509:
258:
36:
533:
212:, a term especially applied in non-biologic contexts. A wide variety of physical/biophysical, chemical/biochemical, and computational methods exist for the study of MA; given the scale (molecular dimensions) of MAs, efforts to elaborate their composition and structure and discern mechanisms underlying their functions are at the forefront of modern structure science.
93:
484:
The images above give an indication of the compositions and scale (dimensions) associated with MAs, though these just begin to touch on the complexity of the structures; in principle, each living cell is composed of MAs, but is itself an MA as well. In the examples and other such complexes and
1163:
bilayers. The hydrophobic hydrocarbon region of the lipid is ~30 Å (3.0 nm) as determined by a combination of neutron and X-ray scattering methods; likewise, the polar/interface region (glyceryl, phosphate, and headgroup moieties, with their combined hydration) is ~15 Å (1.5 nm)
444:, intra- and inter-cellular exchange of material between compartments, etc.). In each of these roles, complex mixtures of become organized in specific structural and spatial ways. While the individual macromolecules are held together by a combination of covalent bonds and
275:
species. Bottom to top: dark blue, repeating FliM and FliN, motor/switch proteins; red, FliG motor/switch proteins; yellow, FliF transmembrane coupling proteins; light blue, L and P ring proteins; and (at top), dark blue, the cap, hook-filament junction, hook, and rod
243:
and other factors involved in light blue, the growing polypeptide chain as a black thread growing vertically from the curve of the mRNA. At end of the animation, the polypeptide produced is extruded through a light blue SecY pore into the gray interior of the
622:
The study of MA structure and function is challenging, in particular because of their megadalton size, but also because of their complex compositions and varying dynamic natures. Most have had standard chemical and biochemical methods applied (methods of
1328:
Snustad DP (August 1968). "Dominance interactions in
Escherichia coli cells mixedly infected with bacteriophage T4D wild-type and amber mutants and their possible implications as to type of gene-product function: catalytic vs. stoichiometric".
675:
molecule and >2000 coat protein molecules). The crystallization and structure solution for the ribosome, MW ~ 2.5 MDa, an example of part of the protein synthetic 'machinery' of living cells, was object of the 2009
614:. Phage T4 encoded proteins that determine virion structure include major structural components, minor structural components and non-structural proteins that catalyze specific steps in the morphogenesis sequence
609:
interact with each other in a characteristic sequence. Maintaining an appropriate balance in the amounts of each of these proteins produced during viral infection appears to be critical for normal phage T4
431:
Complexes of macromolecules occur ubiquitously in nature, where they are involved in the construction of viruses and all living cells. In addition, they play fundamental roles in all basic life processes
161:. They are generally of more than one of these types, and the mixtures are defined spatially (i.e., with regard to their chemical shape), and with regard to their underlying chemical composition and
708:
each have areas that have developed to elaborate and extend the principles first demonstrated in biologic MAs. Of particular interest in these areas has been elaborating the fundamental processes of
200:), and can be in either non-repeating structures (e.g., as in the ribosome (image) and cell membrane architectures), or in repeating linear, circular, spiral, or other patterns (e.g., as in
216:
1905:
493:) at some level of precision. As alluded to in the image legends, when properly prepared, MAs or component subcomplexes of MAs can often be crystallized for study by
1706:
Perino A, Ghigo A, Damilano F, Hirsch E (August 2006). "Identification of the macromolecular complex responsible for PI3Kgamma-dependent regulation of cAMP levels".
1546:"Förster resonance energy transfer - an approach to visualize the spatiotemporal regulation of macromolecular complex formation and compartmentalized cell signaling"
1234:
Hydrocarbon dimensions vary with temperature, mechanical stress, PL structure and coformulants, etc. by single- to low double-digit percentages of these values.
540:, with 30 copies of each of its coat proteins, the small coat protein (S, yellow) and the large coat protein (L, green), which, along with 2 molecules of
405:
1180:"Structure of a fluid dioleoylphosphatidylcholine bilayer determined by joint refinement of x-ray and neutron diffraction data. III. Complete structure"
1933:
424:
into the much larger structures of biomolecular complexes obtained by lower resolution techniques like electron microscopy, electron tomography, and
1842:
Barhoum S, Palit S, Yethiraj A (May 2016). "Diffusion NMR studies of macromolecular complex formation, crowding and confinement in soft materials".
762:
Ban N, Nissen P, Hansen J, Moore PB, Steitz TA (August 2000). "The complete atomic structure of the large ribosomal subunit at 2.4 A resolution".
