20:
117:
1324:
116:
93:
The term protein family has broad usage and can be applied to large groups of proteins with barely detectable sequence similarity as well as narrow groups of proteins with near identical sequence, function, and structure. To distinguish between these cases, a hierarchical terminology is in use. At
1086:
Gerlt, John A.; Allen, Karen N.; Almo, Steven C.; Armstrong, Richard N.; Babbitt, Patricia C.; Cronan, John E.; Dunaway-Mariano, Debra; Imker, Heidi J.; Jacobson, Matthew P.; Minor, Wladek; Poulter, C. Dale; Raushel, Frank M.; Sali, Andrej; Shoichet, Brian K.; Sweedler, Jonathan V. (2011-11-22).
278:
uses protein families and superfamilies as the basis for development of a sequence/structure-based strategy for large scale functional assignment of enzymes of unknown function. The algorithmic means for establishing protein families on a large scale are based on a notion of similarity.
254:
73:
methods. Proteins that do not share a common ancestor are unlikely to show statistically significant sequence similarity, making sequence alignment a powerful tool for identifying the members of protein families. Families are sometimes grouped together into larger
204:
or polarity of the amino-acid residues. Functionally constrained regions of proteins evolve more slowly than unconstrained regions such as surface loops, giving rise to blocks of conserved sequence when the sequences of a protein family are compared (see
193:. Due to evolutionary shuffling, different domains in a protein have evolved independently. This has led to a focus on families of protein domains. Several online resources are devoted to identifying and cataloging these domains.
217:
According to current consensus, protein families arise in two ways. First, the separation of a parent species into two genetically isolated descendant species allows a gene/protein to independently accumulate variations
69:. Sequence similarity (usually amino-acid sequence) is one of the most common indicators of homology, or common evolutionary ancestry. Some frameworks for evaluating the significance of similarity between sequences use
245:
and for protein domains whose hydrophobic amino acids are further from the optimal degree of dispersion along the primary sequence. This expansion and contraction of protein families is one of the salient features of
209:). These blocks are most commonly referred to as motifs, although many other terms are used (blocks, signatures, fingerprints, etc.). Several online resources are devoted to identifying and cataloging protein motifs.
98:, which group distantly related proteins, often based on their structural similarity. Next are protein families, which refer to proteins with a shared evolutionary origin exhibited by significant
270:
analysis, an effort is ongoing to organize proteins into families and to describe their component domains and motifs. Reliable identification of protein families is critical to
408:
85:
Currently, over 60,000 protein families have been defined, although ambiguity in the definition of "protein family" leads different researchers to highly varying numbers.
200:
of an enzyme requires certain amino-acid residues to be precisely oriented. A protein–protein binding interface may consist of a large surface with constraints on the
230:). Because the original gene is still able to perform its function, the duplicated gene is free to diverge and may acquire new functions (by random mutation).
123:
845:
Holm, Liisa; Heger, Andreas (2013). "Automated
Sequence-Based Approaches for Identifying Domain Families". In Orengo, Christine; Bateman, Alex (eds.).
106:
can be defined within families to denote closely related proteins that have similar or identical functions. For example, a superfamily like the
945:
Bateman, Alex (2013). "Sequence
Classification of Protein Families: Pfam and other Resources". In Orengo, Christine; Bateman, Alex (eds.).
322:- Library of HMMs representing superfamilies and database of (superfamily and family) annotations for all completely sequenced organisms
177:
Protein families were first recognised when most proteins that were structurally understood were small, single-domain proteins such as
150:
54:, in which each gene encodes a corresponding protein with a 1:1 relationship. The term "protein family" should not be confused with
274:
analysis, functional annotation, and the exploration of the diversity of protein function in a given phylogenetic branch. The
325:
226:
proteins, usually with conserved sequence motifs. Second, a gene duplication may create a second copy of a gene (termed a
1203:"OrthoFinder: Solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy"
1146:"PASS2 version 4: An update to the database of structure-based sequence alignments of structural domain superfamilies"
1356:
1328:
962:
862:
454:
242:
1640:
669:
Dayhoff, MO; McLaughlin, PJ; Barker, WC; Hunt, LT (1975). "Evolution of sequences within protein superfamilies".
