427:. Fluorescently tagged DNA, by means of base analogues, permits one to study the local changes of a DNA sequence. Studies on the effects of the length of the primer strand reveal that an equilibrium mixture of four hybridization conformations was observed when template bases looped-out as a bulge, i.e. a structure flanked on both sides by duplex DNA. In contrast, a double-loop structure with an unusual unstacked DNA conformation at its downstream edge was observed when the extruded bases were positioned at the primer–template junction, showing that misalignments can be modified by neighboring DNA secondary structure.
532:
expansion mutations, as frame shifting of the original SCA3 gene product encoding CAG/polyglutamines to GCA/polyalanines. Ribosomal slippage during translation of the SCA3 protein has been proposed as the mechanism resulting in shifting from the polyglutamine to the polyalanine-encoding frame. A dinucleotide deletion or single nucleotide insertion within the polyglutamine tract of huntingtin exon 1 would shift the CAG, polyglutamineen coding frame by +1 (+1 frame shift) to the GCA, polyalanine-encoding frame and introduce a novel epitope to the C terminus of Htt exon 1 (APAAAPAATRPGCG).
436:
332:
583:. As stated previously, frameshift mutations are more likely to occur in a region of repeat sequence. When DNA mismatch repair does not fix the addition or deletion of bases, these mutations are more likely to be pathogenic. This may be in part because the tumor is not told to stop growing. Experiments in yeast and bacteria help to show characteristics of microsatellites that may contribute to defective DNA mismatch repair. These include the length of the
220:
31:
553:
545:
380:
reads the mRNA for protein synthesis, to misinterpret the genetic data. Consequently, an entirely different series of amino acids is generated, resulting in the generation of an altered protein sequence. In most instances, the new reading frame results in an early encounter with a stop codon, leading to the formation of a shortened and usually inactive protein. This form of mutation is termed an early stop codon or a nonsense mutation.
486:(5,958,684) in 1999 by Leeuwen, details the methods and reagents for diagnosis of diseases caused by or associated with a gene having a somatic mutation giving rise to a frameshift mutation. The methods include providing a tissue or fluid sample and conducting gene analysis for frameshift mutation or a protein from this type of mutation. The nucleotide sequence of the suspected gene is provided from published gene sequences or from
363:
376:
errors in DNA.In an unaltered gene, codons (triplets of nucleotides) are sequentially interpreted, with each codon encoding a specific amino acid. This is known as the standard reading frame. However, in cases of frame shift mutations, an extra nucleotide (or more) is inserted into the DNA sequence, disrupting the typical reading frame and causing a shift in the sequence.
135:
491:
amplify the specific region containing the mutation for subsequent analysis.Multiplex
Ligation-dependent Probe Amplification (MLPA): MLPA is a technique used to detect copy number variations and small insertions or deletions.Comparative Genomic Hybridization (CGH): CGH is used to detect chromosomal imbalances, which may include large insertions or deletions.
603:, there is also a genetic component. During testing of coding regions to identify mutations, 116 genetic variants were discovered, including 61 frameshift mutations. There are over 500 mutations on chromosome 17 that seem to play a role in the development of breast and ovarian cancer in the BRCA1 gene, many of which are frameshift.
759:. This process allows for passing over the mutation so that the rest of the sequence remains in frame and the function of the protein stays intact. This, however, does not cure the disease, just treats symptoms, and is only practical in structural proteins or other repetitive genes. A third form of repair is
473:
of only about 1 kilobase. Several technologies are available to perform this test and it is being looked at to be used in clinical applications. When testing for different carcinomas, current methods only allow for looking at one gene at a time. Massively
Parallel Sequencing can test for a variety of
358:
is defined by contiguous triplets. Codons are key to translation of genetic information for the synthesis of proteins. The reading frame is set when translating the mRNA begins and is maintained as it reads one triplet to the next. The reading of the genetic code is subject to three rules the monitor
94:
mutation will in general cause the reading of the codons after the mutation to code for different amino acids. The frameshift mutation will also alter the first stop codon ("UAA", "UGA" or "UAG") encountered in the sequence. The polypeptide being created could be abnormally short or abnormally long,
709:
not previously shown for RAI1 mutation, including hearing loss, absence of self-abusive behaviours, and mild global delays. Sequencing of RAI1 revealed mutation of a heptamericC-tract (CCCCCCC) in exon 3 resulting in frameshift mutations. Of the seven reported frameshift mutations occurring in poly
656:
is one of the cell entry co-factors associated with HIV, most frequently involved with nonsyncytium-inducing strains, is most apparent in HIV patients as opposed to AIDS patients. A 32 base pair deletion in CCR5 has been identified as a mutation that negates the likelihood of an HIV infection. This
499:
Despite the rules that govern the genetic code and the various mechanisms present in a cell to ensure the correct transfer of genetic information during the process of DNA replication as well as during translation, mutations do occur; frameshift mutation is not the only type. There are at least two
634:
conductance regulator (CFTR) gene. There are over 1500 mutations identified, but not all cause the disease. Most cases of cystic fibrosis are a result of the ∆F508 mutation, which deletes the entire amino acid. Two frameshift mutations are of interest in diagnosing CF, CF1213delT and CF1154-insTC.
