7269:
7264:
7260:
7256:
7252:
7245:
7237:
7233:
7228:
7223:
7219:
7215:
7211:
7207:
7203:
7196:
7194:
7185:
7181:
7176:
7171:
7167:
7163:
7159:
7155:
7151:
7144:
7132:
7124:
7118:
7113:
7112:
7107:
7103:
7099:
7095:
7088:
7084:
7079:
7074:
7069:
7064:
7060:
7056:
7052:
7048:
7044:
7037:
7029:
7023:
7019:
7018:
7013:
7006:
6998:
6994:
6990:
6986:
6982:
6978:
6974:
6970:
6963:
6955:
6951:
6946:
6941:
6937:
6933:
6929:
6922:
6914:
6910:
6905:
6900:
6896:
6892:
6888:
6884:
6880:
6873:
6865:
6861:
6856:
6851:
6846:
6841:
6837:
6833:
6829:
6822:
6814:
6810:
6805:
6800:
6796:
6792:
6788:
6784:
6780:
6776:
6772:
6765:
6757:
6753:
6748:
6743:
6739:
6735:
6731:
6727:
6723:
6719:
6715:
6708:
6700:
6696:
6691:
6686:
6682:
6678:
6674:
6670:
6666:
6659:
6651:
6647:
6643:
6639:
6634:
6629:
6625:
6621:
6618:(4): 545–62.
6617:
6613:
6609:
6602:
6594:
6590:
6586:
6582:
6578:
6574:
6567:
6559:
6555:
6550:
6545:
6541:
6537:
6533:
6529:
6525:
6518:
6516:
6507:
6503:
6498:
6493:
6489:
6485:
6481:
6474:
6466:
6462:
6457:
6452:
6447:
6442:
6438:
6434:
6430:
6423:
6415:
6411:
6406:
6401:
6397:
6393:
6389:
6385:
6381:
6374:
6372:
6370:
6361:
6357:
6353:
6349:
6345:
6341:
6337:
6333:
6329:
6325:
6318:
6310:
6306:
6301:
6296:
6292:
6288:
6284:
6280:
6276:
6269:
6267:
6265:
6256:
6252:
6247:
6242:
6239:(5): 623–34.
6238:
6234:
6230:
6223:
6215:
6211:
6206:
6201:
6196:
6191:
6187:
6183:
6179:
6175:
6171:
6164:
6156:
6152:
6147:
6142:
6138:
6134:
6130:
6126:
6122:
6115:
6107:
6103:
6098:
6093:
6088:
6083:
6079:
6075:
6071:
6067:
6063:
6056:
6048:
6044:
6039:
6034:
6029:
6024:
6020:
6016:
6012:
6005:
6003:
5994:
5990:
5985:
5980:
5976:
5972:
5968:
5964:
5960:
5953:
5945:
5941:
5936:
5931:
5927:
5923:
5919:
5915:
5911:
5904:
5896:
5892:
5887:
5882:
5878:
5874:
5870:
5863:
5855:
5851:
5846:
5841:
5838:(2): 209–16.
5837:
5833:
5829:
5822:
5814:
5810:
5805:
5800:
5796:
5792:
5788:
5781:
5773:
5769:
5764:
5759:
5755:
5751:
5747:
5740:
5738:
5729:
5725:
5721:
5717:
5713:
5709:
5705:
5701:
5694:
5686:
5682:
5677:
5672:
5668:
5664:
5660:
5656:
5652:
5645:
5637:
5633:
5629:
5625:
5621:
5617:
5610:
5608:
5598:
5593:
5589:
5585:
5581:
5574:
5566:
5562:
5557:
5552:
5549:(2): 262–72.
5548:
5544:
5540:
5533:
5525:
5521:
5517:
5511:
5507:
5503:
5499:
5492:
5484:
5480:
5475:
5470:
5466:
5462:
5458:
5454:
5450:
5443:
5435:
5431:
5426:
5421:
5417:
5413:
5409:
5402:
5400:
5398:
5389:
5382:
5378:
5374:
5370:
5366:
5362:
5358:
5354:
5348:
5346:
5337:
5333:
5329:
5325:
5321:
5317:
5310:
5302:
5298:
5294:
5290:
5286:
5282:
5279:(3): 228–34.
5278:
5274:
5267:
5259:
5255:
5250:
5245:
5241:
5237:
5233:
5229:
5225:
5218:
5216:
5207:
5203:
5198:
5193:
5189:
5185:
5181:
5177:
5173:
5166:
5158:
5154:
5149:
5144:
5140:
5136:
5132:
5128:
5124:
5120:
5116:
5109:
5101:
5097:
5092:
5087:
5083:
5079:
5075:
5071:
5067:
5063:
5059:
5052:
5044:
5040:
5036:
5032:
5027:
5022:
5018:
5014:
5010:
5003:
4995:
4991:
4986:
4981:
4977:
4973:
4970:(4): 844–58.
4969:
4965:
4961:
4954:
4946:
4942:
4937:
4932:
4929:(2): 542–54.
4928:
4924:
4920:
4913:
4905:
4901:
4896:
4891:
4887:
4883:
4879:
4872:
4864:
4860:
4855:
4850:
4846:
4842:
4838:
4834:
4830:
4823:
4815:
4811:
4807:
4801:
4797:
4793:
4789:
4782:
4774:
4770:
4766:
4762:
4758:
4754:
4750:
4746:
4742:
4738:
4731:
4723:
4719:
4714:
4709:
4705:
4701:
4698:(11): 663–7.
4697:
4693:
4689:
4682:
4674:
4670:
4665:
4660:
4655:
4650:
4646:
4642:
4638:
4634:
4630:
4623:
4615:
4611:
4606:
4601:
4597:
4593:
4589:
4585:
4581:
4574:
4572:
4570:
4561:
4557:
4552:
4547:
4543:
4539:
4535:
4531:
4527:
4520:
4512:
4508:
4503:
4498:
4494:
4490:
4487:(3): 775–83.
4486:
4482:
4478:
4471:
4463:
4459:
4455:
4451:
4446:
4441:
4437:
4433:
4429:
4422:
4414:
4410:
4405:
4400:
4396:
4392:
4388:
4384:
4380:
4373:
4371:
4369:
4367:
4358:
4354:
4349:
4344:
4340:
4336:
4332:
4328:
4324:
4317:
4315:
4306:
4300:
4292:
4288:
4283:
4278:
4274:
4270:
4266:
4262:
4258:
4254:
4250:
4243:
4235:
4231:
4227:
4223:
4219:
4215:
4211:
4207:
4203:
4199:
4192:
4184:
4180:
4176:
4172:
4168:
4164:
4160:
4156:
4152:
4148:
4141:
4139:
4130:
4126:
4121:
4116:
4112:
4108:
4104:
4100:
4096:
4089:
4081:
4077:
4073:
4069:
4065:
4061:
4057:
4053:
4046:
4038:
4034:
4030:
4026:
4022:
4018:
4015:(6s): S8–13.
4014:
4010:
4003:
3995:
3991:
3986:
3981:
3977:
3973:
3969:
3965:
3961:
3954:
3946:
3942:
3937:
3932:
3929:(7): 787–98.
3928:
3924:
3920:
3913:
3911:
3909:
3900:
3896:
3891:
3886:
3881:
3876:
3872:
3868:
3864:
3857:
3849:
3845:
3840:
3835:
3830:
3825:
3821:
3817:
3813:
3806:
3798:
3794:
3789:
3784:
3780:
3776:
3772:
3768:
3764:
3757:
3749:
3745:
3740:
3735:
3731:
3727:
3723:
3719:
3715:
3708:
3700:
3696:
3692:
3686:
3682:
3678:
3674:
3667:
3659:
3655:
3650:
3645:
3642:(5): 855–62.
3641:
3637:
3633:
3626:
3618:
3614:
3609:
3604:
3599:
3594:
3590:
3586:
3585:F1000Research
3582:
3575:
3567:
3563:
3559:
3555:
3552:(1): 92–104.
3551:
3547:
3540:
3532:
3528:
3523:
3518:
3514:
3510:
3507:(4): 747–57.
3506:
3502:
3498:
3491:
3483:
3479:
3474:
3469:
3464:
3459:
3455:
3451:
3447:
3440:
3432:
3428:
3423:
3418:
3413:
3408:
3404:
3400:
3396:
3392:
3388:
3381:
3373:
3369:
3364:
3359:
3355:
3351:
3348:(3): 209–17.
3347:
3343:
3339:
3332:
3324:
3320:
3316:
3312:
3307:
3302:
3298:
3294:
3290:
3286:
3282:
3278:
3271:
3263:
3259:
3255:
3251:
3247:
3243:
3239:
3235:
3231:
3227:
3220:
3212:
3208:
3203:
3198:
3195:(3): 327–32.
3194:
3190:
3186:
3179:
3171:
3167:
3162:
3157:
3153:
3149:
3145:
3138:
3130:
3126:
3122:
3118:
3114:
3110:
3103:
3101:
3092:
3088:
3084:
3080:
3076:
3072:
3068:
3064:
3060:
3056:
3049:
3041:
3037:
3033:
3029:
3025:
3021:
3017:
3013:
3009:
3005:
2998:
2996:
2994:
2985:
2981:
2977:
2973:
2969:
2965:
2961:
2957:
2953:
2949:
2942:
2940:
2938:
2929:
2925:
2921:
2917:
2912:
2907:
2903:
2899:
2895:
2891:
2887:
2883:
2876:
2874:
2872:
2863:
2859:
2855:
2851:
2847:
2843:
2839:
2835:
2831:
2827:
2820:
2818:
2809:
2805:
2801:
2797:
2793:
2789:
2785:
2781:
2777:
2773:
2766:
2764:
2755:
2751:
2746:
2741:
2738:(5): 843–54.
2737:
2733:
2729:
2722:
2720:
2718:
2709:
2705:
2700:
2695:
2691:
2687:
2683:
2679:
2675:
2668:
2660:
2656:
2651:
2646:
2642:
2638:
2635:(1): 92–105.