138:
535:"A comprehensive review and comparison of different computational methods for protein remote homology detection"
1650:
1512:
740:
189:. Since then, many proteins have been found with multiple independent structural and functional units called
237:, undergo extreme expansions and contractions in the course of evolution, sometimes in concert with whole
206:
275:
580:
Kunin, Victor; Cases, Ildefonso; Enright, Anton J.; de
Lorenzo, Victor; Ouzounis, Christos A. (2003).
19:
1645:
1613:
1600:
1587:
1574:
1561:
1548:
1535:
1497:
437:
Orengo, Christine; Bateman, Alex (2013). "Introduction". In Orengo, Christine; Bateman, Alex (eds.).
291:
catalog protein families and allow users to match query sequences to known families. These include:
1507:
1461:
1404:
1635:
1409:
639:
257:
Phylogenetic tree of RAS superfamily: This tree was created using FigTree (free online software).
1430:
1349:
756:
389:
340:
127:
99:
95:
1502:
678:
288:
141:). Below, sequence conservation of 70 members of the C04 protease family: Arrows indicate
8:
1466:
376:
299:
79:
1063:
1038:
682:
1399:
1296:
1261:
1237:
1202:
1178:
1121:
1088:
922:
887:
868:
694:
507:
474:
238:
196:
Different regions of a protein have differing functional constraints. For example, the
70:
1014:
979:
822:
787:
616:
581:
1301:
1283:
1242:
1224:
1183:
1165:
1126:
1108:
1068:
1039:"Differential Retention of Pfam Domains Contributes to Long-term Evolutionary Trends"
1019:
1001:
958:
927:
909:
858:
827:
809:
768:
760:
721:
651:
621:
603:
562:
554:
512:
494:
450:
371:
172:
103:
66:
62:
996:
872:
698:
82:
based on structural similarity, even if there is no identifiable sequence homology.
1445:
1440:
1414:
1342:
1291:
1273:
1232:
1214:
1173:
1157:
1116:
1100:
1058:
1050:
1009:
991:
950:
917:
899:
850:
817:
799:
752:
686:
611:
593:
546:
502:
486:
442:
247:
168:
55:
446:
1492:
1476:
1389:
490:
142:
712:
Dayhoff, MO (August 1976). "The origin and evolution of protein superfamilies".
332:- Classifications of protein structures into superfamilies, families and domains
1530:
904:
201:
190:
164:
61:
Proteins in a family descend from a common ancestor and typically have similar
31:
1278:
1219:
954:
854:
1629:
1435:
1394:
1287:
1228:
1169:
1112:
1054:
1005:
913:
813:
764:
607:
558:
498:
598:
534:
266:
As the total number of sequenced proteins increases and interest expands in
110:
of proteases has less sequence conservation than the C04 family within it.
1384:
1305:
1246:
1187:
1130:
1072:
1023:
931:
831:
772:
625:
566:
516:
271:
186:
1161:
980:"Tools and resources for identifying protein families, domains and motifs"
655:
1608:
1543:
1379:
725:
550:
366:
352:
319:
197:
51:
24:
1262:"OrthoFinder: Phylogenetic orthology inference for comparative genomics"
804:
690:
234:
182:
1145:
1104:
1582:
1556:
316:
PASS2 - Protein
Alignment as Structural Superfamilies v2 - PASS2@NCBS
219:
178:
146:
134:
43:
28:
947:
Protein
Families: Relating Protein Sequence, Structure, and Function
847:
Protein
Families: Relating Protein Sequence, Structure, and Function
441:. Hoboken, New Jersey: John Wiley & Sons, Inc. pp. vii–xi.
439:
Protein
Families: Relating Protein Sequence, Structure, and Function
949:. Hoboken, New Jersey: John Wiley & Sons, Inc. pp. 25–36.
533:
Chen, Junjie; Guo, Mingyue; Wang, Xiaolong; Liu, Bin (2018-03-01).
413:
310:
267:
223:
886:
Wang, Yan; Zhang, Hang; Zhong, Haolin; Xue, Zhidong (2021-01-01).
849:. Hoboken, New Jersey: John Wiley & Sons, Inc. pp. 1–24.
336:
Similarly, many database-searching algorithms exist, for example:
1037:
James, Jennifer E; Nelson, Paul G; Masel, Joanna (4 April 2023).
304:
227:
131:
107:
47:
1595:
1365:
1323:
741:"Protein Families and Their Evolution—A Structural Perspective"
475:"An Introduction to Sequence Similarity ("Homology") Searching"
346:
75:
250:, but its importance and ramifications are currently unclear.