1051:
Russell RB, Alber F, Aloy P, Davis FP, Korkin D, Pichaud M, et al. (June 2004). "A structural perspective on protein-protein interactions".
489:
in molecular weight (megadaltons, i.e., millions of times the weight of a single, simple atom), though still having measurable component ratios (
558:
were among the first studied MAs; other biologic examples include ribosomes (partial image above), proteasomes, and translation complexes (with
524:). Bilayer/liposome dimensions (obscured in graphic): hydrophobic and polar regions, each ~30 Å (3.0 nm) "thick"—the polar from ~15 Å (1.5 nm)
816:
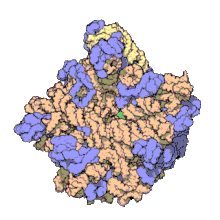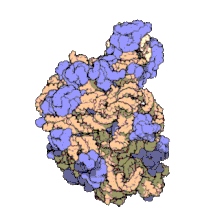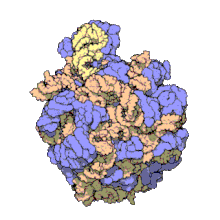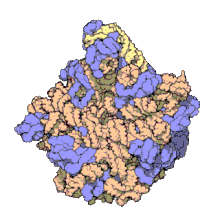
109:. Of the 31 component proteins, 27 are shown (blue), along with its 2 RNA strands (orange/yellow). Scale: assembly is approx. 24 nm across.
739:: the broadest definition of "organelle" includes not only membrane bound cellular structures, but also very large biomolecular complexes.
667:
in
Chemistry for his work on structural elucidation using electron microscopy, in particular for protein-nucleic acid MAs including the
578:
are also generally considered MAs, though the requirement for structural and spatial definition is modified to accommodate the inherent
177:). Assemblies of these can likewise be biologic or non-biologic, though the MA term is more commonly applied in biology, and the term
1875:
Nobel Prizes in
Chemistry (2012), The Nobel Prize in Chemistry 2009, Venkatraman Ramakrishnan, Thomas A. Steitz, Ada E. Yonath,
1799:"Putting the pieces together: integrative modeling platform software for structure determination of macromolecular assemblies"
1926:
1688:
1615:
1596:
1429:
Perrakis A, Musacchio A, Cusack S, Petosa C (August 2011). "Investigating a macromolecular complex: the toolkit of methods".
742:
721:
79:
57:
50:
1919:
652:
1168:, for a total thickness about equal to the hydrocarbon region. See S.H. White references, preceding and following.
1086:
van Dijk AD, Boelens R, Bonvin AM (January 2005). "Data-driven docking for the study of biomolecular complexes".
17:
1876:
648:
1883:
1634:
Valle M (May 2011). "Almost lost in translation. Cryo-EM of a dynamic macromolecular complex: the ribosome".
457:
328:
165:. Macromolecules are found in living and nonliving things, and are composed of many hundreds or thousands of
1797:
Russel D, Lasker K, Webb B, Velázquez-Muriel J, Tjioe E, Schneidman-Duhovny D, et al. (January 2012).
644:
425:
332:
1906:
Dynamics of
Macromolecular Assembly Section | National Institute of Biomedical Imaging and Bioengineering
420:. The atomic structure models obtained by X-ray crystallography and biomolecular NMR spectroscopy can be
2028:
473:
193:
1897:
Beck Group (2019), Structure and function of large macromolecular assemblies (Beck group home page),
541:
449:
841:
Osborne AR, Rapoport TA, van den Berg B (2005). "Protein translocation by the Sec61/SecY channel".
784:
701:
681:
656:
413:
409:
182:
44:
820:
1942:
494:
209:
178:
1746:"Molecular architecture of the 26S proteasome holocomplex determined by an integrative approach"
231:
molecules into proteins. The animation presents the elongation and membrane targeting stages of
1744:
Lasker K, Förster F, Bohn S, Walzthoeni T, Villa E, Unverdorben P, et al. (January 2012).
779:
421:
232:
61:
1671:
Monie TP (2017). "The
Canonical Inflammasome: A Macromolecular Complex Driving Inflammation".