2634:
2630:
2626:
2619:
2617:
2615:
2606:
2602:
2597:
2592:
2588:
2584:
2580:
2573:
2571:
2569:
2560:
2556:
2551:
2546:
2542:
2538:
2534:
2530:
2526:
2519:
2517:
2508:
2504:
2499:
2494:
2490:
2486:
2482:
2478:
2474:
2467:
2459:
2455:
2450:
2445:
2441:
2437:
2433:
2429:
2425:
2418:
2410:
2406:
2401:
2396:
2392:
2388:
2384:
2380:
2376:
2369:
2363:
2359:
2357:
2351:
2343:
2339:
2334:
2329:
2324:
2319:
2315:
2311:
2307:
2303:
2299:
2295:
2291:
2284:
2276:
2272:
2268:
2264:
2260:
2256:
2252:
2248:
2241:
2233:
2229:
2225:
2221:
2217:
2213:
2209:
2205:
2198:
2190:
2186:
2181:
2176:
2172:
2168:
2165:(2): 215–33.
2164:
2160:
2156:
2149:
2147:
2145:
2136:
2132:
2127:
2122:
2118:
2114:
2110:
2106:
2102:
2095:
2087:
2083:
2078:
2073:
2069:
2065:
2061:
2054:
2052:
2050:
2041:
2037:
2032:
2027:
2023:
2019:
2015:
2011:
2007:
2000:
1998:
1996:
1994:
1989:
1980:
1977:
1975:
1972:
1970:
1967:
1965:
1962:
1960:
1957:
1955:
1952:
1950:
1947:
1945:
1942:
1940:
1937:
1935:
1932:
1930:
1927:
1925:
1922:
1920:
1917:
1915:
1912:
1911:
1907:
1901:
1896:
1889:
1887:
1880:
1870:
1868:
1867:
1855:
1846:
1837:
1835:
1831:
1827:
1823:
1819:
1815:
1811:
1807:
1802:
1800:
1796:
1792:
1788:
1784:
1780:
1776:
1772:
1768:
1764:
1760:
1756:
1752:
1748:
1744:
1734:
1732:
1728:
1724:
1720:
1716:
1712:
1708:
1704:
1700:
1696:
1692:
1688:
1684:
1680:
1676:
1671:
1669:
1664:
1660:
1656:
1652:
1651:cell adhesion
1648:
1644:
1640:
1636:
1632:
1628:
1618:
1616:
1606:
1604:
1600:
1596:
1592:
1591:schizophrenia
1588:
1584:
1580:
1576:
1572:
1568:
1564:
1560:
1556:
1552:
1542:
1540:
1536:
1532:
1528:
1524:
1520:
1516:
1512:
1508:
1504:
1503:transcriptome
1500:
1496:
1492:
1488:
1484:
1474:
1471:
1462:
1458:
1454:
1451:
1447:
1443:
1433:
1431:
1428:
1418:
1416:
1412:
1408:
1399:
1397:
1392:
1385:Heart disease
1382:
1379:
1377:
1373:
1369:
1364:
1360:
1358:
1354:
1350:
1345:
1341:
1340:glioblastomas
1338:In 29–66% of
1336:
1334:
1330:
1326:
1322:
1318:
1314:
1310:
1306:
1302:
1300:
1297:. Defects in
1296:
1292:
1288:
1284:
1274:
1270:
1268:
1264:
1260:
1254:
1250:
1248:
1244:
1240:
1237:samples, low
1236:
1231:
1229:
1225:
1221:
1212:
1203:
1200:
1197:
1189:
1180:
1178:
1174:
1170:
1166:
1161:
1159:
1155:
1151:
1147:
1143:
1139:
1135:
1131:
1127:
1122:
1120:
1116:
1105:
1101:
1099:
1095:
1094:
1089:
1085:
1081:
1080:
1074:
1072:
1068:
1063:
1061:
1057:
1056:
1050:
1045:
1042:
1037:
1035:
1031:
1027:
1017:
1015:
1011:
1006:
1003:
999:
995:
992:
991:extracellular
987:
985:
981:
976:
972:
967:
965:
961:
957:
952:
950:
944:
942:
934:
931:
928:
925:
921:
918:
915:
912:
909:
906:
905:
904:
901:
899:
895:
891:
886:
884:
880:
876:
872:
868:
867:deadenylation
862:
859:
855:
851:
850:complementary
843:
837:
828:
826:
822:
817:
813:
809:
808:
802:
800:
797:
793:
792:
786:
777:
774:
770:
766:
755:
753:
749:
746:
742:
738:
734:
730:
725:
722:
718:
714:
705:
701:
697:
692:
682:
680:
679:Hua-Enhancer1
676:
672:
661:
657:
655:
651:
647:
643:
639:
635:
625:
623:
619:
615:
611:
602:
598:
593:
589:
584:
575:
573:
569:
565:
561:
557:
552:
550:
546:
542:
538:
533:
531:
527:
523:
519:
514:
506:
502:
497:
493:
489:
485:
482:
478:
475:of the human
474:
469:
460:
458:
455:(tRNAs), and
454:
453:transfer RNAs
450:
449:Alu sequences
446:
442:
438:
434:
430:
426:
422:
415:Transcription
412:
410:
405:
403:
399:
391:
382:
378:
369:
367:
361:
359:
355:
351:
346:
333:
324:
323:
322:
319:
317:
313:
309:
306:
302:
298:
287:
286:
282:
277:
274:
270:
266:
265:
260:
256:
252:
247:
245:
241:
237:
233:
229:
228:
222:
220:
216:
212:
208:
204:
200:
196:
192:
188:
187:
182:
181:
176:
172:
167:
164:
153:
149:
147:
143:
139:
135:
131:
126:
120:
116:
113:
109:
106:
105:
104:
102:
98:
94:
90:
86:
82:
81:RNA silencing
78:
74:
70:
66:
58:
52:
47:
43:
39:
38:messenger RNA
35:
30:
19:
15712:Bacterial
15687:Lambda phage
15457:
15351:Constituents
15301:List of RNAs
15192:
15111:Transfer RNA
13791:
13757:
13666:
13662:
13652:
13620:
13616:
13608:
13578:(3): 277–9.
13575:
13571:
13519:
13515:
13505:
13468:
13464:
13454:
13419:
13415:
13405:
13396:
13386:
13351:
13347:
13337:
13302:
13298:
13292:
13255:
13251:
13241:
13200:
13196:
13190:
13153:
13149:
13139:
13102:
13098:
13052:
13048:
13042:
13015:
13011:
13001:
12974:
12970:
12930:
12926:
12916:
12871:
12867:
12857:
12825:(1): 81–94.
12822:
12818:
12808:
12775:
12771:
12765:
12738:
12734:
12724:
12689:
12685:
12678:
12645:
12641:
12635:
12600:
12596:
12538:
12534:
12524:
12489:
12485:
12475:
12430:
12426:
12416:
12381:
12377:
12367:
12332:
12328:
12278:
12274:
12264:
12229:
12225:
12215:
12203:. Retrieved
12198:
12189:
12152:
12148:
12138:
12094:(7): e6121.
12091:
12087:
12077:
12038:(1): 79–97.
12035:
12031:
12018:
11991:
11987:
11977:
11940:
11936:
11926:
11899:
11895:
11885:
11840:
11836:
11826:
11781:
11777:
11767:
11734:
11730:
11724:
11687:
11683:
11673:
11636:
11632:
11622:
11587:
11583:
11529:
11525:
11519:
11482:
11478:
11422:
11418:
11350:
11346:
11340:
11297:
11293:
11283:
11242:
11238:
11232:
11202:(5): 613–8.
11199:
11195:
11189:
11156:
11152:
11146:
11132:{{
11124:{{
11117:
11115:,
11065:
11061:
11051:
11010:
11006:
11000:
10965:
10961:
10951:
10906:
10902:
10892:
10865:
10861:
10851:
10824:
10820:
10762:
10758:
10751:Murchison EP
10744:
10717:
10713:
10703:
10666:
10662:
10652:
10617:
10613:
10603:
10576:
10572:
10562:
10532:(1–3): 1–8.
10529:
10526:FEBS Letters
10525:
10519:
10487:(1): 28–36.
10484:
10480:
10470:
10438:(6): 712–9.
10435:
10431:
10371:
10367:
10357:
10330:
10326:
10316:
10281:
10277:
10267:
10232:
10228:
10218:
10199:
10193:
10166:
10162:
10152:
10142:15 September
10140:. Retrieved
10106:
10102:
10089:
10054:
10050:
10040:
10005:
10001:
9991:
9956:
9952:
9942:
9915:
9911:
9901:
9892:
9847:
9843:
9806:(2): 311–8.
9803:
9799:
9789:
9756:
9752:
9742:
9708:(16): 3976.
9705:
9701:
9691:
9646:
9642:
9632:
9587:
9583:
9573:
9538:
9534:
9524:
9489:
9485:
9475:
9442:
9438:
9431:
9396:
9392:
9382:
9337:
9333:
9323:
9288:
9284:
9274:
9239:
9235:
9225:
9190:
9186:
9176:
9139:
9135:
9125:
9100:
9097:Biochemistry
9096:
9090:
9058:(6): 566–9.
9055:
9052:Biology Open
9051:
9041:
9006:
9002:
8992:
8949:
8945:
8935:
8900:
8896:
8886:
8872:{{
8864:{{
8807:
8804:PLOS Biology
8803:
8793:
8758:
8754:
8744:
8709:
8705:
8695:
8660:
8656:
8646:
8609:
8606:BMC Genomics
8605:
8595:
8560:
8556:
8546:
8514:(20): e179.
8511:
8507:
8497:
8464:
8460:
8454:
8419:
8415:
8405:
8393:. Retrieved
8389:the original
8384:
8380:
8370:
8333:
8330:BMC Genomics
8329:
8319:
8284:
8280:
8270:
8235:
8231:
8221:
8186:
8182:
8172:
8139:
8135:
8125:
8092:
8088:
8082:
8047:
8043:
8033:
8006:
8002:
7992:
7959:
7955:
7949:
7914:
7910:
7850:
7846:
7800:
7796:
7790:
7760:(1): 50–68.