1569:
888:"Protein domain identification methods and online resources"
355:- Method for clustering proteins into families (orthogroups)
307:- Database of protein domains, families and functional sites
668:
642:(December 1974). "Computer analysis of protein sequences".
579:
329:
295:
1334:
253:
241:. Expansions are less likely, and losses more likely, for
1143:
261:
785:
1085:
739:
Orengo, Christine A.; Thornton, Janet M. (2005-06-01).
1144:
Gandhimathi, A.; Nair, Anu G.; Sowdhamini, R. (2012).
50:. In many cases, a protein family has a corresponding
788:"Visualizing Sequence Similarity of Protein Families"
222:) in these two lineages. This results in a family of
786:
Veeramachaneni, Vamsi; Makałowski, Wojciech (2004).
409:"What are protein families? Protein classification"
892:Computational and Structural Biotechnology Journal
885:
582:"Myriads of protein families, and still counting"
1627:
1036:
978:Mulder, Nicola J.; Apweiler, Rolf (2001-12-19).
212:
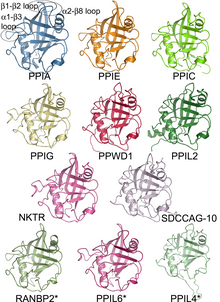
27:family, as represented by the structures of the
977:
738:
532:
298:- Protein families database of alignments and
1350:
468:
466:
436:
432:
430:
233:Certain gene/protein families, especially in
158:
1260:Emms, David M.; Kelly, Steven (2019-11-14).
1201:Emms, David M.; Kelly, Steven (2015-08-06).
282:
1357:
1343:
632:
528:
526:
463:
427:
1295:
1277:
1259:
1236:
1218:
1200:
1177:
1120:
1062:
1013:
995:
921:
903:
844:
821:
803:
705:
615:
597:
506:
757:10.1146/annurev.biochem.74.082803.133029
252:
94:the highest level of classification are
88:
18:
16:Group of evolutionarily-related proteins
944:
711:
638:
523:
472:
1628:
262:Use and importance of protein families
1338:
349:- Protein sequence similarity search
479:Current Protocols in Bioinformatics
383:
313:- SuperFamily Classification System
13:
145:residues, aligned on the basis of
14:
1662:
1316:
243:intrinsically disordered proteins
1322:
1089:"The Enzyme Function Initiative"
343:- DNA sequence similarity search
115:
1253:
1194:
1137:
1079:
1043:Molecular Biology and Evolution
1030:
997:10.1186/gb-2001-3-1-reviews2001
971:
938:
879:
838:
779:
732:
662:
573:
401:
1:
745:Annual Review of Biochemistry
447:10.1002/9781118743089.fmatter
394:
213:Evolution of protein families
65:, functions, and significant
491:10.1002/0471250953.bi0301s42
473:Pearson, William R. (2013).
63:three-dimensional structures
7:
1364:
539:Briefings in Bioinformatics
359:
207:multiple sequence alignment
58:as it is used in taxonomy.
10:
1669:
905:10.1016/j.csbj.2021.01.041
387:
276:Enzyme Function Initiative
162:
159:Protein domains and motifs
1521:
1513:Michaelis–Menten kinetics
1485:
1454:
1423:
1372:
1279:10.1186/s13059-019-1832-y
1220:10.1186/s13059-015-0721-2
955:10.1002/9781118743089.ch2
855:10.1002/9781118743089.ch1
1405:Diffusion-limited enzyme
283:Protein family resources
671:Die Naturwissenschaften
599:10.1186/gb-2003-4-2-401
1641:Protein classification
1150:Nucleic Acids Research
1055:10.1093/molbev/msad073
714:Federation Proceedings
644:Federation Proceedings
258:
130:of 250 members of the
35:
34:of some of its members
1651:Protein superfamilies
1498:Eadie–Hofstee diagram
1431:Allosteric regulation
390:List of gene families
256:
128:sequence conservation
96:protein superfamilies
89:Terminology and usage
22:
1508:Lineweaver–Burk plot
1331:at Wikimedia Commons
990:(1): reviews2001.1.
