1629:
1306:
1641:
226:
are diagrams that depict evolutionary relationships within groups of taxa. These illustrations are accurate predictive device in modern genetics. They are usually depicted in either tree or ladder form. Synapomorphies then create evidence for historical relationships and their associated hierarchical
213:
and paired appendages in both sharks and dogs, but not in lampreys or close invertebrate relatives, identifies these traits as synapomorphies. This supports the hypothesis that dogs and sharks are more closely related to each other than to lampreys.
227:
structure. Evolutionarily, a synapomorphy is the marker for the most recent common ancestor of the monophyletic group consisting of a set of taxa in a cladogram. What counts as a synapomorphy for one clade may well be a primitive character or
267:
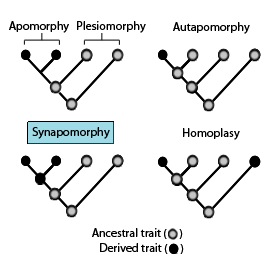
These phylogenetic terms are used to describe different patterns of ancestral and derived character or trait states as stated in the above diagram in association with apomorphies and synapomorphies.
364:
is the case where a character that appears homoplastic given the species tree actually has a single origin on the associated gene tree. Hemiplasy reflects gene tree-species tree discordance due to the
255:) the apomorphy: mammary glands are evolutionarily newer than vertebral column, so mammary glands are an autapomorphy if vertebral column is an apomorphy, but if mammary glands are the
31:
1154:
Copetti D, Búrquez A, Bustamante E, Charboneau JL, Childs KL, Eguiarte LE, Lee S, Liu TL, McMahon MM, Whiteman NK, Wing RA, Wojciechowski MF, Sanderson MJ (November 2017).
1013:
Archie JW (September 1989). "Homoplasy Excess Ratios: New
Indices for Measuring Levels of Homoplasy in Phylogenetic Systematics and a Critique of the Consistency Index".
305:– a derived trait that is found in some or all terminal groups of a clade, and inherited from a common ancestor, for which it was an autapomorphy (i.e., not present in
298:– a derived trait. Apomorphy shared by two or more taxa and inherited from a common ancestor is synapomorphy. Apomorphy unique to a given taxon is autapomorphy.
247:
for mammals in relation to one another—rodents and primates, for example. So the concept can be understood as well in terms of "a character newer than" (
315:– a synapomorphy that has been lost again in many members of the clade. If lost in all but one, it can be hard to distinguish from an autapomorphy.
138:
gait and lack of fur. Thus, these derived traits are also synapomorphies of mammals in general as they are not shared by other vertebrate animals.
676:
Novick LR, Catley KM. Understanding phylogenies in biology: the influence of a
Gestalt perceptual principle. J Exp Psychol Appl. 2007;13:197–223.
811:
338:
leads to species independently sharing a trait that is different from the trait inferred to have been present in their common ancestor.
630:
1672:
1048:
Wake DB, Wake MH, Specht CD (February 2011). "Homoplasy: from detecting pattern to determining process and mechanism of evolution".
108:
1244:
948:
908:"Chapter 3: Introduction of Phylogenetics and its Molecular Aspects." World Scientific Publishing Company, 1st edition. 2009.
881:
836:
755:
720:
696:
653:
559:
400:
1439:
284:
Pseudoplesiomorphy – is a trait that cannot be identified as neither a plesiomorphy nor an apomorphy that is a reversal.
532:
1399:
782:
591:
499:
1479:
252:
66:
523:
Hillis, David M.; Sadava, David; Hill, Richard W.; Price, Mary V. (2014). "Reconstructing and using phylogenies".
1484:
1417:
365:
1645:
1494:
1424:
182:
631:"Trait Evolution on a Phylogenetic Tree: Relatedness, Similarity, and the Myth of Evolutionary Advancement"
289:
Reversal – is a loss of derived trait present in ancestor and the reestablishment of a plesiomorphic trait.
1404:
1311:
1267:
1237:
479:
82:
361:
205:
share some features, like a nervous system, that are not synapomorphic because they are also shared by
135:
97:
1594:
1520:
449:
17:
804:
608:
1156:"Extensive gene tree discordance and hemiplasy shaped the genomes of North American columnar cacti"
172:
37:
showing the terminology used to describe different patterns of ancestral and derived character or
1474:
1272:
1093:
873:
Bioinformatics for
Beginners: Genes, Genomes, Molecular Evolution, Databases and Analytical Tools
356:– trait present in an ancestor but not in direct descendants that reappears in later descendants.