7757:
7753:
7707:
7703:
7693:
7658:
7654:
7644:
7609:
7605:
7563:
7559:
7552:
7519:
7515:
7509:
7474:
7470:
7464:
7429:
7425:
7415:
7405:Science News
7404:
7348:
7344:
7334:
7299:
7296:eBioMedicine
7295:
7285:
7258:
7254:
7244:
7209:
7205:
7160:(3): 353–8.
7157:
7153:
7143:
7129:{{
7121:{{
7110:
7108:,
7050:
7046:
7036:
7016:
7005:
6972:
6968:
6962:
6935:
6931:
6921:
6889:(5): 602–7.
6886:
6882:
6872:
6835:
6831:
6821:
6778:
6774:
6764:
6721:
6717:
6707:
6675:(1): 21–32.
6672:
6668:
6658:
6615:
6611:
6601:
6576:
6572:
6566:
6531:
6527:
6490:(1): 211–2.
6487:
6483:
6473:
6436:
6432:
6422:
6387:
6383:
6327:
6323:
6317:
6282:
6278:
6236:
6232:
6222:
6177:
6173:
6163:
6131:(6): 720–8.
6128:
6124:
6114:
6072:(2): 512–7.
6069:
6065:
6055:
6018:
6014:
5966:
5962:
5952:
5917:
5913:
5903:
5876:
5872:
5862:
5835:
5831:
5821:
5794:
5790:
5780:
5753:
5749:
5706:(1): 23–36.
5703:
5699:
5693:
5661:(1): 14–23.
5658:
5654:
5644:
5622:(3): 616–7.
5619:
5615:
5587:
5583:
5573:
5546:
5542:
5532:
5497:
5491:
5456:
5452:
5442:
5415:
5411:
5367:(3): 223–9.
5364:
5360:
5353:Murchison EP
5319:
5315:
5309:
5276:
5272:
5266:
5231:
5227:
5179:
5175:
5165:
5122:
5118:
5108:
5065:
5061:
5051:
5016:
5013:FEBS Letters
5012:
5002:
4967:
4963:
4953:
4926:
4923:Cell Reports
4922:
4912:
4885:
4881:
4871:
4836:
4832:
4822:
4787:
4781:
4740:
4736:
4730:
4695:
4691:
4681:
4636:
4632:
4622:
4587:
4583:
4536:(3): 241–7.
4533:
4529:
4519:
4484:
4480:
4470:
4438:(1): 59–73.
4435:
4431:
4421:
4386:
4382:
4330:
4326:
4299:cite journal
4256:
4252:
4242:
4201:
4197:
4191:
4150:
4146:
4102:
4098:
4088:
4055:
4051:
4045:
4012:
4008:
4002:
3967:
3963:
3953:
3926:
3922:
3873:(1): 19–53.
3870:
3867:BMC Genomics
3866:
3856:
3819:
3815:
3805:
3770:
3766:
3756:
3724:(3): 277–9.
3721:
3717:
3707:
3672:
3666:
3639:
3635:
3625:
3588:
3584:
3574:
3549:
3545:
3539:
3504:
3500:
3490:
3453:
3449:
3439:
3394:
3390:
3380:
3345:
3341:
3331:
3280:
3276:
3270:
3229:
3225:
3219:
3192:
3188:
3178:
3154:(1): 25–36.
3151:
3147:
3137:
3112:
3108:
3058:
3054:
3048:
3007:
3003:
2951:
2947:
2885:
2881:
2829:
2825:
2775:
2771:
2735:
2731:
2681:
2677:
2667:
2632:
2628:
2589:(1): 15–20.
2586:
2582:
2535:(9): 735–9.
2532:
2528:
2480:
2476:
2466:
2431:
2427:
2417:
2382:
2378:
2368:
2356:Homo sapiens
2355:
2350:
2323:1721.1/72447
2297:
2293:
2283:
2250:
2246:
2240:
2207:
2203:
2197:
2162:
2158:
2108:
2104:
2094:
2067:
2063:
2016:(1): 20–51.
2013:
2009:
1885:
1882:
1864:
1861:
1852:
1843:
1803:
1763:stromal cell
1755:adipogenesis
1740:
1672:
1624:
1612:
1548:
1480:
1472:
1468:
1459:
1455:
1439:
1430:
1424:
1405:
1388:
1380:
1368:polypeptides
1361:
1337:
1309:colon cancer
1303:
1280:
1271:
1255:
1251:
1232:
1217:
1201:
1198:
1195:
1186:
1162:
1132:followed by
1123:
1111:
1102:
1091:
1077:
1075:
1064:
1053:
1046:
1041:phylogenetic
1038:
1023:
1007:
993:
988:
968:
953:
948:
945:
940:
938:
902:
887:
878:
863:
847:
805:
803:
789:
787:
783:
773:feed-forward
761:
726:
710:
698:
694:
667:
658:
646:G-quadruplex
642:G-quadruplex
631:
607:
553:
548:
544:
534:
515:
511:
418:
406:
395:
379:
375:
362:
357:
353:
350:Homo sapiens
349:
347:
343:
335:hsa-miR-181b
326:hsa-miR-181a
320:
315:
311:
307:
304:
300:
296:
293:
290:Nomenclature
278:
272:
268:
262:
258:
254:
250:
248:
243:
239:
235:
231:
225:
223:
214:
210:
206:
194:
190:
184:
178:
168:
159:
150:
142:human genome
127:
124:
112:poly(A) tail
68:
64:
63:
42:
29:
15707:P1-derived
15475:Antisense
15368:Nucleotides
15363:Nucleosides
15358:Nucleobases
12707:10852/40423
11300:(1): 4511.
10862:Circulation
8845:, see
8810:(8): e203.
6285:(1): 5–10.
3822:(2): 90–8.
3306:1721.1/7483
1759:bone marrow
1355:3'UTR (the
1138:microarrays
1071:brown algae
973:that share
971:pseudogenes
825:hepatitis C
671:Dicer-like1
556:RNA editing
305:Drosophila,
119:translation
77:nucleotides
15757:Categories
15659:Morpholino
15572:Organellar
15480:Processual
15445:Regulatory
15389:non-coding
15239:Riboswitch
15183:CRISPR RNA
13705:miRTarBase
13522:: e05005.
13393:"VIRmiRNA"
13354:: bau103.
13258:(3): 319.
13156:(1): 120.
13018:: 107676.
12205:5 December
11943:(3): 425.
11639:(6): 842.
9009:(8): e60.
8336:(1): 714.
7710:(4): 221.
7261:: 101732.
6883:Cell Cycle
6439:(9): R65.
5963:Cell Cycle
4639:(3): e37.
2684:: 213–42.
2111:: bau103.
1985:References
1954:miR-324-5p
1877:See also:
1840:Hemostasis
1830:antagomirs
1747:adipocytes
1725:and delta
1643:cell cycle
1639:withdrawal
1631:alcoholism
1621:Alcoholism
1495:arterioles
1299:DNA repair
1283:cell cycle
1243:metastasis
1154:Morpholino
1060:arthropods
1002:biomarkers
879:Drosophila
733:interferon
610:Exportin-5
592:nucleotide
549:C. elegans
545:Drosophila
481:C-terminal
433:adenosines
385:Biogenesis
358:Drosophila
354:Ovis aries
318:in human.
301:C. elegans
285:oncogenes.
264:Drosophila
259:C. elegans
236:C. elegans
186:C. elegans
15619:Analogues
15604:Hachimoji
15387:(coding,
15342:Types of
15296:Vault RNA
15270:RNA virus
15253:Parasites
15152:RNase MRP
15142:Guide RNA
15080:Types of
13952:46/47/281
13837:8/141/200
13391:Kumar M.
13233:233685660
13225:1336-9563
11134:retracted
11126:retracted
10749:Chen JF,
10137:249070106
9683:255034585
8089:BioEssays
7917:: 180–9.
7606:BioEssays
7479:CiteSeerX
7393:230713808
7115:. If the
6838:(1): 12.
5418:: 59–66.
5357:Hannon GJ
5316:Biochimie
5301:205286318
3115:: 19–53.
1886:in silico
1816:, is the
1761:-derived
1743:stem cell
1719:accumbens
1695:cytokines
1691:microglia
1647:apoptosis
1627:addiction
1571:forebrain
1539:apoptosis
1519:Trp53inp1
1499:mesangial
1436:Mechanism
1402:miRNA-712
1291:mutations
1158:antagomir
1026:conserved
1020:Evolution
865:speed up
765:TNF alpha
735:-induced
713:Argonaute
650:cytoplasm
634:cytoplasm
624:protein.
568:adenosine
562:known as
441:stem-loop
146:MirGeneDB
117:Reducing
89:base-pair
87:. miRNAs
15763:MicroRNA
15742:Category
15677:Phagemid
15528:Ribozyme
15193:MicroRNA
13726:Archived
13645:82531727
13637:12242426
13602:12592000
13548:26267216
13497:23591837
13446:22114334
13378:25380780
13348:Database
13329:21504877
13284:32192095
13197:Biologia
13182:22838835
13131:36499082
13069:22886416
13034:32898547
12993:30207063
12949:25400249
12908:22160727
12849:21962509
12792:26811207
12757:16431920
12716:21756067
12670:30646787
12662:21844119
12627:23873704
12575:23951048
12535:PLOS ONE
12516:24672003
12467:24358208
12427:PLOS ONE
12408:22614244
12359:21651580
12305:23170800
12256:20841499
12181:19721432
12130:19568434
12088:PLOS ONE
12066:Archived
12061:25088900
12052:25395183
12010:27839653
11969:35327979
11918:20660113
11877:18385371
11818:36656826
11778:PLOS ONE
11751:19888283
11716:22661924
11665:27240359
11614:26438731
11554:12490960
11511:23144462
11457:24346612
11425:: 3000.
11367:19110086
11332:28674420
11267:17379774
11224:10097893
11216:17468766
11173:17401374
11113:21961173
11094:17344217
11035:15951802
10992:17498736
10943:17108080
10884:17606841
10843:17397913
10799:18256189
10736:25313246
10695:22494821
10669:(1): 3.