289:biological databases
1162:10.1093/nar/gkr1096
683:1975NW.....62..154D
377:Sequence clustering
239:genome duplications
100:sequence similarity
67:sequence similarity
1467:Enzyme superfamily
1400:Enzyme promiscuity
805:10.1101/gr.2079204
691:10.1007/BF00608697
551:10.1093/bib/bbw108
259:
71:sequence alignment
36:
1623:
1622:
1327:Media related to
1156:(D1): D531–D534.
1105:10.1021/bi201312u
1099:(46): 9950–9962.
372:Genome annotation
173:Protein structure
1658:
1646:Protein families
1503:Hanes–Woolf plot
1446:Enzyme activator
1441:Enzyme inhibitor
1415:Enzyme catalysis
1359:
1352:
1345:
1336:
1335:
1329:Protein families
1326:
1310:
1309:
1299:
1281:
1257:
1251:
1250:
1240:
1222:
1198:
1192:
1191:
1181:
1141:
1135:
1134:
1124:
1083:
1077:
1076:
1066:
1034:
1028:
1027:
1017:
999:
975:
969:
968:
942:
936:
935:
925:
907:
883:
877:
876:
842:
836:
835:
825:
807:
798:(6): 1160–1169.
783:
777:
776:
736:
730:
729:
709:
703:
702:
666:
660:
659:
636:
630:
629:
619:
601:
577:
571:
570:
530:
521:
520:
510:
470:
461:
460:
434:
425:
424:
422:
421:
405:
384:Protein families
248:genome evolution
169:Structural motif
119:
1668:
1667:
1661:
1660:
1659:
1657:
1656:
1655:
1626:
1625:
1624:
1619:
1531:Oxidoreductases
1517:
1493:Enzyme kinetics
1481:
1477:List of enzymes
1450:
1419:
1390:Catalytic triad
1368:
1363:
1319:
1314:
1313:
1258:
1254:
1199:
1195:
1142:
1138:
1084:
1080:
1035:
1031:
976:
972:
965:
943:
939:
884:
880:
865:
843:
839:
792:Genome Research
784:
780:
737:
733:
710:
706:
667:
663:
637:
633:
578:
574:
531:
524:
485:: 3.1.1–3.1.8.
471:
464:
457:
435:
428:
419:
417:
407:
406:
402:
397:
392:
386:
381:
362:
285:
264:
215:
175:
161:
156:
155:
154:
143:catalytic triad
125:
120:
91:
17:
12:
11:
5:
1666:
1665:
1654:
1653:
1648:
1643:
1638:
1636:Bioinformatics
1621:
1620:
1618:
1617:
1604:
1591:
1578:
1565:
1552:
1539:
1525:
1523:
1519:
1518:
1516:
1515:
1510:
1505:
1500:
1495:
1489:
1487:
1483:
1482:
1480:
1479:
1474:
1469:
1464:
1458:
1456:
1455:Classification
1452:
1451:
1449:
1448:
1443:
1438:
1433:
1427:
1425:
1421:
1420:
1418:
1417:
1412:
1407:
1402:
1397:
1392:
1387:
1382:
1376:
1374:
1370:
1369:
1362:
1361:
1354:
1347:
1339:
1333:
1332:
1318:
1317:External links
1315:
1312:
1311:
1266:Genome Biology
1252:
1207:Genome Biology
1193:
1136:
1078:
1049:(4): msad073.
1029:
984:Genome Biology
970:
963:
937:
878:
863:
837:
778:
751:(1): 867–900.
731:
720:(10): 2132–8.
704:
677:(4): 154–161.
661:
650:(12): 2314–6.
631:
586:Genome Biology
572:
545:(2): 231–244.