2033:
854:
640:
433:
401:
245:
224:
102:
1364:
508:
1757:
1191:
771:
731:
726:
668:
624:
599:
453:
947:
448:
molecular non-covalent forces (i.e., associations between parts within each molecule, via
257:
8:
1160:
537:
465:
441:
417:
320:
1761:
1195:
775:
1825:
1798:
1780:
1745:
1659:
1570:
1545:
1527:
1502:
1484:
1459:
1270:
1245:
1212:
1179:
1121:
911:
886:
579:
240:
162:
1203:
1978:
1855:
1830:
1785:
1723:
1694:
1684:
1651:
1611:
1592:
1575:
1532:
1489:
1446:
1417:
1346:
1342:
1310:
1306:
1275:
1217:
1113:
1099:
1068:
1033:
992:
951:
916:
858:
797:
709:
379:
371:
114:
1663:
1503:"Studying macromolecular complex stoichiometries by peptide-based mass spectrometry"
1125:
1986:
1847:
1820:
1810:
1775:
1765:
1715:
1676:
1643:
1565:
1557:
1522:
1514:
1479:
1471:
1438:
1409:
1338:
1302:
1293:
Floor E (February 1970). "Interaction of morphogenetic genes of bacteriophage T4".
1265:
1257:
1207:
1199:
1103:
1095:
1060:
1023:
982:
943:
906:
898:
850:
789:
685:
1139:
1982:
1882:
Nobel Prizes in
Chemistry (2012), The Nobel Prize in Chemistry 1982, Aaron Klug,
1815:
1561:
793:
632:
555:
490:
316:
201:
1851:
1680:
1544:
Sinha C, Arora K, Moon CS, Yarlagadda S, Woodrooffe K, Naren AP (October 2014).
1990:
1750:
Proceedings of the
National Academy of Sciences of the United States of America
1261:
987:
970:
705:
628:
186:
154:
150:
1898:
1647:
1475:
1442:
1064:
1028:
1011:
2022:
2002:
1904:
DMA Group (2019), Dynamics of macromolecular assembly (DMA Group home page),
689:
635:
characterization, etc.). In addition, their methods of study include modern
611:
587:
547:(RNA-1 and RNA-2, not visible) constitute the virion. The assembly is highly
486:
461:
437:
348:
197:
170:
158:
138:
106:
1770:
1859:
1834:
1789:
1727:
1698:
1655:
1579:
1536:
1518:
1493:
1450:
1421:
1279:
1117:
1072:
1037:
996:
955:
920:
862:
801:
567:
563:
498:
352:
312:). The interactions between these biomolecules are non-covalent. Examples:
305:
190:
130:
1350:
1314:
1221:
1012:"Large macromolecular complexes in the Protein Data Bank: a status report"
639:
approaches, computational and atomic-resolution structural methods (e.g.,
571:
2006:
1966:
1413:
1400:
Williamson JR (August 2008). "Cooperativity in macromolecular assembly".
927:
902:
677:
664:
575:
517:
513:
393:
367:
261:
146:
92:
1899:
Beck Group - Structure and function of large molecular assemblies - EMBL
878:
1970:
1719:
1108:
887:"POPSCOMP: an automated interaction analysis of biomolecular complexes"
660:
502:
386:
324:
289:
98:
532:
215:
27:
Large chemical complexes composed of polymers and other macromolecules
1994:
1962:
1954:
736:
636:
574:
that allow material passage between cells and cellular compartments.
566:
components), procaryotic and eukaryotic transcription complexes, and
268:
205:
134:
1911:
1974:
548:
363:
356:
265:
239:
subunits in green and yellow, tRNAs in dark blue, proteins such as
236:
220:
174:
142:
962:
712:, and extending known machine designs to new types and processes.
173:; they are often characterized by repeating units (i.e., they are
559:
520:
lipid tails; black and white spheres represent PL polar regions (
497:
and related methods, or studied by other physical methods (e.g.,
464:), by definition MAs themselves are held together solely via the
293:
1796:
105:
model of 29 of the 33 native components, from the laboratory of
700:
Finally, biology is not the sole domain of MAs. The fields of
336:
208:, image). The process by which MAs are formed has been termed
96:
Structure of nucleoprotein MA: The 50S ribosomal subunit from
1958:
1464:
1428:
672:
602:
583:
375:
344:
340:
309:
126:
934:
Moore PB (2012). "How should we think about the ribosome?".
400:
The biomacromolecular complexes are studied structurally by
189:). MAs of macromolecules are held in their defined forms by
1743:
840:
606:
536:
A graphical representation of the structure of a viral MA,
228:
166:
1675:. Subcellular Biochemistry. Vol. 83. pp. 43–73.
1705:
544:
301:
297:
181:
is more often applied in non-biologic contexts (e.g., in
1543:
1460:"So how do you know you have a macromolecular complex?"
1550:
Biochimica et
Biophysica Acta (BBA) - General Subjects
1085:
1050:
1841:
1500:
761:
1844:
Progress in
Nuclear Magnetic Resonance Spectroscopy
884:
551:, and is ~280 Å (28 nm) across at its widest point.