10644:14627817
10595:15150086
10554:28903539
10546:15358530
10511:20150365
10462:22570426
10408:20351277
10349:15887099
10308:20420945
10259:21183529
10185:10688858
10129:35623329
10081:20603801
10073:24608427
10051:Oncogene
10032:24604117
9975:29521426
9934:24222179
9884:27271572
9844:PLOS ONE
9822:21573504
9773:21748820
9734:36010971
9675:36541726
9624:15284443
9565:21892160
9516:21996275
9467:11113852
9459:19363479
9423:18927107
9374:22723856
9334:PLOS ONE
9315:19420067
9266:17991681
9217:17612493
9168:19689821
9117:16752924
9082:23213449
9033:16670427
8984:30461594
8976:17761850
8927:24068553
8859:35486652
8836:17676975
8785:14970398
8736:17220889
8687:15585662
8638:21171994
8587:16043497
8538:16314309
8481:18388938
8446:16717284
8362:23256903
8311:24337300
8262:21317375
8213:20520714
8164:26319627
8117:20548122
8109:20486135
8074:20407027
8025:18296705
7984:11997028
7976:15565108
7941:20624724
7887:18287013
7825:12025420
7817:23980889
7782:14924603
7774:19196333
7736:21554756
7685:18715673
7636:15364875
7628:19472371
7580:17465887
7536:17230196
7501:15136036
7456:15849273
7385:33443164
7326:26629549
7277:31816349
7236:27264337
7184:21802130
7106:29555737
7087:18227514
6989:15211354
6954:16337999
6913:18256543
6864:19232136
6813:22499947
6756:22422859
6699:19029310
6642:17965831
6633:11131689
6593:17532529
6558:18653800
6506:17150553
6465:15345049
6414:22850425
6352:19734881
6309:20051982
6255:15766526
6214:16289642
6155:11914277
6106:18178619
6047:19342379
5993:18769156
5944:16005165
5920:: 32–8.
5895:14567917
5854:14567918
5813:15292246
5772:12000786
5720:17183358
5685:20808519
5636:24508494
5565:25641166
5524:18268841
5483:21753850
5434:17381281
5381:15145345
5336:17628290
5293:19255566
5258:18684997
5206:17964270
5157:17589500
5100:17589500
5043:28899778
5035:23063642
4994:23415231
4945:25310978
4904:16751099
4863:15574589
4814:16957365
4765:14508493
4722:18778799
4673:17352530
4614:15372072
4560:15701730
4511:17255951
4462:32923462
4454:15634332
4413:15525708
4357:15364901
4291:18668037
4226:18668040
4175:15685193
4129:21652644
4080:22672750
4072:15806104
4037:23496396
4029:16736023
3994:21441577
3945:14697198
3899:31345162
3848:21296742
3797:16381832
3748:12592000
3699:26658994
3617:24627783
3566:19896977
3531:19876744
3482:19956180
3473:10033140
3450:Oncogene
3431:16040801
3372:17521938
3315:14657504
3254:15538371
3211:16423811
3170:12679032
3129:16669754
3091:38939571
3083:15919954
3040:33480585
3032:11679672
2984:43262684
2976:11679671
2928:18101169
2920:11679670
2854:11081512
2800:10706289
2708:26473382
2659:18955434
2605:15652477
2559:12007417
2507:12672692
2458:31598695
2409:30820533
2342:20703300
2275:24892348
2267:26077373
2232:24892348
2224:26077373
2189:19167326
2135:25380780
2105:Database
2086:14744438
2040:29570994
1934:MicroDNA
1892:See also
1826:diabetes
1814:diabetes
1753:play in
1587:epilepsy
1567:striatal
1446:collagen
1372:AT hooks
1319:protein
1305:Germline
1224:oncomirs
1098:Placozoa
924:P-bodies
898:promoter
780:Turnover
745:helicase
604:.
541:mirtrons
508:.
400:or even
219:nematode
65:MicroRNA
15682:Plasmid
15147:RNase P
15024:BHRF1-3
15019:BHRF1-2
15014:BHRF1-1
14660:501-600
14524:401-500
14363:301-400
14237:201-300
14198:192/215
14113:148/152
14023:103/107
14011:101-200
13717:ONCO.IO
13711:semirna
13685:8252621
13617:Science
13610:Science
13593:1370393
13539:4532895
13488:3645737
13437:3264354
13369:4224276
13275:7140873
13205:Bibcode
13173:3443071
13122:9740008
13077:6402014
12899:3248488
12876:Bibcode
12840:3353524
12800:3614126
12618:3799494
12566:3739772
12543:Bibcode
12507:3965783
12458:3865091
12435:Bibcode
12399:3546132
12350:3170679
12296:3673881
12247:3110894
12172:2990188
12121:2699475
12096:Bibcode
11960:8951370
11868:2291142
11845:Bibcode
11809:9851566
11786:Bibcode
11759:3507952
11707:3356852
11656:4926376
11605:4632944
11534:Bibcode
11502:3531724
11448:3923891
11427:Bibcode
11323:5495799
11302:Bibcode
11275:1927839
11247:Bibcode
11239:Science
11181:1935811
11085:3151106
11043:4340449
11015:Bibcode
10983:1934409
10934:1838739
10911:Bibcode
10790:2542870
10767:Bibcode
10686:3351028
10502:2940563
10453:3367855
10399:2872463
10376:Bibcode
10299:3057468
10250:3104011
10023:4144705
9983:3766576
9875:4894642
9852:Bibcode
9781:9746594
9725:9406077
9702:Cancers
9592:Bibcode
9556:3184212
9507:3213395
9414:2686559
9365:3378551
9342:Bibcode
9306:2703907
9257:2238936
9208:3800283
9159:2736963
9073:3509442
9024:1456327
8954:Bibcode
8946:Science
8918:3874169
8827:1925136
8776:1370948
8727:3982799
8629:3022920
8612:: 716.
8578:1370829
8529:1292995
8489:2441105
8437:1464411
8395:10 July
8353:3563456
8302:3920664
8281:Science
8253:3077775
8191:Bibcode
8144:Bibcode
8065:2879733
7932:2942034
7878:2268565
7855:Bibcode
7727:3218855
7676:2695246
7447:1143068
7376:7817116
7353:Bibcode
7317:4634783
7227:5343750
7175:3235919
7131:erratum
7123:erratum
7117:erratum
7078:2234192
7055:Bibcode
6997:5270062
6904:2877389
6855:2680403
6804:3971879
6783:Bibcode
6775:Science
6747:3547538
6726:Bibcode
6718:Science
6690:2612776
6650:5708394
6549:2556272
6405:3425779
6360:4414841
6332:Bibcode
6300:6417416
6182:Bibcode
6097:2206567
6074:Bibcode
6038:2709356
5984:2697033
5935:1788082
5728:8966239
5676:2851111
5474:4693635
5249:2532740
5197:2763384
5148:2475599
5127:Bibcode
5091:2475599
5070:Bibcode
4985:3707628
4773:4421030
4745:Bibcode
4713:2633599
4664:1817659
4641:Bibcode
4551:1370713
4502:1794378
4404:1370684
4282:2745094
4261:Bibcode
4234:4429008
4206:Bibcode
4183:4430576
4155:Bibcode
4120:3167600
3985:3087955
3890:6657096
3839:3051107
3788:1347474
3739:1370393
3658:8252622
3608:3917658
3591:: 136.
3522:2782079
3422:1182454
3399:Bibcode
3363:1978064
3323:7044929
3285:Bibcode
3277:Science
3262:4415988
3234:Bibcode
3063:Bibcode
3055:Science
3012:Bibcode
3004:Science
2956:Bibcode
2948:Science
2890:Bibcode
2882:Science
2862:4401732
2834:Bibcode
2808:4384503
2780:Bibcode
2754:8252621
2699:4743252
2650:2612969
2537:Bibcode
2449:6943042
2400:6468295
2333:2990499
2302:Bibcode
2180:3794896
2126:4224276
2031:6091663
1944:MiRNEST
1858:Viruses
1834:obesity
1810:obesity
1775:miR-222
1771:miR-221
1767:miR-155
1737:Obesity
1675:miR-206
1521:, Jun,
1507:Bcl2L11
1487:nephron
1470:TIMP3.
1450:elastin
1327:of the
1239:miR-324
1067:genomes
1034:animals
871:miR-430
821:miR-122
677:called
632:In the
588:Ran-GTP
572:inosine
560:enzymes
537:spliced
486:of two
484:helices
437:spliced
409:IsomiRs
398:introns
372:Targets
205:of the
163:tissues
156:History
132:of the
97:silence
40:(mRNA).
15697:Fosmid
15692:Cosmid
15642:Hexose
15564:acids
15516:Others
15275:Viroid
13683:
13643:
13635:
13600:
13590:
13546:
13536:
13495:
13485:
13444:
13434:
13376:
13366:
13327:
13282:
13272:
13231:
13223:
13180:
13170:
13129:
13119:
13075:
13067:
13032:
12991:
12947:
12906:
12896:
12847:
12837:
12798:
12790:
12755:
12714:
12668:
12660:
12625:
12615:
12573:
12563:
12514:
12504:
12465:
12455:
12406:
12396:
12357:
12347:
12303:
12293:
12254:
12244:
12179:
12169:
12128:
12118:
12058:
12050:
12008:
11967:
11957:
11916:
11875:
11865:
11816:
11806:
11757:
11749:
11714:
11704:
11690:: 71.
11663:
11653:
11612:
11602:
11562:407449
11560:
11552:
11526:Nature
11509:
11499:
11455:
11445:
11365:
11330:
11320:
11273:
11265:
11222:
11214:
11179:
11171:
11111:
11107:,
11092:
11082:
11041:
11033:
11007:Nature
10990:
10980:
10941:
10931:
10882:
10841:
10797:
10787:
10734:
10720:(11).
10693:
10683:
10642:
10635:290254
10632:
10593:
10552:
10544:
10509:
10499:
10460:
10450:
10406:
10396:
10347:
10306:
10296:
10257:
10247:
10206:
10183:
10135:
10127:
10109:(11).