375:
Frameshift mutations can occur randomly or be caused by an external stimulus. The detection of frameshift mutations can occur via several different methods. Frameshifts are just one type of mutation that can lead to incomplete or incorrect proteins, but they account for a significant percentage of
531:
is one of the nine codon reiteration disorders caused by polyglutamine expansion mutations that include spino-cerebellar ataxia (SCA) 1, 2, 6, 7 and 3, spinobulbar muscular atrophy and dentatorubal-pallidoluysianatrophy. There may be a link between diseases caused by polyglutamine and polyalanine
490:
and sequencing of the suspect gene. The amino acid sequence encoded by the gene is then predicted. NA Sequencing: Sanger sequencing or Next-Generation
Sequencing (NGS) can be used to directly sequence the DNA and identify insertions or deletions.Polymerase Chain Reaction (PCR): PCR can be used to
458:
have been identified through Sanger sequencing that do not overlap with other databases. When a frameshift mutation is observed it is compared against the Human Genome
Mutation Database (HGMD) to determine if the mutation has a damaging effect. This is done by looking at four features. First, the
130:
The information contained in DNA determines protein function in the cells of all organisms. Transcription and translation allow this information to be communicated into making proteins. However, an error in reading this communication can cause protein function to be incorrect and eventually cause
677:
but symptoms do not appear until approximately 6 months of age. There is no cure for the disease. Mutations in the β-hexosaminidase A (Hex A) gene are known to affect the onset of Tay-Sachs, with 78 mutations of different types being described, 67 of which are known to cause disease. Most of the
379:
This insertion prompts a shift in the reading frame due to the triplet nature of the genetic code. For instance, the addition of an extra "A" leads to a sequence shift, triggering the reading of an entirely different set of codons. This deviation in genetic information causes the ribosome, which
540:
Several diseases have frameshift mutations as at least part of the cause. Knowing prevalent mutations can also aid in the diagnosis of the disease. Currently there are attempts to use frameshift mutations beneficially in the treatment of diseases, changing the reading frame of the amino acids.
751:, but this is a highly risky treatment and can often lead to other diseases, such as leukemia. Gene therapy procedures include modifying the zinc fringer nuclease fusion protein, cleaving both ends of the mutation, which in turn removes it from the sequence. Antisense-oligonucleotide mediated
661:
contains a frameshift mutation leading to a premature stop codon. This leads to the loss of the HIV-coreceptor function in vitro. CCR5-1 is considered the wild type and CCR5-2 is considered to be the mutant allele. Those with a heterozygous mutation for the CCR5 were less susceptible to the
259:(protein) has been synthesised and is released. For every 1000 amino acid incorporated into the protein, no more than one is incorrect. This fidelity of codon recognition, maintaining the importance of the proper reading frame, is accomplished by proper base pairing at the ribosome A site,
678:
mutations observed (65/78) are single base substitutions or SNPs, 11 deletions, 1 large and 10 small, and 2 insertions. 8 of the observed mutations are frameshift, 6 deletions and 2 insertions. A 4 base pair insertion in exon 11 is observed in 80% of Tay-Sachs disease presence in the
587:, the makeup of the genetic material and how pure the repeats are. Based on experimental results longer microsatellites have a higher rate of frameshift mutations. The flanking DNA can also contribute to frameshift mutations. In prostate cancer a frameshift mutation changes the
618:
at position 3020. This leads to a premature stop codon, shortening the protein that is supposed to be transcribed. When the protein is able to form normally, it responds to bacterial liposaccharides, where the 3020insC mutation prevents the protein from being responsive.
524:
looked at the difference in frequency of the mutation by both adding and deleting a base pair. It was shown that there was no difference in the frequency between the addition and deletion of a base pair. There is however, a difference in the result of the protein.
786:
A European patent (EP1369126A1) in 2003 by Bork records a method used for prevention of cancers and for the curative treatment of cancers and precancers such as DNA-mismatch repair deficient (MMR) sporadic tumours and HNPCC associated tumours. The idea is to use
936:
Zimmerman PA, Buckler-White A, Alkhatib G, Spalding T, Kubofcik J, Combadiere C, Weissman D, Cohen O, Rubbert A, Lam G, Vaccarezza M, Kennedy PE, Kumaraswami V, Giorgi JV, Detels R, Hunter J, Chopek M, Berger EA, Fauci AS, Nutman TB, Murphy PM (January 1997).
735:) are a rare genetic cause of hypertrophic cardiomyopathy. A recent study has indicated that a frameshift mutation (c.363dupG or p.Gln122AlafsX30) in Troponin C was the cause of hypertrophic cardiomyopathy (and sudden cardiac death) in a 19-year-old male.
698:
involving intellectual disabilities, sleep disturbance, behavioural problems, and a variety of craniofacial, skeletal, and visceral anomalies. The majority of SMS cases harbor an ~3.5 Mb common deletion that encompasses the retinoic acid induced-1
508:. A frameshift mutation can drastically change the coding capacity (genetic information) of the message. Small insertions or deletions (those less than 20 base pairs) make up 24% of mutations that manifest in currently recognized genetic disease.
131:
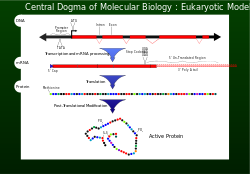
disease even as the cell incorporates a variety of corrective measures.Genetic information is conveyed by DNA for protein synthesis within cells. Misinterpretation can lead to faulty function and disease, despite cellular correction mechanisms.
398:-induced frameshift mutations by DNA polymerases deficient in 3′ → 5′ exonuclease activity was done. The normal sequence 5′ GTC GTT TTA CAA 3′ was changed to GTC GTT T TTA CAA (MIDT) of GTC GTT C TTA CAA (MIDC) to study frameshifts.
682:
Jewish population. The frameshift mutations lead to an early stop codon which is known to play a role in the disease in infants. Delayed onset disease appears to be caused by 4 different mutations, one being a 3 base pair deletion.
1335:
Walsh, T.; Casadei, S.; Lee, M. K.; Pennil, C. C.; Nord, A. S.; Thornton, A. M.; Roeb, W.; Agnew, K. J.; Stray, S. M.; Wickramanayake, A.; Norquist, B.; Pennington, K. P.; Garcia, R. L.; King, M.-C.; Swisher, E. M. (2011).