1605:
1586:
605:, the morphogenetic proteins encoded by the phage
288:, is any biological complex made of more than one
512:Cross-sections of phospholipid (PLs) relevant to
2020:
125:) refers to massive chemical structures such as
875:Legend, cover art, J. Bacteriol., October 2006.
843:Annual Review of Cell and Developmental Biology
1927:
1501:Wohlgemuth I, Lenz C, Urlaub H (March 2015).
1003:
1626:
695:
378:. Such complexes in cell nucleus are called
1610:(Fourth ed.). New York: W.H. Freeman.
1321:
1286:
1177:
1009:
834:
485:assemblies, MAs are each often millions of
1934:
1920:
1399:
1824:
1814:
1779:
1769:
1569:
1526:
1483:
1269:
1211:
1107:
1027:
986:
910:
783:
479:
80:Learn how and when to remove this message
1591:(5th ed.). New York: W.H. Freeman.
855:10.1146/annurev.cellbio.21.012704.133214
531:
507:
256:
214:
91:
43:This article includes a list of general
1606:Lehninger AL, Cox M, Nelson DL (2005).
1457:
1327:
885:Kleinjung J, Fraternali F (July 2005).
308:) or large non-polymeric biomolecules (
271:"motor" and partial rod structure of a
252:
235:, showing the mRNA as a black arc, the
14:
2021:
1587:Berg JM, Tymoczko J, Stryer L (2002).
968:
1941:
1915:
1670:
1633:
1292:
1243:
1178:Wiener MC, White SH (February 1992).
1053:Current Opinion in Structural Biology
948:10.1146/annurev-biophys-050511-102314
933:
227:the information content contained in
1608:Lehninger principles of biochemistry
971:"The Complex Macromolecular Complex"
743:Multi-state modeling of biomolecules
722:Multi-state modeling of biomolecules
671:(a structure containing a 6400 base
617:
145:, etc. that are complex mixtures of
29:
1365:"The Nobel Prize in Chemistry 2009"
1142:. Blanco.biomol.uci.edu. 2009-11-10
1140:"Structure of Fluid Lipid Bilayers"
814:
24:
1736:
1392:
1387:
755:
49:it lacks sufficient corresponding
25:
2045:
1891:
1884:The Nobel Prize in Chemistry 1982
1877:The Nobel Prize in Chemistry 2009
1010:Dutta S, Berman HM (March 2005).
593:
1868:
1708:Biochemical Society Transactions
1673:Macromolecular Protein Complexes
1246:"Essay on Biomembrane Structure"
1100:10.1111/j.1742-4658.2004.04473.x
653:transmission electron microscopy
651:(SANS), force spectroscopy, and
34:
1357:
1250:The Journal of Membrane Biology
1237:
1228:
1171:
1153:
1132:
1079:
897:(Web Server issue): W342–W346.
1371:. Nobel Prize Outreach AB 2021
1044:
975:Trends in Biochemical Sciences
869:
808:
649:small-angle neutron scattering
13:
1:
1431:Journal of Structural Biology
1204:10.1016/S0006-3495(92)81849-0
1159:Experimental system, dioleoyl
748:
663:was recognized with the 1982
516:MAs. Yellow-orange indicates
329:DNA polymerase III holoenzyme
1816:10.1371/journal.pbio.1001244
1562:10.1016/j.bbagen.2014.07.015
1343:10.1016/0042-6822(68)90285-7
1307:10.1016/0022-2836(70)90303-7
1295:Journal of Molecular Biology
794:10.1126/science.289.5481.905
645:small-angle X-ray scattering
426:small-angle X-ray scattering
406:NMR spectroscopy of proteins
333:RNA polymerase II holoenzyme
264:model of the structure of a
7:
1852:10.1016/j.pnmrs.2016.01.004
1681:10.1007/978-3-319-46503-6_2
1636:European Biophysics Journal
1458:Dafforn TR (January 2007).
936:Annual Review of Biophysics
715:
474:intermolecular interactions
468:forces, except now exerted
194:intermolecular interactions
10:
2050:
1262:10.1007/s00232-019-00061-w
988:10.1016/j.tibs.2015.11.006
458:dipole–dipole interactions
450:charge-charge interactions
1949:
1648:10.1007/s00249-011-0683-6
1627:Reviews on particular MAs
1476:10.1107/S0907444906047044
1443:10.1016/j.jsb.2011.05.014
1065:10.1016/j.sbi.2004.04.006
1029:10.1016/j.str.2005.01.008
969:Neuman N (January 2016).