522:
462:
455:
426:
399:
398:
396:
393:
388:Main article:
385:
382:
380:
379:
374:
369:
363:
361:
358:
357:
356:
350:
344:
334:
333:
323:
317:
314:
308:
302:
284:
281:
263:
260:
214:
211:
202:hydrophobicity
165:Protein domain
160:
157:
122:
121:
114:
113:
112:
90:
87:
44:evolutionarily
42:is a group of
40:protein family
15:
9:
6:
4:
3:
2:
1664:
1663:
1652:
1649:
1647:
1644:
1642:
1639:
1637:
1634:
1633:
1631:
1615:
1611:
1610:
1605:
1602:
1598:
1597:
1592:
1589:
1585:
1584:
1579:
1576:
1572:
1571:
1566:
1563:
1559:
1558:
1553:
1550:
1546:
1545:
1540:
1537:
1533:
1532:
1527:
1526:
1524:
1520:
1514:
1511:
1509:
1506:
1504:
1501:
1499:
1496:
1494:
1491:
1490:
1488:
1484:
1478:
1475:
1473:
1472:Enzyme family
1470:
1468:
1465:
1463:
1460:
1459:
1457:
1453:
1447:
1444:
1442:
1439:
1437:
1436:Cooperativity
1434:
1432:
1429:
1428:
1426:
1422:
1416:
1413:
1411:
1408:
1406:
1403:
1401:
1398:
1396:
1395:Oxyanion hole
1393:
1391:
1388:
1386:
1383:
1381:
1378:
1377:
1375:
1371:
1367:
1360:
1355:
1353:
1348:
1346:
1341:
1340:
1337:
1330:
1325:
1321:
1320:
1307:
1303:
1298:
1293:
1289:
1285:
1280:
1275:
1271:
1267:
1263:
1256:
1248:
1244:
1239:
1234:
1230:
1226:
1221:
1216:
1212:
1208:
1204:
1197:
1189:
1185:
1180:
1175:
1171:
1167:
1163:
1159:
1155:
1151:
1147:
1140:
1132:
1128:
1123:
1118:
1114:
1110:
1106:
1102:
1098:
1094:
1090:
1082:
1074:
1070:
1065:
1060:
1056:
1052:
1048:
1044:
1040:
1033:
1025:
1021:
1016:
1011:
1007:
1003:
998:
993:
989:
985:
981:
974:
966:
964:9781118743089
960:
956:
952:
948:
941:
933:
929:
924:
919:
915:
911:
906:
901:
898:: 1145–1153.
897:
893:
889:
882:
874:
870:
866:
864:9781118743089
860:
856:
852:
848:
841:
833:
829:
824:
819:
815:
811:
806:
801:
797:
793:
789:
782:
774:
770:
766:
762:
758:
754:
750:
746:
742:
735:
727:
723:
719:
715:
708:
700:
696:
692:
688:
684:
680:
676:
672:
665:
657:
653:
649:
645:
641:
635:
627:
623:
618:
613:
609:
605:
600:
595:
591:
587:
583:
576:
568:
564:
560:
556:
552:
548:
544:
540:
536:
529:
527:
518:
514:
509:
504:
500:
496:
492:
488:
484:
480:
476:
469:
467:
458:
456:9781118743089
452:
448:
444:
440:
433:
431:
416:
415:
410:
404:
400:
391:
378:
375:
373:
370:
368:
365:
364:
354:
351:
348:
345:
342:
339:
338:
337:
331:
327:
324:
321:
318:
315:
312:
309:
306:
303:
301:
297:
294:
293:
292:
290:
280:
277:
273:
269:
255:
251:
249:
244:
240:
236:
231:
229:
225:
221:
210:
208:
203:
199:
194:
192:
188:
184:
180:
174:
170:
166:
152:
148:
144:
140:
136:
133:
129:
124:
118:
111:
109:
105:
101:
97:
86:
83:
81:
80:superfamilies
77:
72:
68:
64:
59:
57:
53:
49:
45:
41:
33:
30:
26:
21:
1609:Translocases
1606:
1593:
1580:
1567:
1554:
1544:Transferases
1541:
1528:
1471:
1385:Binding site
1269:
1265:
1255:
1210:
1206:
1196:
1153:
1149:
1139:
1096:
1093:Biochemistry
1092:
1081:
1046:
1042:
1032:
987:
983:
973:
946:
940:
895:
891:
881:
846:
840:
795:
791:
781:
748:
744:
734:
717:
713:
707:
674:
670:
664:
647:
643:
634:
589:
585:
575:
542:
538:
482:
478:
438:
418:. Retrieved
412:
403:
335:
286:
272:phylogenetic
265:
232:
216:
195:
187:cytochrome c
176:
92:
84:
60:
39:
37:
1380:Active site
640:Dayhoff, MO
367:Gene family
353:OrthoFinder
320:SUPERFAMILY
224:orthologous
198:active site
139:superfamily
104:Subfamilies
52:gene family
25:cyclophilin
1630:Categories
1583:Isomerases
1557:Hydrolases
1424:Regulation
1272:(1): 238.
1213:(1): 157.
592:(2): 401.