1667:
1633:
1357:
1230:
162:
853:
826:
745:
710:
583:
549:
1609:
1287:
938:
871:
772:
434:
Futuyma, Douglas J.; Kirkpatrick, Mark (2017). "Phylogeny: The unity and diversity of life".
78:
148:
1446:
1352:
1216:
1167:
1057:
346:
335:
8:
1489:
1371:
1171:
1061:
1412:
1366:
1282:
1190:
1155:
1081:
1030:
792:
491:
302:
90:
1340:
1195:
1136:
1073:
995:
954:
944:
919:
887:
877:
832:
778:
751:
726:
716:
692:
587:
576:
555:
528:
495:
396:
1085:
1429:
1383:
1185:
1175:
1126:
1065:
1022:
985:
487:
345:– derived trait present in two groups or species without a common ancestor due to
331:
58:
38:
334:
has been gained or lost independently in separate lineages during evolution. This
1530:
686:
390:
272:
244:
1504:
1160:
Proceedings of the
National Academy of Sciences of the United States of America
1131:
1114:
1094:"Homoplasy: A good thread to pull to understand the evolutionary ball of yarn"
730:
1661:
1499:
1469:
1376:
1253:
958:
891:
232:
157:
112:
46:
1180:
1069:
990:
973:
1604:
1550:
1545:
1540:
1525:
1333:
1328:
1199:
1140:
1077:
999:
318:
292:
Convergence – independent evolution of a similar trait in two or more taxa.
248:
228:
206:
153:
222:
The concept of synapomorphy depends on a given clade in the tree of life.
1599:
1292:
527:(2nd ed.). Sunderland, Mass.: Sinauer Associates. pp. 325–342.
34:
438:(4th ed.). Sunderland, Mass.: Sinauer Associates. pp. 401–429.
321:– a distinctive derived trait that is unique to a given taxon or group.
1277:
1034:
327:
120:
86:
420:(4th ed.). Sunderland, Mass.: Sinauer Associates. pp. 27–53.
1614:
1578:
1573:
1568:
1463:
1345:
353:
342:
223:
127:
62:
1026:
281:– a symplesiomorphy discussed in reference to a more derived state.
256:
240:
1153:
747:
The Future of
Phylogenetic Systematics: The Legacy of Willi Hennig
388:
231:
at a less inclusive or nested clade. For example, the presence of
236:
198:
131:
1222:
1305:
654:"Gills, fins and the evolution of vertebrate paired appendages"
416:
Futuyma, Douglas J.; Kirkpatrick, Mark (2017). "Tree of life".
123:
116:
30:
478:
Kitching, Ian J.; Forey, Peter L.; Williams, David M. (2001).
1434:
1362:
456:. University of California Museum of Paleontology. 5 May 2021
202:
74:
972:
Brandley MC, Warren DL, Leaché AD, McGuire JA (April 2009).
971:
100:
685:
Roderick D.M. Page; Edward C. Holmes (14 July 2009).
607:
Barton N, Briggs D, Eisen J, Goldstein D, Patel N (2007).
606:
259:
being considered then vertebral column is a plesiomorphy.
210:
104:
1115:"Hemiplasy: a new term in the lexicon of phylogenetics"
743:
522:
477:
389:
Roderick D.M. Page; Edward C. Holmes (14 July 2009).
1301:
940:
Homoplasy: The
Recurrence of Similarity in Evolution
824:
712:
Encyclopedia of
Ecology and Environmental Management
582:. Tubingen, DEU: Walter de Gruyter. 1996. p.
575:
473:
471:
433:
415:
134:, which have retained their ancestral traits of a
275:– an ancestral trait shared by two or more taxa.
1659:
936:
859:. Washington D.C.: George Washington University.
384:
382:
468:
1047:
920:"Similarity Happens! The Problem of Homoplasy"
744:Williams D, Schmitt M, Wheeler Q (July 2016).
516:
442:
429:
427:
1238:
1112:
379:
1147:
1106:
1041:
876:(1st ed.). Academic Press. p. 51.