10079:
10071:
10030:
10020:
9981:
9973:
9932:
9882:
9872:
9820:
9779:
9771:
9732:
9722:
9681:
9673:
9622:
9615:511048
9612:
9563:
9553:
9514:
9504:
9465:
9457:
9421:
9411:
9395:. 37.
9372:
9362:
9313:
9303:
9264:
9254:
9215:
9205:
9166:
9156:
9142:: 98.
9115:
9080:
9070:
9031:
9021:
8982:
8974:
8925:
8915:
8857:
8853:,
8834:
8824:
8783:
8773:
8734:
8724:
8685:
8678:535676
8675:
8636:
8626:
8585:
8575:
8536:
8526:
8487:
8479:
8444:
8434:
8360:
8350:
8309:
8299:
8260:
8250:
8211:
8183:Nature
8162:
8115:
8107:
8072:
8062:
8023:
7982:
7974:
7939:
7929:
7885:
7875:
7823:
7815:
7780:
7772:
7734:
7724:
7683:
7673:
7634:
7626:
7578:
7544:174231
7542:
7534:
7499:
7481:
7454:
7444:
7391:
7383:
7373:
7324:
7314:
7275:
7234:
7224:
7182:
7172:
7104:
7100:,
7085:
7075:
7024:
6995:
6987:
6952:
6911:
6901:
6862:
6852:
6811:
6801:
6754:
6744:
6697:
6687:
6648:
6640:
6630:
6591:
6556:
6546:
6504:
6463:
6456:522872
6453:
6412:
6402:
6358:
6350:
6324:Nature
6307:
6297:
6253:
6212:
6153:
6146:155365
6143:
6104:
6094:
6045:
6035:
5991:
5981:
5942:
5932:
5893:
5852:
5811:
5770:
5726:
5718:
5683:
5673:
5634:
5563:
5522:
5512:
5481:
5471:
5453:Nature
5432:
5379:
5334:
5299:
5291:
5256:
5246:
5204:
5194:
5155:
5145:
5119:Nature
5098:
5088:
5062:Nature
5041:
5033:
4992:
4982:
4943:
4902:
4861:
4854:535913
4851:
4812:
4802:
4771:
4763:
4737:Nature
4720:
4710:
4671:
4661:
4612:
4605:524334
4602:
4558:
4548:
4509:
4499:
4460:
4452:
4411:
4401:
4355:
4348:524413
4345:
4289:
4279:
4253:Nature
4232:
4224:
4198:Nature
4181:
4173:
4147:Nature
4127:
4117:
4078:
4070:
4035:
4027:
3992:
3982:
3943:
3897:
3887:
3846:
3836:
3795:
3785:
3746:
3736:
3697:
3687:
3656:
3615:
3605:
3564:
3529:
3519:
3480:
3470:
3429:
3419:
3370:
3360:
3321:
3313:
3260:
3252:
3226:Nature
3209:
3168:
3127:
3089:
3081:
3038:
3030:
2982:
2974:
2926:
2918:
2860:
2852:
2826:Nature
2806:
2798:
2772:Nature
2752:
2706:
2696:
2657:
2647:
2603:
2557:
2505:
2498:196042
2495:
2456:
2446:
2407:
2397:
2340:
2330:
2294:Nature
2273:
2265:
2230:
2222:
2187:
2177:
2133:
2123:
2084:
2038:
2028:
1949:MIR222
1849:Plants
1812:, and
1789:, and
1773:, and
1707:reward
1609:Stroke
1601:, and
1581:, and
1531:Arid3a
1529:, and
1523:Cdkn1a
1421:Origin
1407:Murine
1391:murine
1313:causal
1206:Cancer
1148:(LNA)
1130:RT-PCR
1115:RNases
1084:Drosha
1030:plants
980:ceRNAs
858:3' UTR
812:uracil
769:GM-CSF
752:TNRC6B
526:Drosha
477:Drosha
425:capped
276:RNAs.
238:. The
232:lin-41
211:lin-14
207:lin-14
203:3' UTR
191:lin-14
175:Ruvkun
171:Ambros
140:. The
18:MMChip
15722:Human
15500:Y RNA
15284:Other
15157:Y RNA
15039:Lsy-6
15034:Lin-4
15029:Let-7
15009:BART2
15004:BART1
14997:Other
13800:1-100
13792:miRNA
13723:MirOB
13653:lin-4
13641:S2CID
13516:eLife
13252:Genes
13229:S2CID
13099:Genes
13073:S2CID
12796:S2CID
12666:S2CID
12069:(PDF)
12056:S2CID
12028:(PDF)
11937:Genes
11755:S2CID
11558:S2CID
11271:S2CID
11220:S2CID
11177:S2CID
11130:with
11039:S2CID
10550:S2CID
10133:S2CID
10077:S2CID
9979:S2CID
9777:S2CID
9679:S2CID
9463:S2CID
8980:S2CID
8870:with
8485:S2CID
8113:S2CID
7980:S2CID
7821:S2CID
7778:S2CID
7632:S2CID
7540:S2CID
7389:S2CID
7127:with
6993:S2CID
6646:S2CID
6356:S2CID
5724:S2CID
5297:S2CID
5039:S2CID
4769:S2CID
4458:S2CID
4230:S2CID
4179:S2CID
4076:S2CID
4033:S2CID
3319:S2CID
3258:S2CID
3087:S2CID
3036:S2CID
2980:S2CID
2924:S2CID
2858:S2CID
2804:S2CID
2271:S2CID
2228:S2CID
1822:aging
1818:let-7
1799:CEBPA
1795:PPARγ
1563:Dicer
1491:renin
1483:FoxD1
1376:ERCC1
1235:NSCLC
1165:mRNAs
1150:oligo
1119:RNase
1088:Pasha
875:miR-9
748:MOV10
638:Dicer
488:DGCR8
402:exons
366:sense
316:MIR25
297:mir-1
273:let-7
269:lin-4
255:let-7
251:lin-4
244:let-7
240:let-7
227:let-7
215:lin-4
195:lin-4
180:lin-4
99:said
69:miRNA
14988:3180
14791:601+
14254:203a
13842:9/79
13681:PMID
13663:Cell
13633:PMID
13598:PMID
13544:PMID
13493:PMID
13442:PMID
13374:PMID
13352:2014
13325:PMID
13280:PMID
13221:ISSN
13178:PMID
13127:PMID
13065:PMID
13030:PMID
12989:PMID
12945:PMID
12904:PMID
12845:PMID
12819:Cell
12788:PMID
12753:PMID
12712:PMID
12658:PMID
12623:PMID
12571:PMID
12512:PMID
12463:PMID
12404:PMID
12355:PMID
12301:PMID
12252:PMID
12207:2019
12177:PMID
12126:PMID
12048:PMID
12006:PMID
11965:PMID
11914:PMID
11873:PMID
11814:PMID
11747:PMID
11712:PMID
11661:PMID
11610:PMID
11550:PMID
11507:PMID
11453:PMID
11363:PMID
11328:PMID
11263:PMID
11212:PMID
11169:PMID
11109:PMID
11090:PMID
11031:PMID
10988:PMID
10939:PMID
10880:PMID
10839:PMID
10821:Cell
10795:PMID
10732:PMID
10691:PMID
10640:PMID
10591:PMID
10542:PMID
10507:PMID
10458:PMID
10404:PMID
10345:PMID
10304:PMID
10255:PMID
10204:ISBN
10181:PMID
10144:2022
10125:PMID
10103:Cell
10069:PMID
10028:PMID
9971:PMID
9930:PMID
9880:PMID
9818:PMID
9769:PMID
9730:PMID
9671:PMID
9620:PMID
9561:PMID
9512:PMID
9455:PMID
9419:PMID
9370:PMID
9311:PMID
9262:PMID
9213:PMID
9164:PMID
9113:PMID
9078:PMID
9029:PMID
8972:PMID
8923:PMID
8855:PMID
8832:PMID
8781:PMID
8732:PMID
8683:PMID
8634:PMID
8583:PMID
8534:PMID
8477:PMID
8442:PMID
8397:2010
8358:PMID
8307:PMID
8258:PMID
8209:PMID
8160:PMID
8105:PMID
8070:PMID
8021:PMID
7972:PMID
7937:PMID
7883:PMID
7813:PMID
7770:PMID
7732:PMID
7681:PMID
7624:PMID
7576:PMID
7532:PMID
7497:PMID
7452:PMID
7381:PMID
7322:PMID
7273:PMID
7232:PMID
7210:1862
7180:PMID
7154:Cell
7102:PMID
7083:PMID
7022:ISBN
6985:PMID
6950:PMID
6932:Cell
6909:PMID
6860:PMID
6809:PMID
6752:PMID
6695:PMID
6638:PMID
6589:PMID
6554:PMID
6502:PMID
6461:PMID
6410:PMID
6348:PMID
6305:PMID
6251:PMID
6233:Cell
6210:PMID
6151:PMID
6102:PMID
6043:PMID
5989:PMID
5940:PMID
5914:Gene
5891:PMID
5873:Cell
5850:PMID
5832:Cell
5809:PMID
5768:PMID
5716:PMID
5681:PMID
5632:PMID
5561:PMID
5520:PMID
5510:ISBN
5479:PMID
5430:PMID
5377:PMID
5332:PMID
5289:PMID
5254:PMID
5202:PMID
5153:PMID
5096:PMID
5031:PMID
4990:PMID
4964:Cell
4941:PMID
4900:PMID
4882:Cell
4859:PMID
4810:PMID
4800:ISBN
4761:PMID
4718:PMID
4696:1779
4669:PMID
4610:PMID
4556:PMID
4507:PMID
4450:PMID
4409:PMID
4353:PMID
4305:link
4287:PMID
4222:PMID
4171:PMID
4125:PMID
4068:PMID
4025:PMID
3990:PMID
3941:PMID
3923:Cell
3895:PMID
3844:PMID
3793:PMID
3744:PMID
3695:PMID
3685:ISBN
3654:PMID
3636:Cell
3613:PMID
3562:PMID
3527:PMID
3478:PMID
3427:PMID
3368:PMID
3311:PMID
3250:PMID
3207:PMID
3166:PMID
3148:Cell
3125:PMID
3079:PMID
3028:PMID
2972:PMID
2916:PMID
2850:PMID
2796:PMID
2750:PMID
2732:Cell
2704:PMID
2655:PMID
2601:PMID
2583:Cell
2555:PMID
2503:PMID
2454:PMID
2405:PMID
2338:PMID
2263:PMID
2220:PMID
2185:PMID
2159:Cell
2131:PMID
2109:2014
2082:PMID
2064:Cell