474:
cancer causing mutations at once as opposed to several specific tests. An experiment to determine the accuracy of this newer sequencing method tested for 21 genes and had no false positive calls for frameshift mutations.
459:
ratio between the affected and conserved DNA, second the location of the mutation relative to the transcript, third the ratio of conserved and affected amino acids and finally the distance of the indel to the end of the
359:
codons in mRNA. First, codons are read in a 5' to 3' direction. Second, codons are nonoverlapping and the message has no gaps. The last rule, as stated above, that the message is translated in a fixed reading frame.
519:
can be run to determine the frequency of the frameshift mutation by adding or removing a pre-set number of nucleotides. Experiments have been run by adding four basepairs, called the +4 experiments, but a team from
1620:
Xu, XiaoLin; Zhu, KaiChang; Liu, Feng; Wang, Yue; Shen, JianGuo; Jin, Jizhong; Wang, Zhong; Chen, Lin; Li, Jiadong; Xu, Min (May 2013). "Identification of somatic mutations in human prostate cancer by RNA-Seq".
939:"Inherited resistance to HIV-1 conferred by an inactivating mutation in CC chemokine receptor 5: studies in populations with contrasting clinical phenotypes, defined racial background, and quantified risk"
106:; in 1997, a frameshift mutation was linked to resistance to infection by the HIV retrovirus. Frameshift mutations have been proposed as a source of biological novelty, as with the alleged creation of
1089:
Sagher, Daphna; Turkington, Edith; Acharya, Sonia; Strauss, Bernard (July 1994). "Production of UV-induced
Frameshift Mutations in Vitro by DNA Polymerases Deficient in 3′ → 5′ Exonuclease Activity".
747:(PID), an inherited condition which can lead to an increase in infections. There are 120 genes and 150 mutations that play a role in primary immunodeficiencies. The standard treatment is currently
1682:
Ogura Y, Bonen DK, Inohara N, Nicolae DL, Chen FF, Ramos R, Britton H, Moran T, Karaliuskas R, Duerr RH, Achkar JP, Brant SR, Bayless TM, Kirschner BS, Hanauer SB, Nuñez G, Cho JH (May 31, 2001).
511:
Frameshift mutations are found to be more common in repeat regions of DNA. A reason for this is because of slipping of the polymerase enzyme in repeat regions, allowing for mutations to enter the
394:
This is a genetic mutation at the level of nucleotide bases. Why and how frameshift mutations occur are continually being sought after. An environmental study, specifically the production of
298:, is thought to be a stronger cause of the occurrence of frameshift mutations. In experiments only 3–13% of all frameshift mutations occurred because of RNA Polymerase II. In
454:
are two methods that have been used to detect frameshift mutations, however, it is likely that data generated will not be of the highest quality. Even still, 1.96 million
791:
with combinatorial mixtures of tumour-specific frameshift mutation-derived peptides to elicit a cytotoxic T-cell response specifically directed against tumour cells.
763:, which is naturally occurring by creating a reverse mutation or a mutation at a second site that corrects the reading frame. This reversion may happen by intragenic
305:
There are several biological processes that help to prevent frameshift mutations. Reverse mutations occur which change the mutated sequence back to the original
729:
in young people, including trained athletes, and is caused by mutations in genes encoding proteins of the cardiac sarcomere. Mutations in the
Troponin C gene (
1656:
313:. This offsets the effect of the original mutation by creating a secondary mutation, shifting the sequence to allow for the correct amino acids to be read.
1954:
Chung WK, Kitner C, Maron BJ (June 2011). "Novel frameshift mutation in
Troponin C ( TNNC1) associated with hypertrophic cardiomyopathy and sudden death".
86:
from the original. The earlier in the sequence the deletion or insertion occurs, the more altered the protein. A frameshift mutation is not the same as a
469:
is a newer method that can be used to detect mutations. Using this method, up to 17 gigabases can be sequenced at once, as opposed to limited ranges for
662:
development of HIV. In a study, despite high exposure to the HIV virus, there was no one homozygous for the CCR5 mutation that tested positive for HIV.
673:
is a fatal disease affecting the central nervous system. It is most frequently found in infants and small children. Disease progression begins in the
1338:"From the Cover: Mutations in 12 genes for inherited ovarian, fallopian tube, and peritoneal carcinoma identified by massively parallel sequencing"
1750:
Farrell PM, Rosenstein BJ, White TB, Accurso FJ, Castellani C, Cutting GR, Durie PR, Legrys VA, Massie J, Parad RB, Rock MJ, Campbell PW (2008).
1395:
Walsh, T.; Lee, M. K.; Casadei, S.; Thornton, A. M.; Stray, S. M.; Pennil, C.; Nord, A. S.; Mandell, J. B.; Swisher, E. M.; King, M.-C. (2010).
247:. The genetic information carried in the codons of the mRNA are now read (decoded) by anticodons of the tRNA. As each codon (triplet) is read,
423:
The effects of neighboring bases and secondary structure to detect the frequency of frameshift mutations has been investigated in depth using
635:
Both of these mutations commonly occur in tandem with at least one other mutation. They both lead to a small decrease in the function of the
710:
C-tracts in RAI1, four cases (~57%) occur at this heptameric C-tract. The results indicate that this heptameric C-tract is a preferential
980:"X-ray crystallographic analysis of 6-aminohexanoate-dimer hydrolase: molecular basis for the birth of a nylon oligomer-degrading enzyme"
772:
1880:
1863:
908:
1799:
Iannuzzi, MC; Stern, RC; Collins, FS; Hon, CT; Hidaka, N; Strong, T; Becker, L; Drumm, ML; White, MB; Gerrard, B (February 1991).