696:Non-biologic counterparts
586:, and of proteins within
392:Protein-lipid complexes:
286:biomacromolecular complex
1908:, accessed 13 June 2011.
1901:, accessed 13 June 2011.
1886:, accessed 13 June 2011.
1879:, accessed 13 June 2011.
702:supramolecular chemistry
682:Venkatraman Ramakrishnan
680:in Chemistry awarded to
657:cryo-electron microscopy
600:bacteriophage (phage) T4
414:single particle analysis
410:cryo-electron microscopy
183:supramolecular chemistry
1771:10.1073/pnas.1120559109
1402:Nature Chemical Biology
598:During assembly of the
495:protein crystallography
385:DNA-protein complexes:
362:RNA-protein complexes:
210:molecular self-assembly
179:supramolecular assembly
119:macromolecular assembly
64:more precise citations.
1519:10.1002/pmic.201400466
891:Nucleic Acids Research
817:"50S Ribosome Subunit"
552:
529:
480:MA scales and examples
277:
249:
233:eukaryotic translation
223:, which catalytically
110:
103:X-ray crystallographic
1244:Gerle C (June 2019).
641:X-ray crystallography
570:and other biological
535:
511:
402:X-ray crystallography
339:, chaperonin complex
321:multienzyme complexes
260:
218:
95:
1999:Biomolecular complex
1414:10.1038/nchembio.102
732:Multiprotein complex
727:Quaternary structure
669:tobacco mosaic virus
625:protein purification
454:van der Waals forces
319:, some of which are
282:biomolecular complex
253:Biomolecular complex
1762:2012PNAS..109.1380L
1196:1992BpJ....61..434W
1184:Biophysical Journal
1161:phosphatidylcholine
776:2000Sci...289..905B
538:cowpea mosaic virus
442:vesicle trafficking
434:protein translation
418:electron tomography
157:or other polymeric
1720:10.1042/BST0340502
903:10.1093/nar/gki369
710:molecular machines
580:molecular dynamics
553:
530:
380:ribonucleoproteins
335:, symmetric viral
278:
250:
111:
2029:Molecular biology
2016:
2015:
1943:Hierarchy of life
1690:978-3-319-46501-2
1617:978-0-7167-4339-2
1598:978-0-7167-4955-4
770:(5481): 905–920.
618:Research into MAs
472:molecules (i.e.,
317:Protein complexes
169:held together by
129:and non-biologic
115:molecular biology
90:
89:
82:
16:(Redirected from
2041:
2009:
1936:
1929:
1922:
1913:
1912:
1863:
1838:
1828:
1818:
1793:
1783:
1773:
1756:(5): 1380–1387.
1731:
1702:
1667:
1621:
1602:
1583:
1573:
1540:
1530:
1497:
1487:
1454:
1425:
1381:
1380:
1378:
1376:
1361:
1355:
1354:
1325:
1319:
1318:
1290:
1284:
1283:
1273:
1256:(2–3): 115–130.
1241:
1235:
1232:
1226:
1225:
1215:
1175:
1169:
1157:
1151:
1150:
1148:
1147:
1136:
1130:
1129:
1111:
1088:The FEBS Journal
1083:
1077:
1076:
1048:
1042:
1041:
1031:
1007:
1001:
1000:
990:
966:
960:
959:
931:
925:
924:
914:
882:
876:
873:
867:
866:
838:
832:
831:
829:
828:
819:. Archived from
812:
806:
805:
787:
759:
686:Thomas A. Steitz
556:Virus structures
284:, also called a
85:
78:
74:
71:
65:
60:this article by
51:inline citations
38:
37:
30:
21:
2049:
2048:
2044:
2043:
2042:
2040:
2039:
2038:
2019:
2018:
2017:
2012:
1953:
1945:
1940:
1894:
1889:
1871:
1866:
1846:. 94–95: 1–10.
1809:(1): e1001244.
1739:
1737:Primary sources
1734:
1714:(Pt 4): 502–3.
1691:
1629:
1624:
1618:
1599:
1556:(10): 3067–72.
1513:(5–6): 862–79.
1470:(Pt 1): 17–25.