2036:PMID
2010:Cell
1727:fosB
1723:DRD1
1715:fosB
1679:BDNF
1657:and
1527:Mmp2
1448:and
1363:HMGA
1353:mRNA
1344:MGMT
1333:MLH1
1321:MLH1
1177:mRNA
1173:mRNA
1169:mRNA
1152:, a
1086:and
1032:and
960:RNAa
949:ITPR
892:and
799:XRN2
729:TRBP
717:PIWI
601:3A6P
505:5B16
496:zinc
314:and
308:Mir1
303:and
271:and
253:and
114:, or
101:mRNA
93:mRNA
51:mRNA
15773:RNA
15082:RNA
14983:939
14978:938
14973:885
14968:874
14963:873
14958:872
14953:854
14948:828
14943:824
14938:802
14933:786
14928:767
14923:765
14918:764
14913:744
14908:720
14903:711
14898:708
14893:675
14888:672
14883:671
14878:663
14873:661
14868:657
14863:652
14858:650
14853:638
14848:636
14843:633
14838:632
14833:625
14828:624
14823:612
14818:612
14813:615
14808:612
14803:605
14798:601
14782:598
14777:592
14772:590
14767:589
14762:584
14757:580
14752:575
14747:572
14742:569
14737:562
14732:558
14727:552
14722:550
14717:549
14712:548
14707:544
14702:542
14697:541
14692:535
14687:529
14682:506
14677:505
14672:504
14667:503
14651:500
14646:498
14641:497
14636:492
14631:491
14626:489
14621:488
14616:484
14611:474
14606:455
14601:454
14596:452
14591:451
14586:449
14581:448
14576:444
14571:434
14566:433
14561:432
14556:431
14551:430
14546:425
14541:423
14536:408
14531:403
14515:399
14510:398
14505:397
14500:396
14495:395
14490:394
14485:390
14480:384
14475:383
14470:374
14465:370
14460:367
14455:365
14450:363
14445:361
14440:351
14435:350
14430:346
14425:345
14420:344
14415:340
14410:339
14405:338
14400:337
14395:331
14390:330
14385:326
14380:322
14375:305
14370:301
14354:299
14349:296
14344:282
14339:281
14334:279
14329:278
14324:277
14319:275
14314:241
14309:224
14304:223
14299:221
14294:219
14289:218
14284:216
14279:214
14274:212
14269:210
14264:208
14259:205
14249:203
14244:202
14228:200
14223:199
14218:198
14213:196
14208:194
14203:193
14193:191
14188:190
14183:188
14178:187
14173:186
14168:185
14163:184
14158:181
14153:172
14148:166
14143:160
14138:156
14133:155
14128:154
14123:153
14118:150
14108:146
14103:145
14098:144
14093:143
14088:141
14083:138
14078:137
14073:135
14068:134
14063:133
14058:132
14053:130
14048:129
14043:127
14038:126
14033:124
14028:122
14018:101
13671:doi
13625:doi
13621:297
13588:PMC
13580:doi
13572:RNA
13534:PMC
13524:doi
13483:PMC
13473:doi
13432:PMC
13424:doi
13364:PMC
13356:doi
13315:hdl
13307:doi
13270:PMC
13260:doi
13213:doi
13168:PMC
13158:doi
13117:PMC
13107:doi
13057:doi
13053:303
13020:doi
13016:218
12979:doi
12935:doi
12894:PMC
12884:doi
12872:108
12835:PMC
12827:doi
12823:147
12780:doi
12743:doi
12739:281
12702:hdl
12694:doi
12650:doi
12646:236
12613:PMC
12605:doi
12561:PMC
12551:doi
12502:PMC
12494:doi
12453:PMC
12443:doi
12394:PMC
12386:doi
12345:PMC
12337:doi
12291:PMC
12283:doi
12242:PMC
12234:doi
12167:PMC
12157:doi
12116:PMC
12106:doi
12040:doi
12036:122
11996:doi
11955:PMC
11945:doi
11904:doi
11863:PMC
11853:doi
11841:105
11804:PMC
11794:doi
11739:doi
11702:PMC
11692:doi
11651:PMC
11641:doi
11600:PMC
11592:doi
11542:doi
11530:420
11497:PMC
11487:doi
11483:287
11443:PMC
11435:doi
11355:doi
11351:122
11318:PMC
11310:doi
11255:doi
11243:316
11204:doi
11161:doi
11101:doi
11080:PMC
11070:doi
11066:282
11023:doi
11011:436
10978:PMC
10970:doi
10929:PMC
10919:doi
10907:103
10870:doi
10866:116
10829:doi
10825:129
10785:PMC
10775:doi
10763:105
10722:doi
10718:106
10681:PMC
10671:doi
10630:PMC
10622:doi
10581:doi
10534:doi
10530:574
10497:PMC
10489:doi
10448:PMC
10440:doi
10394:PMC
10384:doi
10372:107
10335:doi
10331:128
10294:PMC
10286:doi
10282:138
10245:PMC
10237:doi
10171:doi
10115:doi
10107:185
10059:doi
10018:PMC
10010:doi
10006:233
9961:doi
9957:233
9920:doi
9870:PMC
9860:doi
9808:doi
9761:doi
9720:PMC
9710:doi
9661:hdl
9651:doi
9647:152
9610:PMC
9600:doi
9588:101
9551:PMC
9543:doi
9502:PMC
9494:doi
9447:doi
9409:PMC
9401:doi
9360:PMC
9350:doi
9301:PMC
9293:doi
9252:PMC
9244:doi
9203:PMC
9195:doi
9154:PMC
9144:doi
9105:doi
9068:PMC
9060:doi
9019:PMC
9011:doi
8962:doi
8950:318
8913:PMC
8905:doi
8847:doi
8822:PMC
8812:doi
8771:PMC
8763:doi
8755:RNA
8722:PMC
8714:doi
8673:PMC
8665:doi
8624:PMC
8614:doi
8573:PMC
8565:doi
8557:RNA
8524:PMC
8516:doi
8469:doi
8432:PMC
8424:doi
8348:PMC
8338:doi
8297:PMC
8289:doi
8285:342
8248:PMC
8240:doi
8199:doi
8187:465
8152:doi
8097:doi
8060:PMC
8052:doi
8011:doi
7964:doi
7927:PMC
7919:doi
7873:PMC
7863:doi
7851:105
7805:doi
7762:doi
7722:PMC
7712:doi
7671:PMC
7663:doi
7614:doi
7568:doi
7524:doi
7489:doi
7475:339
7442:PMC
7434:doi
7371:PMC
7361:doi
7349:118
7312:PMC
7304:doi
7263:doi
7259:185
7222:PMC
7214:doi
7170:PMC
7162:doi
7158:146
7094:doi
7073:PMC
7063:doi
7051:105
6977:doi
6940:doi
6936:123
6899:PMC
6891:doi
6850:PMC
6840:doi
6799:PMC
6791:doi
6779:336
6742:PMC
6734:doi
6722:336
6685:PMC
6677:doi
6669:RNA
6628:PMC
6620:doi
6581:doi
6544:PMC
6536:doi
6492:doi
6451:PMC
6441:doi
6400:PMC
6392:doi
6384:RNA
6340:doi
6328:461
6295:PMC
6287:doi
6241:doi
6237:120
6200:hdl
6190:doi
6141:PMC
6133:doi
6092:PMC
6082:doi
6070:105
6033:PMC
6023:doi
6019:284
5979:PMC
5971:doi
5930:PMC
5922:doi
5918:356
5881:doi
5877:115
5840:doi
5836:115
5799:doi
5795:279
5758:doi
5708:doi
5671:PMC
5663:doi
5624:doi
5620:172
5592:doi
5551:doi
5502:doi
5469:PMC
5461:doi
5457:475
5420:doi
5369:doi
5324:doi
5281:doi
5244:PMC
5236:doi
5192:PMC
5184:doi
5143:PMC
5135:doi
5123:448
5086:PMC
5078:doi
5066:448
5021:doi
5017:586
4980:PMC
4972:doi
4968:152
4931:doi
4890:doi
4886:125
4849:PMC
4841:doi
4792:doi
4753:doi
4741:425
4708:PMC
4700:doi
4659:PMC
4649:doi
4600:PMC
4592:doi
4546:PMC
4538:doi
4530:RNA
4497:PMC
4489:doi
4440:doi
4436:272
4399:PMC
4391:doi
4383:RNA
4343:PMC
4335:doi
4277:PMC
4269:doi
4257:455
4214:doi
4202:455
4163:doi
4151:433
4115:PMC
4107:doi
4060:doi
4017:doi
3980:PMC
3972:doi
3931:doi
3927:115
3885:PMC
3875:doi
3834:PMC
3824:doi
3783:PMC
3775:doi
3734:PMC
3726:doi
3718:RNA
3677:doi
3644:doi
3603:PMC
3593:doi
3554:doi
3550:125
3517:PMC
3509:doi
3468:PMC
3458:doi
3417:PMC
3407:doi
3395:102
3358:PMC
3350:doi
3301:hdl
3293:doi
3281:303
3242:doi
3230:432
3197:doi
3193:187
3156:doi
3152:113
3117:doi
3071:doi
3059:309
3020:doi
3008:294
2964:doi
2952:294
2906:hdl
2898:doi
2886:294
2842:doi
2830:408
2788:doi
2776:403
2740:doi
2694:PMC
2686:doi
2645:PMC
2637:doi
2591:doi
2587:120
2545:doi
2493:PMC
2485:doi
2444:PMC
2436:doi
2395:PMC
2387:doi
2360:at
2328:PMC
2318:hdl
2310:doi
2298:466
2255:doi
2212:doi
2175:PMC
2167:doi
2163:136
2121:PMC
2113:doi
2072:doi
2068:116
2026:PMC
2018:doi
2014:173
1791:222
1787:221
1783:155
1605:).