410:
proficient counterparts. The data indicates that loss of proofreading activity increases the frequency of UV-induced frameshifts.
114:(2006) found that a frameshift mutation was unlikely to have been the cause and that rather a two amino acid substitution in the
169:
as the central dogma. For a cell to properly function, proteins are required to be produced accurately for structural and for
2117:
1073:
978:
Negoro S, Ohki T, Shibata N, Mizuno N, Wakitani Y, Tsurukame J, Matsumoto K, Kawamoto I, Takeo M, Higuchi Y (November 2005).
892:
2048:(December 10, 2003) "Use of coding microsatellite region frameshift mutation-derived peptides for treating cancer" by Bork
1752:"Guidelines for Diagnosis of Cystic Fibrosis in Newborns through Older Adults: Cystic Fibrosis Foundation Consensus Report"
317:
can also be used to insert or delete
Uridine into the mRNA after transcription, this allows for the correct reading frame.
17:
1397:"Detection of inherited mutations for breast and ovarian cancer using genomic capture and massively parallel sequencing"
743:
Finding a cure for the diseases caused by frameshift mutations is rare. Research into this is ongoing. One example is a
714:
insertion/deletions (SNindels) and therefore a primary target for analysis in patients suspected for mutations in RAI1.
2241:
152:
2163:— "a central repository for both single base nucleotide substitutions and short deletion and insertion polymorphisms"
1185:"DNA models of trinucleotide frameshift deletions: the formation of loops and bulges at the primer–template junction"
771:
gene conversion, second site DNA slipping or site-specific reversion. This is possible in several diseases, such as
1660:
439:
A deletion mutation alters every codon following it, and can make protein synthesis stop prematurely by forming a
2183:
883:
Losick, Richard; Watson, James D.; Baker, Tania A.; Bell, Stephen; Gann, Alexander; Levine, Michael W. (2008).
103:
87:
1126:"Low-energy circular dichroism of 2-aminopurine dinucleotide as a probe of local conformation of DNA and RNA"
466:
354:. The first codon establishes the reading frame, whereby a new codon begins. A protein's amino acid backbone
776:
2195:
722:
1905:"Frameshift mutation hotspot identified in Smith-Magenis syndrome: case report and review of literature"
1683:
406:
devoid of 3′ → 5′ exonuclease activity produce UV-induced revertants at higher frequency than did their
274:
translation, producing different proteins from overlapping open reading frames, such as the gag-pol-env
2400:
691:
580:
235:. Nucleotides containing the genetic information are now on a single strand messenger template called
2350:
639:
and occur in about 1% of patients tested. These mutations were identified through Sanger sequencing.
2154:
810:
800:
744:
232:
231:
After DNA replication, the reading of a selected section of genetic information is accomplished by
210:
2395:
528:
302:
the error rate inducing frameshift mutations is only somewhere in the range of .0001 and .00001.
1183:
Baase, Walter A.; Davis Jose; Benjamin C. Ponedel; Peter H. von Hippel; Neil P. Johnson (2009).
2234:
2207:
2150:
260:
83:
34:
Different types of indel mutation. Panel C is simply a deletion and not a frameshift mutation.
2355:
2319:
2282:
1475:"Removal of frameshift intermediates by mismatch repair proteins in Saccharomyces cerevisiae"
845:
815:
764:
711:
670:
291:
224:
214:
99:
1999:"New approaches to treatment of primary immunodeficiencies: fixing mutations with chemicals"
2264:
1902:
1698:
1408:
1349:
1137:
726:
2127:
8:
2269:
310:
2390:
2380:
2375:
1702:
1572:
1412:
1353:
1141:
1021:"Host RNA polymerase II makes minimal contributions to retroviral frame-shift mutations"
2023:
1998:
1979:
1931:
1904:
1817:
1800:
1776:
1751:
1732:
1548:
1523:
1431:
1396:
1372:
1337:
1312:
1287:
1263:
1236:
1209:
1184:
1062:
955:
938:
840:
756:
658:
611:
588:
2091:
2066:
1160:
1125:
2421:
2314:
2309:
2227:
2113:
2096:
2028:
1971:
1936:
1885:
1822:
1781:
1736:
1724:
1638:
1602:
1588:
1553:
1504:
1499:
1474:
1436:
1377:
1317:
1268:
1214:
1182:
1165:
1106:
1088:
1069:
1042:
1001:
960:
888:
505:
501:
470:
447:
295:
1983:
435:
2385:
2334:
2086:
2082:
2078:
2046:
2018:
2010:
1963:
1926:
1916:
1875:
1812:
1771:
1763:
1714:
1706:
1630:
1592:
1584:
1543:
1535:
1494:
1486:
1426:
1416:
1367:
1357:
1307:
1299:
1258:
1248:
1204:
1196:
1155:
1145:
1098:
1032:
991:
950:
521:
63:
51:
2324:
2199:
2014:
1903:
Truong, Hoa T; Dudding, Tracy; Blanchard, Christopher L.; Elsea, Sarah H (2010).
1864:"Tay-Sachs disease-causing mutations and neutral polymorphisms in the Hex A gene"
1684:"A frameshift mutation in NOD2 associated with susceptibility to Crohn's disease"
627:
600:
198:
194:
71:
1767:
267:
a form of kinetic stability, and a proofreading mechanism as EF-Tu is released.