1395:
1393:General reviews
1390:
1388:Further reading
1385:
1384:
1374:
1372:
1369:The Nobel Prize
1363:
1362:
1358:
1326:
1322:
1291:
1287:
1242:
1238:
1233:
1229:
1176:
1172:
1158:
1154:
1145:
1143:
1138:
1137:
1133:
1084:
1080:
1049:
1045:
1008:
1004:
967:
963:
932:
928:
883:
879:
874:
870:
839:
835:
826:
824:
813:
809:
760:
756:
751:
718:
698:
633:electrochemical
631:, chemical and
620:
596:
491:stoichiometries
482:
412:and successive
255:
206:flagellar motor
202:actin filaments
86:
75:
69:
66:
56:Please help to
55:
39:
35:
28:
23:
22:
18:Biological unit
15:
12:
11:
5:
2047:
2037:
2036:
2031:
2014:
2013:
2011:
2010:
1950:
1947:
1946:
1939:
1938:
1931:
1924:
1916:
1910:
1909:
1902:
1893:
1892:External links
1890:
1888:
1887:
1880:
1872:
1870:
1867:
1865:
1864:
1839:
1794:
1740:
1738:
1735:
1733:
1732:
1703:
1689:
1668:
1630:
1628:
1625:
1623:
1622:
1616:
1603:
1597:
1584:
1541:
1498:
1455:
1426:
1408:(8): 458–465.
1396:
1394:
1391:
1389:
1386:
1383:
1382:
1356:
1320:
1301:(3): 293–306.
1285:
1236:
1227:
1190:(2): 434–447.
1170:
1152:
1131:
1094:(2): 293–312.
1078:
1059:(3): 313–324.
1043:
1022:(3): 381–388.
1002:
961:
926:
877:
868:
833:
807:
785:10.1.1.58.2271
753:
752:
750:
747:
746:
745:
740:
734:
729:
724:
717:
714:
706:nanotechnology
697:
694:
629:centrifugation
619:
616:
595:
594:Virus assembly
592:
588:lipid bilayers
542:positive-sense
481:
478:
462:hydrogen bonds
398:
397:
390:
383:
360:
254:
251:
198:covalent bonds
187:nanotechnology
171:covalent bonds
159:macromolecules
155:polysaccharide
151:polynucleotide
99:H. marismortui
88:
87:
42:
40:
33:
26:
9:
6:
4:
3:
2:
2046:
2035:
2032:
2030:
2027:
2026:
2024:
2008:
2004:
2003:Macromolecule
2000:
1996:
1992:
1988:
1984:
1980:
1976:
1972:
1968:
1964:
1960:
1956:
1952:
1951:
1948:
1944:
1937:
1932:
1930:
1925:
1923:
1918:
1917:
1914:
1907:
1903:
1900:
1896:
1895:
1885:
1881:
1878:
1874:
1873:
1869:Other sources
1861:
1857:
1853:
1849:
1845:
1840:
1836:
1832:
1827:
1822:
1817:
1812:
1808:
1804:
1800:
1795:
1791:
1787:
1782:
1777:
1772:
1767:
1763:
1759:
1755:
1751:
1747:
1742:
1741:
1729:
1725:
1721:
1717:
1713:
1709:
1704:
1700:
1696:
1692:
1686:
1682:
1678:
1674:
1669:
1665:
1661:
1657:
1653:
1649:
1645:
1642:(5): 589–97.
1641:
1637:
1632:
1631:
1619:
1613:
1609:
1604:
1600:
1594:
1590:
1585:
1581:
1577:
1572:
1567:
1563:
1559:
1555:
1551:
1547:
1542:
1538:
1534:
1529:
1524:
1520:
1516:
1512:
1508:
1504:
1499:
1495:
1491:
1486:
1481:
1477:
1473:
1469:
1465:
1461:
1456:
1452:
1448:
1444:
1440:
1437:(2): 106–12.
1436:
1432:
1427:
1423:
1419:
1415:
1411:
1407:
1403:
1398:
1397:
1370:
1366:
1360:
1352:
1348:
1344:
1340:
1337:(4): 550–63.