1515:Bax
1511:p53
1329:CpG
896:of
877:in
767:or
622:Ran
597:PDB
570:to
501:PDB
310:in
299:in
15759::
14002:96
13997:92
13992:84
13987:71
13982:67
13977:62
13972:61
13967:58
13962:52
13957:48
13947:42
13942:34
13937:33
13932:32
13927:31
13922:30
13917:29
13912:28
13907:26
13902:25
13897:24
13892:23
13887:22
13882:21
13877:19
13872:17
13867:16
13862:15
13857:14
13852:11
13847:10
13679:.
13667:75
13665:.
13661:.
13639:.
13631:.
13619:.
13596:.
13586:.
13574:.
13570:.
13542:.
13532:.
13518:.
13514:.
13491:.
13481:.
13469:14
13467:.
13463:.
13440:.
13430:.
13420:86
13418:.
13414:.
13395:.
13372:.
13362:.
13350:.
13346:.
13323:.
13313:.
13303:62
13301:.
13278:.
13268:.
13256:11
13254:.
13250:.
13227:.
13219:.
13211:.
13201:76
13199:.
13176:.
13166:.
13154:12
13152:.
13148:.
13125:.
13115:.
13101:.
13097:.
13085:^
13071:.
13063:.
13051:.
13028:.
13014:.
13010:.
12987:.
12975:16
12973:.
12969:.
12957:^
12943:.
12931:13
12929:.
12925:.
12902:.
12892:.
12882:.
12870:.
12866:.
12843:.
12833:.
12821:.
12817:.
12794:.
12786:.
12776:27
12774:.
12751:.
12737:.
12733:.
12710:.
12700:.
12690:21
12688:.
12664:.
12656:.
12644:.
12621:.
12611:.
12599:.
12595:.
12583:^
12569:.
12559:.
12549:.
12537:.
12533:.
12510:.
12500:.
12490:34
12488:.
12484:.
12461:.
12451:.
12441:.
12429:.
12425:.
12402:.
12392:.
12382:13
12380:.
12376:.
12353:.
12343:.
12333:35
12331:.
12327:.
12313:^
12299:.
12289:.
12279:14
12277:.
12273:.
12250:.
12240:.
12230:43
12228:.
12224:.
12197:.
12175:.
12165:.
12153:15
12151:.
12147:.
12124:.
12114:.
12104:.
12090:.
12086:.
12064:.
12054:.
12046:.
12034:.
12030:.
12004:.
11992:15
11990:.
11986:.
11963:.
11953:.
11941:13
11939:.
11935:.
11912:.
11900:19
11898:.
11894:.
11871:.
11861:.
11851:.
11839:.
11835:.
11812:.
11802:.
11792:.
11782:18
11780:.
11776:.
11753:.
11745:.
11735:10
11733:.
11710:.
11700:.
11686:.
11682:.
11659:.
11649:.
11637:17
11635:.
11631:.
11608:.
11598:.
11586:.
11582:.
11570:^
11556:.
11548:.
11540:.
11528:.
11505:.
11495:.
11481:.
11477:.
11465:^
11451:.
11441:.
11433:.
11421:.
11417:.
11375:^
11361:.
11349:.
11326:.
11316:.
11308:.
11296:.
11292:.
11269:.
11261:.
11253:.
11241:.
11218:.
11210:.
11200:13
11198:.
11175:.
11167:.
11157:13
11155:.
11088:.
11078:.
11064:.
11060:.
11037:.
11029:.
11021:.
11009:.
10986:.
10976:.
10966:42
10964:.
10960:.
10937:.
10927:.
10917:.
10905:.
10901:.
10878:.
10864:.
10860:.
10837:.
10823:.
10819:.
10807:^
10793:.
10783:.
10773:.
10761:.
10757:.
10730:.
10716:.
10712:.
10689:.
10679:.
10665:.
10661:.
10638:.
10628:.
10618:31
10616:.
10612:.
10589:.
10577:64
10575:.
10571:.
10548:.
10540:.
10528:.
10505:.
10495:.
10485:12
10483:.
10479:.
10456:.
10446:.
10436:14
10434:.
10430:.
10416:^
10402:.
10392:.
10382:.
10370:.
10366:.
10343:.
10329:.
10325:.
10302:.
10292:.
10280:.
10276:.
10253:.
10243:.
10231:.
10227:.
10179:.
10167:21
10165:.
10161:.
10131:.
10123:.
10105:.
10101:.
10075:.
10067:.
10055:34
10053:.
10049:.
10026:.
10016:.
10004:.
10000:.
9977:.
9969:.
9955:.
9951:.
9928:.
9916:20
9914:.
9910:.
9878:.
9868:.
9858:.
9848:11
9846:.
9842:.
9830:^
9816:.
9804:39
9802:.
9798:.
9775:.
9767:.
9757:50
9755:.
9751:.
9728:.
9718:.
9706:14
9704:.
9700:.
9677:.
9669:.
9659:.
9645:.
9641:.
9618:.
9608:.
9598:.
9586:.
9582:.
9559:.
9549:.
9539:43
9537:.
9533:.
9510:.
9500:.
9490:89
9488:.
9484:.
9461:.
9453:.
9443:41
9441:.
9417:.
9407:.
9397:37
9391:.
9368:.
9358:.
9348:.
9336:.
9332:.
9309:.
9299:.
9289:37
9287:.
9283:.
9260:.
9250:.
9240:36
9238:.
9234:.
9211:.
9201:.
9191:27
9189:.
9185:.
9162:.
9152:.
9140:10
9138:.
9134:.
9111:.
9101:45
9099:.
9076:.
9066:.
9054:.
9050:.
9027:.
9017:.
9007:34
9005:.
9001:.
8978:.
8970:.
8960:.
8948:.
8944:.
8921:.
8911:.
8901:42
8899:.
8895:.
8830:.
8820:.
8806:.
8802:.
8779:.
8769:.
8759:10
8757:.
8753:.
8730:.
8720:.
8710:39
8708:.
8704:.
8681:.
8671:.
8661:32
8659:.
8655:.
8632:.
8622:.
8610:11
8608:.
8604:.
8581:.
8571:.
8561:11
8559:.
8555:.
8532:.
8522:.
8512:33
8510:.
8506:.
8483:.
8475:.
8463:.
8440:.
8430:.
8420:34
8418:.
8414:.
8385:30
8383:.
8379:.
8356:.
8346:.
8334:13
8332:.
8328:.
8305:.
8295:.
8283:.
8279:.
8256:.
8246:.
8236:23
8234:.
8230:.
8207:.
8197:.
8185:.
8181:.
8158:.
8150:.
8140:24
8138:.
8134:.
8111:.
8103:.
8093:32
8091:.
8068:.
8058:.
8048:22
8046:.
8042:.
8019:.
8007:25
8005:.
8001:.
7978:.
7970:.
7960:36
7958:.
7935:.
7925:.
7913:.
7909:.
7895:^
7881:.
7871:.
7861:.
7849:.
7845:.
7833:^
7819:.
7811:.
7801:78
7799:.
7776:.
7768:.
7758:11
7756:.
7744:^
7730:.
7720:.
7708:12
7706:.
7702:.
7679:.
7669:.
7659:23
7657:.
7653:.
7630:.
7622:.
7610:31
7608:.
7604:.
7588:^
7574:.
7564:26
7562:.
7538:.
7530:.
7518:.
7495:.
7487:.
7473:.
7450:.
7440:.
7430:17
7428:.
7424:.
7403:.
7387:.
7379:.
7369:.
7359:.
7347:.
7343:.
7320:.
7310:.
7298:.
7294:.
7271:.
7257:.
7253:.
7230:.
7220:.
7208:.
7204:.
7192:^
7178:.
7168:.
7156:.
7152:.
7081:.
7071:.
7061:.
7049:.
7045:.
6991:.
6983:.
6971:.
6948:.
6934:.
6930:.
6907:.
6897:.
6885:.
6881:.
6858:.
6848:.
6836:10
6834:.
6830:.
6807:.
6797:.
6789:.
6777:.
6773:.
6750:.
6740:.
6732:.
6720:.
6716:.
6693:.
6683:.
6673:15
6671:.
6667:.
6644:.
6636:.
6626:.
6616:65
6614:.
6610:.
6587:.
6577:12
6575:.
6552:.
6542:.
6532:18
6530:.
6526:.
6514:^
6500:.
6488:48
6486:.
6482:.
6459:.
6449:.
6435:.
6431:.
6408:.
6398:.
6388:18
6386:.
6382:.
6368:^
6354:.
6346:.
6338:.
6326:.
6303:.
6293:.
6283:17
6281:.
6277:.
6263:^
6249:.
6235:.
6231:.
6208:.
6198:.
6188:.
6178:15
6176:.
6172:.
6149:.
6139:.
6129:16
6127:.
6123:.
6100:.
6090:.
6080:.
6068:.
6064:.
6041:.
6031:.
6017:.
6013:.
6001:^
5987:.
5977:.
5965:.
5961:.
5938:.
5928:.
5916:.
5912:.
5889:.
5875:.
5871:.
5848:.
5834:.
5830:.
5807:.
5793:.
5789:.
5766:.
5754:16
5752:.
5748:.
5736:^
5722:.
5714:.
5702:.
5679:.
5669:.
5659:11
5657:.
5653:.
5630:.
5618:.
5606:^
5588:10
5586:.
5582:.
5559:.
5547:22
5545:.
5541:.
5518:.
5508:.
5477:.
5467:.
5455:.
5451:.
5428:.
5416:71
5414:.
5410:.
5396:^
5375:.
5365:16
5363:.
5355:,
5344:^
5330:.
5320:89
5318:.
5295:.
5287:.
5277:11
5275:.
5252:.
5242:.
5232:36
5230:.
5226:.
5214:^
5200:.
5190:.
5180:28
5178:.
5174:.
5151:.
5141:.
5133:.
5121:.
5117:.
5094:.
5084:.
5076:.
5064:.
5060:.
5037:.
5029:.