2301:
1659:. National Cancer Institute at the National Institute of Health. Archived from
1634:
1303:
835:
780:
584:
451:
174:
70:
in a DNA sequence that is not divisible by three. Due to the triplet nature of
59:
1967:
1454:
599:. While there are environmental factors that contribute to the progression of
2415:
2166:
2160:
1921:
830:
631:
158:
139:
79:
1539:
1421:
1362:
1253:
1150:
2277:
2186:- compare a DNA sequence to a protein sequence database, allowing gaps and
2032:
1975:
1940:
1785:
1728:
1642:
1557:
1508:
1440:
1381:
1321:
1272:
1218:
1169:
1102:
1046:
1005:
996:
979:
783:. There are no drugs or other pharmacogenomic methods that help with PIDs.
424:
326:
186:
2100:
1889:
1826:
1490:
1200:
1110:
1037:
1020:
964:
1719:
1457:(September 28, 1999) "Diagnosis of Neurodegenerative Disease" by Leeuwen
1123:
614:
has an association with the NOD2 gene. The mutation is an insertion of a
407:
347:
299:
256:
248:
190:
115:
2213:
1597:
1524:"Polyalanine and polyserine frameshift products in Huntington's disease"
331:
173:
activities. An incorrectly made protein can have detrimental effects on
2203:
2187:
2174:
1288:"Massively Parallel Sequencing: The Next Big Thing in Genetic Medicine"
572:
516:
440:
351:
275:
252:
219:
181:
to become unhealthy by abnormal cellular functions. To ensure that the
91:
67:
1881:
10.1002/(SICI)1098-1004(1997)9:3<195::AID-HUMU1>3.0.CO;2-7
935:
309:
sequence. Another possibility for mutation correction is the use of a
90:
in which a nucleotide is replaced, rather than inserted or deleted. A
30:
1710:
1606:
1124:
Johnson, Neil P.; Walter A. Baase; Peter H. von Hippel (March 2004).
706:
679:
592:
552:
544:
314:
306:
170:
98:
Frameshift mutations are apparent in severe genetic diseases such as
1613:
2250:
1798:
805:
695:
648:
615:
512:
389:
355:
283:
271:
240:
178:
119:
110:, however, this interpretation is controversial. A study by Negoro
107:
82:(the grouping of the codons), resulting in a completely different
2170:
1840:
1619:
825:
768:
576:
487:
403:
399:
362:
279:
166:
102:; they increase susceptibility to certain cancers and classes of
2192:
2178:
566:
483:
182:
1570:
1468:
1466:
1749:
820:
731:
636:
596:
455:
343:
336:
287:
264:
75:
55:
1060:
Cox, Michael; Nelson, David R.; Lehninger, Albert L (2008).
909:"DNA Is Constantly Changing through the Process of Mutation"
2219:
1463:
701:
674:
653:
595:
from occurring. This leads to an unregulated growth of the
460:
244:
236:
1792:
1571:
Schmoldt, A; Benthe, HF; Haberland, G (1 September 1975).
1564:
887:(6th ed.). San Francisco: Pearson/Benjamin Cummings.
1230:
1228:
162:
2112:(6th ed.). Boston MA: McGraw Hill. pp. 227–8.
1681:
500:
other types of recognized point mutations, specifically
1472:
1394:
977:
395:
134:
1801:"Two frameshift mutations in the cystic fibrosis gene"
1521:
1286:
Tucker, Tracy; Marra, Marco; Friedman, Jan M. (2009).
1225:
705:) gene. Other cases illustrate variability in the SMS
1334:
882:
556:
Frequency of mutations on BRCA2 gene on chromosome 13
548:
Frequency of mutations on BRCA1 gene on chromosome 17
1855:
1285:
2003:Current Opinion in Allergy and Clinical Immunology
1996:
1061:
1059:
165:to a specific amino acid arrangement for making a
1743:
571:Frameshift mutations are known to be a factor in
239:. The mRNA is incorporated with a subunit of the
2413:
1953:
1328:
1237:"Predicting the effects of frameshifting indels"
1012:
931:
929:
255:(UAG, UGA or UAA) is reached. At this point the
204:
1649:
1279:
878:
876:
874:
872:
870:
868:
866:
864:
862:
860:
630:(CF) is a disease based on mutations in the CF
161:described the flow of genetic information from
1990:
1573:"Digitoxin metabolism by rat liver microsomes"
901:
717:
2235:
926:
366:Example of different types of point mutations
177:viability and in most cases cause the higher
2107:
2064:
1947:
1388:
857:
1473:Harfe, BD; Jinks-Robertson, S (July 1999).
383:
320:
78:, the insertion or deletion can change the
2242:
2228:
1843:. National Human Genome Research Institute
2153:at the U.S. National Library of Medicine
2110:Human Genetics: Concepts and Applications
2090:
2022:
1930:
1920:
1879:
1861:
1833:
1816:
1775:
1718:
1596:
1547:
1498:
1430:
1420:
1371:
1361:
1311:
1262:
1252:
1208:
1159:
1149:
1036:
995:
954:
773:X-linked severe combined immunodeficiency
686:
2369:Mutation with respect to overall fitness
2067:"Programmed translational frameshifting"
1234:
551:
543:
434:
361:
330:
218:
185:successfully passes the information on,
133:
95:and will most likely not be functional.
29:
1997:Hu, Hailiang; Gatti, Richard A (2008).
14:
2414:
1522:Davies, J E; Rubinsztein, D C (2006).
1292:The American Journal of Human Genetics
402:pol I Kf and T7 DNA polymerase mutant
2223:
1018:
1064:Lehninger principles of biochemistry
1053:
755:is another possibility for Duchenne
665:
350:, a triplet that code for a certain
270:Frameshifting may also occur during
27:Mutation that shifts codon alignment
278:proteins. This is fairly common in
24:
2294:Mutation with respect to structure
2057:
1841:"Learning About Tay-Sachs Disease"
1805:American Journal of Human Genetics
622:
606:
153:Central dogma of molecular biology
25:
2433:
2144:
1235:Hu, J; Ng, PC (9 February 2012).