1336:
1332:
1324:
1316:
1312:
1308:
1304:
1300:
1296:
1289:
1281:
1277:
1272:
1267:
1263:
1259:
1255:
1251:
1247:
1240:
1231:
1223:
1219:
1214:
1209:
1205:
1201:
1197:
1193:
1189:
1185:
1181:
1174:
1167:
1162:
1156:
1141:
1135:
1127:
1123:
1119:
1115:
1110:
1105:
1101:
1097:
1093:
1089:
1082:
1074:
1070:
1066:
1062:
1058:
1054:
1047:
1039:
1035:
1030:
1025:
1021:
1017:
1013:
1006:
998:
994:
989:
984:
980:
976:
972:
965:
957:
953:
949:
945:
941:
937:
930:
922:
918:
913:
908:
904:
900:
896:
892:
888:
881:
872:
864:
860:
856:
852:
848:
844:
837:
823:on 2005-11-24
822:
818:
811:
803:
799:
795:
791:
786:
781:
777:
773:
769:
765:
758:
754:
744:
741:
738:
735:
733:
730:
728:
725:
723:
720:
719:
713:
711:
707:
703:
693:
691:
690:Ada E. Yonath
687:
683:
679:
674:
670:
666:
662:
658:
654:
650:
646:
642:
638:
634:
630:
626:
615:
613:
612:morphogenesis
608:
604:
601:
591:
589:
585:
581:
577:
573:
569:
565:
561:
557:
550:
546:
543:
539:
534:
527:
523:
519:
515:
510:
506:
504:
500:
496:
492:
488:
477:
475:
471:
467:
463:
459:
455:
451:
447:
443:
439:
438:cell division
435:
429:
427:
423:
419:
415:
411:
407:
403:
395:
391:
388:
384:
381:
377:
373:
369:
365:
361:
358:
354:
350:
349:photosystem I
346:
342:
338:
334:
330:
326:
322:
318:
315:
314:
313:
311:
307:
303:
299:
295:
291:
287:
283:
274:
270:
267:
263:
259:
247:
242:
238:
234:
230:
226:
222:
219:A eukaryotic
217:
213:
211:
207:
203:
199:
196:(rather than
195:
192:
188:
184:
180:
176:
172:
168:
164:
160:
156:
152:
148:
144:
140:
136:
132:
131:nanoparticles
128:
124:
120:
116:
108:
107:Thomas Steitz
104:
101:
100:
94:
84:
81:
73:
63:
59:
53:
52:
46:
41:
32:
31:
19:
2034:Biochemistry
2005: >
2001: >
1998:
1997: >
1993: >
1989: >
1985: >
1981: >
1979:Organ system
1977: >
1973: >
1969: >
1965: >
1961: >
1957: >
1843:
1806:
1803:PLOS Biology
1802:
1753:
1749:
1711:
1707:
1672:
1639:
1635:
1607:
1589:Biochemistry
1588:
1553:
1549:
1510:
1506:
1467:
1463:
1434:
1430:
1405:
1401:
1373:. Retrieved
1368:
1359:
1334:
1330:
1323:
1298:
1294:
1288:
1253:
1249:
1239:
1230:
1187:
1183:
1173:
1166:on each side
1165:
1155:
1144:. Retrieved
1134:
1091:
1087:
1081:
1056:
1052:
1046:
1019:
1015:
1005:
978:
974:
964:
939:
935:
929:
894:
890:
880:
871:
846:
842:
836:
825:. Retrieved
821:the original
810:
767:
763:
757:
699:
621:
597:
582:of membrane
576:Biomembranes
564:nucleic acid
554:
526:on each side
525:
521:
499:spectroscopy
483:
469:
445:
430:
399:
353:ATP synthase
306:carbohydrate
285:
281:
279:
272:
191:non-covalent
122:
118:
112:
97:
76:
70:October 2019
67:
48:
2007:Biomolecule
1967:Biocoenosis
1109:1874/336958
942:(1): 1–19.
849:: 529–550.
815:McClure W.
678:Nobel Prize
665:Nobel Prize
647:(SAXS) and
518:hydrophobic
514:biomembrane
466:noncovalent
394:lipoprotein
368:spliceosome
147:polypeptide
133:, cellular
117:, the term
62:introducing
2023:Categories
1971:Population
1507:Proteomics
1146:2019-10-09
981:(1): 1–3.
827:2019-10-09
749:References
661:Aaron Klug
503:microscopy
387:nucleosome
325:proteasome
290:biopolymer
273:Salmonella
262:3D printed
241:elongation
135:organelles
45:references
1995:Organelle
1963:Ecosystem
1955:Biosphere
1016:Structure
780:CiteSeerX
737:Organelle
637:proteomic
549:symmetric
276:proteins.
269:flagellum
266:bacterial
225:translate
163:structure
143:ribosomes
139:membranes
1975:Organism
1860:27247282
1835:22272186
1790:22307589
1728:16856844
1699:28271472
1664:26027815
1656:21336521
1580:25086255
1537:25546807
1494:17164522
1451:21620973
1422:18641626
1331:Virology
1280:30877332
1126:20148856
1118:15654870
1073:15193311
1038:15766539
997:26699226
956:22577819
921:15980485
863:16212506
802:10937989
716:See also
460:such as
364:ribosome
357:ferritin
237:ribosome
221:ribosome
204:and the
175:polymers
1826:3260315
1781:3277140
1758:Bibcode
1571:4151567
1528:5024058
1485:2483502
1351:4878023
1315:4907266
1271:6556169
1222:1547331
1213:1260259
1192:Bibcode
912:1160130
772:Bibcode
764:Science
568:nuclear
560:protein
487:daltons
470:between
382:(RNPs).