688:Molecular Evolution: A Phylogenetic Approach
651:
392:Molecular Evolution: A Phylogenetic Approach
937:Sanderson MJ, Hufford L (21 October 1996).
624:
622:
547:
424:
409:
262:
1245:
1231:
486:(2nd ed.). Elsevier. pp. 33–45.
96:Examples of apomorphy are the presence of
1189:
1179:
1130:
989:
869:
825:Russell PJ, Hertz PE, McMillan B (2013).
851:
619:
109:the evolution of three middle ear bones
29:
810:CS1 maint: location missing publisher (
770:
14:
1660:
1012:
917:
904:Appel, Ron D.; Feytmans, Ernest.
615:. Cold Spring Harbor Laboratory Press.
73:is an apomorphy shared by two or more
1226:
708:
1640:
906:Bioinformatics: a Swiss Perspective.
628:
1113:Avise JC, Robinson TJ (June 2008).
1100:(Press release). February 25, 2011.
24:
924:Evolution Today & Science News
492:10.1016/B978-0-12-384719-5.00022-8
450:"Reconstructing trees: Cladistics"
25:
1684:
1252:
1210:
600:
217:
1673:Evolutionary biology terminology
1639:
1628:
1627:
1480:Phylogenetic comparative methods
1304:
251:) and "a character older than" (
27:Two concepts on heritable traits
1485:Phylogenetic niche conservatism
1006:
965:
930:
911:
898:
863:
845:
818:
764:
737:
702:
679:
670:
645:
209:. In contrast, the presence of
152:—coined by German entomologist
854:"Basics of Cladistic Analysis"
750:. Cambridge University Press.
691:. John Wiley & Sons.
568:
541:
13:
1:
974:"Homoplasy and clade support"
609:"Phylogenetic Reconstruction"
372:
170:), meaning "with, together";
828:Biology: The Dynamic Science
771:Simpson MG (9 August 2011).
578:Concise Encyclopedia Biology
484:Encyclopedia of Biodiversity
482:. In Levin, Simon A. (ed.).
180:), meaning "away from"; and
141:
65:from its ancestral form (or
61:or character state that has
7:
1405:Phylogenetic reconciliation
1312:Evolutionary biology portal
1268:Computational phylogenetics
918:Gauger A (April 17, 2012).
652:andrewgillis (2016-04-19).
548:Currie PJ, Padia K (1997).
193:
83:most recent common ancestor
10:
1689:
870:Choudhuri S (2014-05-09).
190:), meaning "shape, form".
181:
171:
161:
1623:
1595:Phylogenetic nomenclature
1587:
1561:
1513:
1455:
1392:
1321:
1299:
1260:
1132:10.1080/10635150802164587
715:. John Wiley & Sons.
554:. Elsevier. p. 543.
551:Encyclopedia of Dinosaurs
395:. John Wiley & Sons.
81:to have evolved in their
263:Relations to other terms
1475:Molecular phylogenetics
1425:Distance-matrix methods
1273:Molecular phylogenetics
1181:10.1073/pnas.1706367114
1070:10.1126/science.1188545
454:Understanding Evolution
366:multispecies coalescent
313:Underlying synapomorphy
89:, synapomorphy implies
1495:Phylogenetics software
1409:Probabilistic methods
1358:Long branch attraction
328:biological systematics
235:is a synapomorphy for
147:
42:
1288:Evolutionary taxonomy
991:10.1093/sysbio/syp019
156:—is derived from the
33:
1447:Three-taxon analysis
1353:Phylogenetic network
831:. Cengage Learning.
629:Baum, David (2008).
347:convergent evolution
336:convergent evolution
309:immediate ancestor).
1490:Phylogenetic signal
1172:2017PNAS..11412003C
1166:(45): 12003–12008.
1062:2011Sci...331.1032W
852:Lipscomb D (1998).
1418:Bayesian inference
1413:Maximum likelihood
1119:Systematic Biology
1015:Systematic Biology
978:Systematic Biology
525:Principles of Life
43:
1655:
1654:
1400:Maximum parsimony
1393:Inference methods
1341:Phylogenetic tree
950:978-0-08-053411-4
883:978-0-12-410471-6
838:978-1-285-41534-5
774:Plant Systematics
757:978-1-107-11764-8
722:978-1-4443-1324-6
709:Calow PP (2009).