657:region on the open reading frame
2216:- Human Genome Mutation Database
2173:against a DNA sequence allowing
146:
2039:
1896:
1675:
1515:
1447:
1068:. San Francisco: W.H. Freeman.
1025:The Journal of General Virology
418:
2083:10.1146/annurev.genet.30.1.507
1479:Molecular and Cellular Biology
1176:
1117:
1082:
971:
104:familial hypercholesterolaemia
88:single-nucleotide polymorphism
13:
1:
885:Molecular biology of the gene
851:
467:Massively Parallel Sequencing
430:
205:Transcription and translation
125:
2249:
2015:10.1097/ACI.0b013e328314b63b
1589:10.1016/0006-2952(75)90094-5
1091:Journal of Molecular Biology
725:is the most common cause of
494:
477:
413:
370:
197:systems are incorporated in
7:
1768:10.1016/j.jpeds.2008.05.005
1528:Journal of Medical Genetics
794:
723:Hypertrophic cardiomyopathy
718:Hypertrophic cardiomyopathy
535:
335:The three letter code, the
10:
2438:
2351:Chromosomal translocations
1635:10.1016/j.gene.2013.01.046
1304:10.1016/j.ajhg.2009.06.022
646:
581:microsatellite instability
564:
387:
324:
208:
150:
2368:
2343:
2300:
2293:
2257:
2202:- tool that compares two
1968:10.1017/S1047951110001927
1756:The Journal of Pediatrics
560:
251:are being joined until a
2155:Medical Subject Headings
1922:10.1186/1471-2350-11-142
1577:Biochemical Pharmacology
1401:Proc Natl Acad Sci U S A
1342:Proc Natl Acad Sci U S A
1130:Proc Natl Acad Sci U S A
1019:Zhang, J (August 2004).
811:Transcription (genetics)
801:Translational frameshift
777:Wiskott–Aldrich syndrome
745:primary immunodeficiency
738:
575:cancer as well as other
384:Genetic or environmental
321:Codon-triplet importance
211:Transcription (genetics)
2391:Nearly neutral mutation
1540:10.1136/jmg.2006.044222
1422:10.1073/pnas.1007983107
1363:10.1073/pnas.1115052108
1254:10.1186/gb-2012-13-2-r9
1151:10.1073/pnas.0400591101
263:hydrolysis activity of
2401:Nonsynonymous mutation
2356:Chromosomal inversions
2258:Mechanisms of mutation
1189:Nucleic Acids Research
1103:10.1006/jmbi.1994.1437
997:10.1074/jbc.m505946200
692:Smith–Magenis syndrome
687:Smith–Magenis syndrome
642:
557:
549:
444:
367:
339:
243:and interacts with an
228:
143:
122:resulted in nylonase.
35:
2381:Advantageous mutation
2320:Conservative mutation
2108:Lewis, Ricki (2005).
2065:Farabaugh PJ (1996).
1862:Myerowitz, R (1997).
1491:10.1128/MCB.19.7.4766
1038:10.1099/vir.0.80081-0
816:Translation (biology)
712:recombination hotspot
555:
547:
438:
365:
334:
292:Reverse transcriptase
222:
215:Translation (biology)
137:
33:
2376:Deleterious mutation
2344:Large-scale mutation
1909:BMC Medical Genetics
529:Huntington's disease
18:Frame-shift mutation
2396:Synonymous mutation
2330:Frameshift mutation
2161:NCBI dbSNP database
2151:Frameshift+Mutation
1703:2001Natur.411..603O
1413:2010PNAS..10712629W
1354:2011PNAS..10818032W
1201:10.1093/nar/gkn1042
1142:2004PNAS..101.3426J
761:revertant mosaicism
694:(SMS) is a complex
591:(ORF) and prevents
311:suppressor mutation
290:(Farabaugh, 1996).
282:and also occurs in
189:mechanisms such as
48:reading frame shift
40:frameshift mutation
2198:2011-07-19 at the
2128:"Nylonase Enzymes"
943:Molecular Medicine
757:muscular dystrophy
589:open reading frame
558:
550:
445:
368:
346:is a set of three
340:
229:
144:
36:
2409:
2408:
2364:
2363:
2315:Missense mutation
2310:Nonsense mutation
2119:978-0-07-111156-0
1657:"Cancer Genomics"
1075:978-0-7167-7108-1
1031:(Pt 8): 2389–95.
894:978-0-8053-9592-1
846:Tay–Sachs disease
671:Tay–Sachs disease
666:Tay–Sachs disease
506:nonsense mutation
502:missense mutation
471:Sanger sequencing
448:Sanger sequencing
296:RNA Polymerase II
100:Tay–Sachs disease
66:) of a number of
16:(Redirected from
2429:
2386:Neutral mutation
2335:Dynamic mutation
2298:
2297:
2244:
2237:
2230:
2221:
2220:
2139:
2137:
2135:
2123:
2104:
2094:
2071:Annu. Rev. Genet
2052:
2045:European Patent
2043:
2037:
2036:
2026:
1994:
1988:
1987:
1951:
1945:
1944:
1934:
1924:
1900:
1894:
1893:
1883:
1859:
1853:
1852:
1850:
1848:
1837:
1831:
1830:
1820:
1796:
1790:
1789:
1779:
1747:
1741:
1740:
1722:
1711:10.1038/35079114
1688:
1679:
1673:
1672:
1670:
1668:
1663:on 18 March 2013
1653:
1647:
1646:
1617:
1611:
1610:
1600:
1568:
1562:
1561:
1551:
1519:
1513:
1512:
1502:
1470:
1461:
1451:
1445:
1444:
1434:
1424:
1407:(28): 12629–33.