5015:.
5011:.
4988:.
4978:.
4966:.
4962:.
4939:.
4925:.
4921:.
4898:.
4884:.
4880:.
4857:.
4847:.
4837:18
4835:.
4831:.
4808:.
4798:.
4767:.
4759:.
4751:.
4739:.
4716:.
4706:.
4694:.
4690:.
4667:.
4657:.
4647:.
4635:.
4631:.
4608:.
4598:.
4588:23
4586:.
4582:.
4568:^
4554:.
4544:.
4534:11
4532:.
4528:.
4505:.
4495:.
4485:26
4483:.
4479:.
4456:.
4448:.
4434:.
4430:.
4407:.
4397:.
4387:10
4385:.
4381:.
4365:^
4351:.
4341:.
4331:14
4329:.
4325:.
4313:^
4301:}}
4297:{{
4285:.
4267:.
4255:.
4251:.
4228:.
4220:.
4212:.
4200:.
4177:.
4169:.
4161:.
4149:.
4137:^
4123:.
4113:.
4103:39
4101:.
4097:.
4074:.
4066:.
4056:37
4054:.
4031:.
4023:.
4013:38
4011:.
3988:.
3978:.
3968:27
3966:.
3962:.
3939:.
3925:.
3921:.
3907:^
3893:.
3883:.
3871:20
3869:.
3865:.
3842:.
3832:.
3818:.
3814:.
3791:.
3781:.
3771:34
3769:.
3765:.
3742:.
3732:.
3720:.
3716:.
3693:.
3683:.
3652:.
3640:75
3638:.
3634:.
3611:.
3601:.
3587:.
3583:.
3560:.
3548:.
3525:.
3515:.
3505:11
3503:.
3499:.
3476:.
3466:.
3454:27
3452:.
3448:.
3425:.
3415:.
3405:.
3393:.
3389:.
3366:.
3356:.
3346:91
3344:.
3340:.
3317:.
3309:.
3299:.
3291:.
3279:.
3256:.
3248:.
3240:.
3228:.
3205:.
3191:.
3187:.
3164:.
3150:.
3146:.
3123:.
3113:57
3111:.
3099:^
3085:.
3077:.
3069:.
3057:.
3034:.
3026:.
3018:.
3006:.
2992:^
2978:.
2970:.
2962:.
2950:.
2936:^
2922:.
2914:.
2904:.
2896:.
2884:.
2870:^
2856:.
2848:.
2840:.
2828:.
2816:^
2802:.
2794:.
2786:.
2774:.
2762:^
2748:.
2736:75
2734:.
2730:.
2716:^
2702:.
2692:.
2682:49
2680:.
2676:.
2653:.
2643:.
2633:19
2631:.
2627:.
2613:^
2599:.
2585:.
2581:.
2567:^
2553:.
2543:.
2533:12
2531:.
2527:.
2515:^
2501:.
2491:.
2481:17
2479:.
2475:.
2452:.
2442:.
2432:48
2430:.
2426:.
2403:.
2393:.
2383:47
2381:.
2377:.
2336:.
2326:.
2316:.
2308:.
2296:.
2292:.
2269:.
2261:.
2251:16
2249:.
2226:.
2218:.
2208:16
2206:.
2183:.
2173:.
2161:.
2157:.
2143:^
2129:.
2119:.
2107:.
2103:.
2080:.
2066:.
2062:.
2048:^
2034:.
2024:.
2012:.
2008:.
1992:^
1808:,
1785:,
1769:,
1653:,
1649:,
1645:,
1597:,
1593:,
1589:,
1577:,
1525:,
1517:,
1249:.
1100:.
1016:.
986:.
943:.
754:.
599::
547:,
503::
471:A
451:,
337::
328::
261:,
148:.
15391:)
15335:e
15328:t
15321:v
15073:e
15066:t
15059:v
13832:7
13827:6
13822:5
13817:3
13812:2
13807:1
13784:e
13777:t
13770:v
13748:.
13687:.
13673::
13647:.
13627::
13604:.
13582::
13576:9
13550:.
13526::
13520:4
13499:.
13475::
13448:.
13426::
13380:.
13358::
13331:.
13317::
13309::
13286:.
13262::
13235:.
13215::
13207::
13184:.
13160::
13133:.
13109::
13103:1
13079:.
13059::
13036:.
13022::
12995:.
12981::
12951:.
12937::
12910:.
12886::
12878::
12851:.
12829::
12802:.
12782::
12759:.
12745::
12718:.
12704::
12696::
12672:.
12652::
12629:.
12607::
12601:5
12577:.
12553::
12545::
12539:8
12518:.
12496::
12469:.
12445::
12437::
12431:8
12410:.
12388::
12361:.
12339::
12307:.
12285::
12258:.
12236::
12209:.
12183:.
12159::
12132:.
12108::
12098::
12092:4
12042::
12012:.
11998::
11971:.
11947::
11920:.
11906::
11879:.
11855::
11847::
11820:.
11796::
11788::
11761:.
11741::
11718:.
11694::
11688:6
11667:.
11643::
11616:.
11594::
11588:3
11564:.
11544::
11536::
11513:.
11489::
11459:.
11437::
11429::
11423:4
11369:.
11357::
11334:.
11312::
11304::
11298:7
11277:.
11257::
11249::
11226:.
11206::
11183:.
11163::
11140:)
11138:.
11103::
11096:.
11072::
11045:.
11025::
11017::
10994:.
10972::
10945:.
10921::
10913::
10886:.
10872::
10845:.
10831::
10801:.
10777::
10769::
10738:.
10724::
10697:.
10673::
10667:3
10646:.
10624::
10597:.
10583::
10556:.
10536::
10513:.
10491::
10464:.
10442::
10410:.
10386::
10378::
10351:.
10337::
10310:.
10288::
10261:.
10239::
10233:3
10212:.
10187:.
10173::
10146:.
10117::
10083:.
10061::
10034:.
10012::
9985:.
9963::
9936:.
9922::
9886:.
9862::
9854::
9824:.
9810::
9783:.
9763::
9736:.
9712::
9685:.
9663::
9653::
9626:.
9602::
9594::
9567:.
9545::
9518:.
9496::
9469:.
9449::
9425:.
9403::
9376:.
9352::
9344::
9338:7
9317:.
9295::
9268:.
9246::
9219:.
9197::
9170:.
9146::
9119:.
9107::
9084:.
9062::
9056:1
9035:.
9013::
8986:.
8964::
8956::
8929:.
8907::
8880:)
8878:.
8849::
8838:.
8814::
8808:5
8787:.
8765::
8738:.
8716::
8689:.
8667::
8640:.
8616::
8589:.
8567::
8540:.
8518::
8491:.
8471::
8465:3
8448:.
8426::
8399:.
8364:.
8340::
8313:.
8291::
8264:.
8242::
8215:.
8201::
8193::
8166:.
8154::
8146::
8119:.
8099::
8076:.
8054::
8027:.
8013::
7986:.
7966::
7943:.
7921::
7915:2
7889:.
7865::
7857::
7827:.
7807::
7784:.
7764::
7738:.
7714::
7687:.
7665::
7638:.
7616::
7582:.
7570::
7546:.
7526::
7520:8
7503:.
7491::
7458:.
7436::
7395:.
7363::
7355::
7328:.
7306::
7300:2
7279:.
7265::
7238:.
7216::
7186:.
7164::
7137:)
7135:.
7096::
7089:.
7065::
7057::
7030:.
6999:.
6979::
6973:5
6956:.
6942::
6915:.
6893::
6887:7
6866:.
6842::
6815:.
6793::
6785::
6758:.
6736::
6728::
6701:.
6679::
6652:.
6622::
6595:.
6583::
6560:.
6538::
6508:.
6494::
6467:.
6443::
6437:5
6416:.
6394::
6362:.
6342::
6334::
6311:.
6289::
6257:.
6243::
6216:.
6202::
6192::
6184::
6157:.
6135::
6108:.
6084::
6076::
6049:.
6025::
5995:.
5973::
5967:7
5946:.
5924::
5897:.
5883::
5856:.
5842::
5815:.
5801::
5774:.
5760::
5730:.
5710::
5704:8
5687:.
5665::
5638:.
5626::
5600:.
5594::
5567:.
5553::
5526:.
5504::
5485:.
5463::
5436:.
5422::
5383:.
5371::
5338:.
5326::
5303:.
5283::
5260:.
5238::
5208:.
5186::
5159:.
5137::
5129::
5102:.
5080::
5072::
5045:.
5023::
4996:.
4974::
4947:.
4933::
4927:9
4906:.
4892::
4865:.
4843::
4816:.
4794::
4775:.
4755::
4747::
4724:.
4702::
4675:.
4651::
4643::
4637:3
4616:.
4594::
4562:.
4540::
4513:.
4491::
4464:.
4442::
4415:.
4393::
4359:.
4337::
4307:)
4293:.
4271::
4263::
4236:.
4216::
4208::
4185:.
4165::
4157::
4131:.
4109::
4082:.
4062::
4039:.
4019::
3996:.
3974::
3947:.
3933::
3901:.
3877::
3850:.
3826::
3820:5
3799:.
3777::
3750:.
3728::
3722:9
3701:.
3679::
3660:.
3646::
3619:.
3595::
3589:2
3568:.
3556::
3533:.
3511::
3484:.
3460::
3433:.
3409::
3401::
3374:.
3352::
3325:.
3303::
3295::
3287::
3264:.
3244::
3236::
3213:.
3199::
3172:.
3158::
3131:.
3119::
3093:.
3073::
3065::
3042:.
3022::
3014::
2986:.
2966::
2958::
2930:.
2908::
2900::
2892::
2864:.
2844::
2836::
2810:.
2790::
2782::
2756:.
2742::
2710:.
2688::
2661:.
2639::
2607:.
2593::
2561:.
2547::
2539::
2509:.
2487::
2460:.
2438::
2411:.
2389::
2344:.
2320::
2312::
2304::
2277:.
2257::
2234:.
2214::
2191:.
2169::
2137:.
2115::
2088:.
2074::
2042:.
2020::
978:(
926:;
364:(
67:(
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.