697:978-1-4443-1336-9
561:978-0-08-049474-6
402:978-1-4443-1336-9
119:but not in other
77:and is therefore
16:(Redirected from
1680:
1643:
1642:
1631:
1630:
1430:Neighbor-joining
1384:Ghost population
1314:
1309:
1308:
1247:
1240:
1233:
1224:
1223:
1204:
1203:
1193:
1183:
1151:
1145:
1144:
1134:
1110:
1104:
1101:
1089:
1056:(6020): 1032–5.
1045:
1039:
1038:
1010:
1004:
1003:
993:
969:
963:
962:
934:
928:
927:
915:
909:
902:
896:
895:
867:
861:
860:
858:
849:
843:
842:
822:
816:
815:
808:
802:
798:
796:
788:
768:
762:
761:
741:
735:
734:
706:
700:
683:
677:
674:
668:
667:
665:
664:
649:
643:
642:
635:Nature Education
626:
617:
616:
604:
598:
597:
581:
572:
566:
565:
545:
539:
538:
520:
514:
512:
510:
508:
475:
466:
465:
463:
461:
446:
440:
439:
431:
422:
421:
413:
407:
406:
386:
185:
175:
165:
21:
1688:
1687:
1683:
1682:
1681:
1679:
1678:
1677:
1658:
1657:
1656:
1651:
1619:
1583:
1557:
1531:Symplesiomorphy
1509:
1451:
1388:
1317:
1310:
1303:
1297:
1261:Relevant fields
1256:
1251:
1213:
1208:
1207:
1152:
1148:
1111:
1107:
1092:
1046:
1042:
1027:10.2307/2992286
1011:
1007:
970:
966:
951:
935:
931:
916:
912:
903:
899:
884:
868:
864:
856:
850:
846:
839:
823:
819:
809:
800:
799:
790:
789:
785:
769:
765:
758:
742:
738:
723:
707:
703:
684:
680:
675:
671:
662:
660:
650:
646:
627:
620:
605:
601:
594:
574:
573:
569:
562:
546:
542:
535:
521:
517:
506:
504:
502:
476:
469:
459:
457:
448:
447:
443:
432:
425:
414:
410:
403:
387:
380:
375:
273:Symplesiomorphy
265:
245:symplesiomorphy
239:in relation to
220:
196:
144:
28:
23:
22:
15:
12:
11:
5:
1686:
1676:
1675:
1670:
1653:
1652:
1650:
1649:
1637:
1624:
1621:
1620:
1618:
1617:
1612:
1607:
1602:
1597:
1591:
1589:
1585:
1584:
1582:
1581:
1576:
1571:
1565:
1563:
1559:
1558:
1556:
1555:
1554:
1553:
1548:
1543:
1535:
1534:
1533:
1528:
1517:
1515:
1511:
1510:
1508:
1507:
1505:Phylogeography
1502:
1497:
1492:
1487:
1482:
1477:
1472:
1467:
1459:
1457:
1456:Current topics
1453:
1452:
1450:
1449:
1444:
1443:
1442:
1437:
1432:
1422:
1421:
1420:
1415:
1407:
1402:
1396:
1394:
1390:
1389:
1387:
1386:
1381:
1380:
1379:
1369:
1360:
1355:
1350:
1349:
1348:
1338:
1337:
1336:
1325:
1323:
1322:Basic concepts
1319:
1318:
1316:
1315:
1300:
1298:
1296:
1295:
1290:
1285:
1280:
1275:
1270:
1264:
1262:
1258:
1257:
1250:
1249:
1242:
1235:
1227:
1221:
1220:
1212:
1211:External links
1209:
1206:
1205:
1146:
1105:
1103:
1102:
1040:
1021:(1): 253–269.