1392:
1386:
1385:
1375:
1365:
1332:
1326:
1325:
1315:
1283:
1277:
1276:
1266:
1256:
1232:
1223:
1222:
1212:
1180:
1174:
1173:
1163:
1153:
1121:
1115:
1114:
1086:
1080:
1079:
1067:
1057:
1051:
1050:
1040:
1016:
1010:
1009:
999:
990:(47): 39644–52.
975:
969:
968:
958:
933:
924:
923:
921:
919:
905:
899:
898:
880:
522:Emory University
294:, as opposed to
118:of an ancestral
52:genetic mutation
21:
2437:
2436:
2432:
2431:
2430:
2428:
2427:
2426:
2412:
2411:
2410:
2405:
2360:
2339:
2325:Silent mutation
2289:
2253:
2248:
2206:proteins (back-
2200:Wayback Machine
2147:
2142:
2133:
2131:
2130:. 20 April 2004
2126:
2120:
2060:
2058:Further reading
2055:
2044:
2040:
1995:
1991:
1952:
1948:
1901:
1897:
1860:
1856:
1846:
1844:
1839:
1838:
1834:
1797:
1793:
1748:
1744:
1697:(6837): 603–6.
1686:
1680:
1676:
1666:
1664:
1655:
1654:
1650:
1618:
1614:
1583:(17): 1639–41.
1569:
1565:
1534:(11): 893–896.
1520:
1516:
1471:
1464:
1452:
1448:
1393:
1389:
1348:(44): 18032–7.
1333:
1329:
1284:
1280:
1233:
1226:
1181:
1177:
1136:(10): 3426–31.
1122:
1118:
1087:
1083:
1076:
1058:
1054:
1017:
1013:
976:
972:
934:
927:
917:
915:
907:
906:
902:
895:
881:
858:
854:
841:Crohn's disease
797:
741:
720:
689:
668:
651:
645:
628:Cystic fibrosis
625:
623:Cystic fibrosis
612:Crohn's disease
609:
607:Crohn's disease
601:prostate cancer
569:
563:
538:
497:
480:
433:
421:
416:
392:
386:
373:
329:
323:
217:
209:Main articles:
207:
199:DNA replication
195:mismatch repair
155:
149:
128:
72:gene expression
42:(also called a
28:
23:
22:
15:
12:
11:
5:
2435:
2425:
2424:
2407:
2406:
2404:
2403:
2398:
2393:
2388:
2383:
2378:
2372:
2370:
2366:
2365:
2362:
2361:
2359:
2358:
2353:
2347:
2345:
2341:
2340:
2338:
2337:
2332:
2327:
2322:
2317:
2312:
2306:
2304:
2302:Point mutation
2295:
2291:
2290:
2288:
2287:
2286:
2285:
2280:
2272:
2267:
2261:
2259:
2255:
2254:
2247:
2246:
2239:
2232:
2224:
2218:
2217:
2211:
2190:
2181:
2164:
2158:
2146:
2145:External links
2143:
2141:
2140:
2124:
2118:
2105:
2061:
2059:
2056:
2054:
2053:
2038:
1989:
1946:
1895:
1874:(3): 195–208.
1868:Human Mutation
1854:
1832:
1791:
1742:
1674:
1648:
1612:
1563:
1514:
1485:(7): 4766–73.
1462:
1446:
1387:
1327:
1298:(2): 142–154.
1278:
1241:Genome Biology
1224:
1175:
1116:
1097:(3): 226–242.
1081:
1074:
1052:
1011:
970:
925:
900:
893:
855:
853:
850:
849:
848:
843:
838:
836:point mutation
833:
828:
823:
818:
813:
808:
803:
796:
793:
781:Bloom syndrome
740:
737:
719:
716:
688:
685:
667:
664:
647:Main article:
644:
641:
624:
621:
608:
605:
585:microsatellite
565:Main article:
562:
559:
537:
534:
496:
493:
479:
476:
452:pyrosequencing
432:
429:
420:
417:
415:
412:
388:Main article:
385:
382:
372:
369:
325:Main article:
322:
319:
206:
203:
151:Main article:
148:
145:
127:
124:
26:
9:
6:
4:
3:
2:
2434:
2423:
2420:
2419:
2417:
2402:
2399:
2397:
2394:
2392:
2389:
2387:
2384:
2382:
2379:
2377:
2374:
2373:
2371:
2367:
2357:
2354:
2352:
2349:
2348:
2346:
2342:
2336:
2333:
2331:
2328:
2326:
2323:
2321:
2318:
2316:
2313:
2311:
2308:
2307:
2305:
2303:
2299:
2296:
2292:
2284:
2281:
2279:
2276:
2275:
2274:Substitution
2273:
2271:
2268:
2266:
2263:
2262:
2260:
2256:
2252:
2245:
2240:
2238:
2233:
2231:
2226:
2225:
2222:
2215:
2212:
2209:
2205:
2201:
2197:
2194:
2191:
2189:
2185:
2182:
2180:
2176:
2172:
2168:
2165:
2162:
2159:
2156:
2152:
2149:
2148:
2129:
2125:
2121:
2115:
2111:
2106:
2102:
2098:
2093:
2088:
2084:
2080:
2077:(1): 507–28.
2076:
2072:
2068:
2063:
2062:
2051:
2047:
2042:
2034:
2030:
2025:
2020:
2016:
2012:
2008:
2004:
2000:
1993:
1985:
1981:
1977:
1973:
1969:
1965:
1961:
1957:
1956:Cardiol Young
1950:
1942:
1938:
1933:
1928:
1923:
1918:
1914:
1910:
1906:
1899:
1891:
1887:
1882:
1877:
1873:
1869:
1865:
1858:
1842:
1836:
1828:
1824:
1819:
1814:
1811:(2): 227–31.