337:capsids
294:protein
127:viruses
58:improve
1987:Tissue
1858:
1833:
1823:
1788:
1778:
1726:
1697:
1687:
1662:
1654:
1614:
1595:
1578:
1568:
1535:
1525:
1492:
1482:
1449:
1420:
1375:10 May
1349:
1313:
1278:
1268:
1220:
1210:
1124:
1116:
1071:
1036:
995:
954:
919:
909:
861:
800:
782:
688:, and
603:virion
584:lipids
456:, and
422:docked
416:, and
47:, but
1983:Organ
1959:Biome
1660:S2CID
1122:S2CID
673:ssRNA
607:genes
572:pores
446:intra
376:SnRNP
372:vault
345:GroES
341:GroEL
310:lipid
167:atoms
1991:Cell
1856:PMID
1831:PMID
1786:PMID
1724:PMID
1695:PMID
1685:ISBN
1652:PMID
1612:ISBN
1593:ISBN
1576:PMID
1554:1840
1533:PMID
1490:PMID
1447:PMID
1418:PMID
1377:2021
1347:PMID
1311:PMID
1276:PMID
1218:PMID
1114:PMID
1069:PMID
1034:PMID
993:PMID
952:PMID
917:PMID
859:PMID
798:PMID
704:and
655:and
627:and
562:and
522:v.i.
229:mRNA
185:and
141:and
137:and
1848:doi
1821:PMC
1811:doi
1776:PMC
1766:doi
1754:109
1716:doi
1677:doi
1644:doi
1566:PMC
1558:doi
1523:PMC
1515:doi
1480:PMC
1472:doi
1439:doi
1435:175
1410:doi
1339:doi
1303:doi
1266:PMC
1258:doi
1254:252
1208:PMC
1200:doi
1104:hdl
1096:doi
1092:272
1061:doi
1024:doi
983:doi
944:doi
907:PMC
899:doi
851:doi
790:doi
768:289
643:),
545:RNA
505:).
476:).
302:DNA
298:RNA
113:In
2025::
1854:.
1829:.
1819:.
1807:10
1805:.
1801:.
1784:.
1774:.
1764:.
1752:.
1748:.
1722:.
1712:34
1710:.
1693:.
1683:.
1658:.
1650:.
1640:40
1638:.
1574:.
1564:.
1552:.
1548:.
1531:.
1521:.
1511:15
1509:.
1505:.
1488:.
1478:.
1468:63
1466:.
1462:.
1445:.
1433:.
1416:.
1404:.
1367:.
1345:.
1335:35
1333:.
1309:.
1299:47
1297:.
1274:.
1264:.
1252:.
1248:.
1216:.
1206:.
1198:.
1188:61
1186:.
1182:.
1120:.
1112:.
1102:.
1090:.
1067:.
1057:14
1055:.
1032:.
1020:13
1018:.
1014:.
991:.
979:41
977:.
973:.
950:.
940:41
938:.
915:.
905:.
895:33
893:.
889:.
857:.
847:21
845:.
796:.
788:.
778:.
766:.
692:.
684:,
659:.
590:.
501:,
452:,
440:,
436:,
428:.
408:,
404:,
374:,
370:,
366:,
355:,
351:,
347:,
331:,
327:,
323::
304:,
300:,
296:,
280:A
246:ER
153:,
149:,
123:MA
1935:e
1928:t
1921:v
1862:.
1850::
1837:.
1813::
1792:.
1768::
1760::
1730:.
1718::
1701:.
1679::
1666:.
1646::
1620:.
1601:.
1582:.
1560::
1539:.
1517::
1496:.
1474::
1453:.
1441::
1424:.
1412::
1406:4
1379:.
1353:.
1341::
1317:.
1305::
1282:.
1260::
1224:.
1202::
1194::
1149:.
1128:.
1106::
1098::
1075:.
1063::
1040:.
1026::
999:.
985::
958:.
946::
923:.
901::
865:.
853::
830:.
804:.
792::
774::
528:.
432:(
396:.
389:.
359:.
343:-
292:(
248:.
121:(
83:)
77:(
72:)
68:(
54:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.