1005:
964:
949:
929:
910:
897:
882:
862:
844:
837:
817:
783:
763:
756:
736:
721:
701:
678:
669:
644:
618:
599:
592:
567:
560:
540:
534:978-1464175121
533:
515:
500:
467:
441:
423:
408:
401:
377:
376:
374:
371:
370:
369:
359:
358:
357:
350:
324:
323:
322:
316:
310:
293:
290:
287:
286:
285:
282:
264:
261:
233:mammary glands
219:
218:Clade analysis
216:
195:
192:
143:
140:
113:mammary glands
26:
9:
6:
4:
3:
2:
1685:
1674:
1671:
1669:
1668:Phylogenetics
1666:
1665:
1663:
1648:
1647:
1638:
1636:
1635:
1626:
1625:
1622:
1616:
1613:
1611:
1608:
1606:
1603:
1601:
1598:
1596:
1593:
1592:
1590:
1586:
1580:
1577:
1575:
1572:
1570:
1567:
1566:
1564:
1560:
1552:
1549:
1547:
1544:
1542:
1539:
1538:
1536:
1532:
1529:
1527:
1524:
1523:
1522:
1519:
1518:
1516:
1512:
1506:
1503:
1501:
1500:Phylogenomics
1498:
1496:
1493:
1491:
1488:
1486:
1483:
1481:
1478:
1476:
1473:
1471:
1470:DNA barcoding
1468:
1466:
1465:
1461:
1460:
1458:
1454:
1448:
1445:
1441:
1440:Least squares
1438:
1436:
1433:
1431:
1428:
1427:
1426:
1423:
1419:
1416:
1414:
1411:
1410:
1408:
1406:
1403:
1401:
1398:
1397:
1395:
1391:
1385:
1382:
1378:
1377:Ghost lineage
1375:
1374:
1373:
1370:
1368:
1364:
1361:
1359:
1356:
1354:
1351:
1347:
1344:
1343:
1342:
1339:
1335:
1332:
1331:
1330:
1327:
1326:
1324:
1320:
1313:
1307:
1302:
1294:
1291:
1289:
1286:
1284:
1281:
1279:
1276:
1274:
1271:
1269:
1266:
1265:
1263:
1259:
1255:
1254:Phylogenetics
1248:
1243:
1241:
1236:
1234:
1229:
1228:
1225:
1218:
1215:
1214:
1201:
1197:
1192:
1187:
1182:
1177:
1173:
1169:
1165:
1161:
1157:
1150:
1142:
1138:
1133:
1128:
1124:
1120:
1116:
1109:
1099:
1095:
1091:
1090:
1087:
1083:
1079:
1075:
1071:
1067:
1063:
1059:
1055:
1051:
1044:
1036:
1032:
1028:
1024:
1020:
1016:
1009:
1001:
997:
992:
987:
984:(2): 184–98.
983:
979:
975:
968:
960:
956:
952:
946:
942:
941:
933:
925:
921:
914:
907:
901:
893:
889:
885:
879:
875:
874:
866:
855:
848:
840:
834:
830:
829:
821:
813:
806:
794:
786:
784:9780080514048
780:
777:. Amsterdam.
776:
775:
767:
759:
753:
749:
748:
740:
732:
728:
724:
718:
714:
713:
705:
698:
694:
690:
689:
682:
673:
659:
655:
648:
640:
636:
632:
625:
623:
614:
610:
603:
595:
593:9783110106619
589:
585:
580:
579:
571:
563:
557:
553:
552:
544:
536:
530:
526:
519:
503:
501:9780123847201
497:
493:
489:
485:
481:
474:
472:
455:
451:
445:
437:
430:
428:
419:
412:
404:
398:
394:
393:
385:
383:
378:
367:
363:
360:
355:
351:
348:
344:
340:
339:
337:
333:
329:
326:Homoplasy in
325:
320:
317:
314:
311:
308:
304:
301:Synapomorphy/
300:
299:
297:
294:
291:
288:
283:
280:
277:
276:
274:
271:
270:
269:
260:
258:
254:
250:
246:
242:
238:
234:
230:
225:
215:
212:
208:
207:invertebrates
204:
200:
191:
189:
184:
179:
174:
169:
164:
159:
158:Ancient Greek
155:
151:
150:
139:
137:
133:
129:
125:
122:
118:
114:
110:
106:
102:
99:
94:
92:
88:
84:
80:
76:
72:
68:
64:
60:
57:) is a novel
56:
55:derived trait
52:
48:
47:phylogenetics
40:
36:
32:
19:
1644:
1632:
1605:Sister group
1588:Nomenclature
1551:Autapomorphy
1546:Synapomorphy
1526:Plesiomorphy
1514:Group traits
1462:
1334:Cladogenesis
1329:Phylogenesis
1163:
1159:
1149:
1125:(3): 503–7.
1122:
1118:
1108:
1098:ScienceDaily
1097:
1053:
1049:
1043:
1018:
1014:
1008:
981:
977:
967:
943:. Elsevier.