420:2023-11-14
395:References
235:eukaryotes
183:hemoglobin
163:See also:
23:The human
1462:EC number
1288:1474-760X
1229:1474-760X
1170:1362-4962
1113:0006-2960
1006:1474-760X
914:2001-0370
814:1088-9051
765:0066-4154
608:1474-760X
559:1477-4054
499:1934-3396
220:mutations
179:myoglobin
147:structure
135:proteases
29:isomerase
1486:Kinetics
1410:Cofactor
1373:Activity
1306:31727128
1247:26243257
1188:22123743
1131:21999478
1073:36947137
1064:10089649
1024:11806833
932:33680357
873:85641264
832:15140831
773:15954844
699:40304076
626:12620116
567:27881430
517:23749753
414:EMBL-EBI
360:See also
268:proteome
48:proteins
46:related
1596:Ligases
1366:Enzymes
1297:6857279
1238:4531804
1179:3245109
1122:3238057
923:7895673
679:Bibcode
656:4435228
508:3820096
305:PROSITE
228:paralog
191:domains
132:PA clan
126:Above,
108:PA clan
78:called
32:domains
1570:Lyases
1304:
1294:
1286:
1245:
1235:
1227:
1186:
1176:
1168:
1129:
1119:
1111:
1071:
1061:
1022:
1015:150457
1012:
1004:
961:
930:
920:
912:
871:
861:
830:
823:419794
820:
812:
771:
763:
726:181273
724:
697:
654:
624:
617:151299
614:
606:
565:
557:
515:
505:
497:
453:
347:BLASTp
185:, and
171:, and
76:clades
56:family
1522:Types
869:S2CID
695:S2CID
341:BLAST
311:PIRSF
287:Many
1614:list
1607:EC7
1601:list
1594:EC6
1588:list
1581:EC5
1575:list
1568:EC4
1562:list
1555:EC3
1549:list
1542:EC2
1536:list
1529:EC1
1302:PMID
1284:ISSN
1243:PMID
1225:ISSN
1184:PMID
1166:ISSN
1127:PMID
1109:ISSN
1069:PMID
1020:PMID
1002:ISSN
959:ISBN
928:PMID
910:ISSN
859:ISBN
828:PMID
810:ISSN
769:PMID
761:ISSN
722:PMID
652:PMID
622:PMID
604:ISSN
563:PMID
555:ISSN
513:PMID
495:ISSN
451:ISBN
330:CATH
328:and
326:SCOP
300:HMMs
296:Pfam
151:DALI
1292:PMC
1274:doi
1233:PMC
1215:doi
1174:PMC
1158:doi
1117:PMC
1101:doi
1059:PMC
1051:doi
1010:PMC
992:doi
951:doi
918:PMC
900:doi
851:doi
818:PMC
800:doi
753:doi
687:doi
612:PMC
594:doi
547:doi
503:PMC
487:doi
443:doi
149:by
1632::
1300:.
1290:.
1282:.
1270:20
1268:.
1264:.
1241:.
1231:.
1223:.
1211:16
1209:.
1205:.
1182:.
1172:.
1164:.
1154:40
1152:.
1148:.
1125:.
1115:.
1107:.
1097:50
1095:.
1091:.
1067:.
1057:.
1047:40
1045:.
1041:.
1018:.
1008:.
1000:.
986:.
982:.
957:.
926:.
916:.
908:.
896:19
894:.
890:.
867:.
857:.
826:.
816:.
808:.
796:14
794:.
790:.
767:.
759:.
749:74
747:.
743:.
718:35
716:.
693:.
685:.
675:62
673:.
648:33
646:.
620:.
610:.
602:.
588:.
584:.
561:.
553:.
543:19
541:.
537:.
525:^
511:.
501:.
493:.
481:.
477:.
465:^
449:.
429:^
411:.
181:,
167:,
102:.
38:A
1616:)
1612:(
1603:)
1599:(
1590:)
1586:(
1577:)
1573:(
1564:)
1560:(
1551:)
1547:(
1538:)
1534:(
1358:e
1351:t
1344:v
1308:.
1276::
1249:.
1217::
1190:.
1160::
1133:.
1103::
1075:.
1053::
1026:.
994::
988:3
967:.
953::
934:.
902::
875:.
853::
834:.
802::
775:.
755::
728:.
701:.
689::
681::
658:.
628:.
596::
590:4
569:.
549::
519:.
489::
483:3
459:.
445::
423:.
218:(
153:.
137:(
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.