939:
932:
923:
913:
905:
900:
872:
865:
847:
827:
820:
773:
766:
746:
739:
711:
704:
687:
681:
672:
661:. Retrieved
657:
647:
638:
634:
612:
602:
577:
570:
550:
543:
524:
518:
505:. Retrieved
483:
480:"Cladistics"
458:. Retrieved
453:
444:
435:
417:
411:
391:
319:Autapomorphy
312:
306:
295:
279:Plesiomorphy
278:
266:
253:plesiomorphy
249:autapomorphy
229:plesiomorphy
221:
197:
187:
177:
167:
154:Willi Hennig
149:synapomorphy
145:
95:
79:hypothesized
71:synapomorphy
70:
67:plesiomorphy
54:
50:
44:
1600:Crown group
1562:Group types
1293:Systematics
801:|work=
35:Phylogenies
1662:Categories
1278:Cladistics
1219:, Berkeley
1217:Cladistics
731:1039167559
663:2024-06-09
460:16 October
373:References
330:is when a
224:Cladograms
128:amphibians
121:vertebrate
87:cladistics
1615:Supertree
1579:Polyphyly
1574:Paraphyly
1569:Monophyly
1541:Apomorphy
1521:Primitive
1464:PhyloCode
1346:Cladogram
959:173520205
892:950546876
803:ignored (
793:cite book
641:(1): 191.
613:Evolution
507:29 August
436:Evolution
418:Evolution
362:Hemiplasy
354:homoplasy
343:homoplasy
341:Parallel
296:Apomorphy
257:apomorphy
243:but is a
241:tetrapods
146:The word
142:Etymology
136:sprawling
59:character
51:apomorphy
18:Apomorphy
1634:Category
1537:Derived
1283:Taxonomy
1200:29078296
1141:18570042
1086:26845473
1078:21350170
1000:20525577
658:the Node
352:Reverse
303:homology
199:Lampreys
194:Examples
132:reptiles
126:such as
91:homology
1646:Commons
1372:Lineage
1191:5692538
1168:Bibcode
1058:Bibcode
1050:Science
1035:2992286
237:mammals
124:animals
117:mammals
63:evolved
41:states.
1198:
1188:
1139:
1084:
1076:
1033:
998:
957:
947:
890:
880:
835:
781:
754:
729:
719:
695:
590:
558:
531:
498:
399:
203:sharks
188:morphḗ
160:words
111:, and
1610:Basal
1435:UPGMA
1367:Grade
1363:Clade
1082:S2CID
1031:JSTOR
857:(PDF)
332:trait
183:μορφή
98:erect
85:. In
69:). A
49:, an
39:trait
1196:PMID
1137:PMID
1074:PMID
996:PMID
955:OCLC
945:ISBN
888:OCLC
878:ISBN
833:ISBN
812:link
805:help
779:ISBN
752:ISBN
727:OCLC
717:ISBN
693:ISBN
588:ISBN
556:ISBN
529:ISBN
509:2021
496:ISBN
462:2021
397:ISBN
211:jaws
201:and
101:gait
75:taxa
53:(or
1365:vs
1186:PMC
1176:doi
1164:114
1127:doi
1066:doi
1054:331
1023:doi
986:doi
584:366
488:doi
307:its
178:apó
173:ἀπό
168:sún
163:σύν
130:or
115:in
105:fur
45:In
1664::
1194:.
1184:.
1174:.
1162:.
1158:.
1135:.
1123:57
1121:.
1117:.
1096:.
1080:.
1072:.
1064:.
1052:.
1029:.
1019:38
1017:.
994:.
982:58
980:.
976:.
953:.
922:.
886:.
797::
795:}}
791:{{
725:.
656:.
637:.
633:.
621:^
611:.
586:.
494:.
470:^
452:.
426:^
381:^
107:,
103:,
93:.
1246:e
1239:t
1232:v
1202:.
1178::
1170::
1143:.
1129::
1088:.
1068::
1060::
1037:.
1025::
1002:.
988::
961:.
926:.
894:.
841:.
814:)
807:)
787:.
760:.
733:.
699:.
666:.
639:1
596:.
564:.
537:.
513:)
511:.
490::
464:.
405:.
368:.
349:.
186:(
176:(
166:(
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.