1810:
1806:
1802:
1795:
1787:
1783:
1778:
1773:
1769:
1765:
1762:(2): S4–S14.
1761:
1757:
1753:
1746:
1738:
1734:
1730:
1726:
1721:
1720:2027.42/62856
1716:
1712:
1708:
1704:
1700:
1696:
1692:
1685:
1678:
1662:
1658:
1652:
1644:
1640:
1636:
1632:
1628:
1624:
1616:
1608:
1604:
1599:
1594:
1590:
1586:
1582:
1578:
1574:
1567:
1559:
1555:
1550:
1545:
1541:
1537:
1533:
1529:
1525:
1518:
1510:
1506:
1501:
1496:
1492:
1488:
1484:
1480:
1476:
1469:
1467:
1460:
1456:
1450:
1442:
1438:
1433:
1428:
1423:
1418:
1414:
1410:
1406:
1402:
1398:
1391:
1383:
1379:
1374:
1369:
1364:
1359:
1355:
1351:
1347:
1343:
1339:
1331:
1323:
1319:
1314:
1309:
1305:
1301:
1297:
1293:
1289:
1282:
1274:
1270:
1265:
1260:
1255:
1250:
1246:
1242:
1238:
1231:
1229:
1220:
1216:
1211:
1206:
1202:
1198:
1195:(5): 1682–9.
1194:
1190:
1186:
1179:
1171:
1167:
1162:
1157:
1152:
1147:
1143:
1139:
1135:
1131:
1127:
1120:
1112:
1108:
1104:
1100:
1096:
1092:
1085:
1077:
1071:
1066:
1065:
1056:
1048:
1044:
1039:
1034:
1030:
1026:
1022:
1015:
1007:
1003:
998:
993:
989:
985:
981:
974:
966:
962:
957:
952:
948:
944:
940:
932:
930:
914:
910:
904:
896:
890:
886:
879:
877:
875:
873:
871:
869:
867:
865:
863:
861:
856:
847:
844:
842:
839:
837:
834:
832:
831:reading frame
829:
827:
824:
822:
819:
817:
814:
812:
809:
807:
804:
802:
799:
798:
792:
790:
789:immunotherapy
784:
782:
778:
774:
770:
766:
765:recombination
762:
758:
754:
753:exon skipping
750:
746:
736:
734:
733:
728:
724:
715:
713:
708:
704:
703:
697:
693:
684:
681:
676:
672:
663:
660:
655:
650:
640:
638:
633:
632:transmembrane
629:
620:
617:
613:
604:
602:
598:
594:
590:
586:
582:
578:
574:
568:
554:
546:
542:
533:
530:
526:
523:
518:
514:
509:
507:
503:
492:
489:
485:
475:
472:
468:
464:
462:
457:
453:
449:
442:
437:
428:
426:
411:
409:
405:
401:
397:
391:
381:
377:
364:
360:
357:
353:
349:
345:
338:
333:
328:
318:
316:
312:
308:
303:
301:
297:
293:
289:
285:
281:
277:
273:
268:
266:
262:
258:
254:
250:
246:
242:
238:
234:
233:transcription
226:
221:
216:
212:
202:
200:
196:
192:
188:
184:
180:
176:
172:
168:
164:
160:
159:Francis Crick
154:
147:Central dogma
141:
140:central dogma
136:
132:
123:
121:
117:
113:
109:
105:
101:
96:
93:
89:
85:
81:
80:reading frame
77:
73:
69:
65:
61:
57:
53:
49:
45:
44:framing error
41:
32:
19:
2329:
2278:Transversion
2132:. Retrieved
2109:
2074:
2070:
2049:
2041:
2009:(6): 540–6.
2006:
2002:
1992:
1962:(3): 345–8.
1959:
1955:
1949:
1912:
1908:
1898:
1871:
1867:
1857:
1845:. Retrieved
1835:
1808:
1804:
1794:
1759:
1755:
1745:
1694:
1690:
1677:
1665:. Retrieved
1661:the original
1651:
1629:(2): 343–7.
1626:
1622:
1615:
1598:10033/333424
1580:
1576:
1566:
1531:
1527:
1517:
1482:
1478:
1458:
1449:
1404:
1400:
1390:
1345:
1341:
1330:
1295:
1291:
1281:
1244:
1240:
1192:
1188:
1178:
1133:
1129:
1119:
1094:
1090:
1084:
1063:
1055:
1028:
1024:
1014:
987:
983:
973:
949:(1): 23–36.
946:
942:
916:. Retrieved
912:
903:
884:
788:
785:
760:
752:
749:gene therapy
748:
742:
730:
727:sudden death
721:
700:
690:
669:
652:
626:
610:
570:
539:
527:
510:
498:
481:
465:
446:
425:fluorescence
422:
419:Fluorescence
393:
378:
374:
341:
327:Genetic code
304:
269:
230:
191:exonucleases
187:proofreading
156:
129:
111:
97:
47:
43:
39:
37:
2208:translation
2188:frameshifts
2175:frameshifts
2169:- aligns a
984:J Biol Chem
517:Experiments
408:exonuclease
348:nucleotides
300:prokaryotes
257:polypeptide
249:amino acids
225:translation
116:active site
84:translation
68:nucleotides
2283:Transition
2210:principle)
2204:frameshift
1915:(1): 142.
1453:US Patent
852:References
573:colorectal
441:stop codon
431:Sequencing
352:amino acid
276:retroviral
253:stop codon
126:Background
92:frameshift
60:insertions
54:caused by
2265:Insertion
1737:205017657
1455:5,958,684
1247:(2): R9.
707:phenotype
680:Ashkenazi
593:apoptosis
495:Frequency
478:Diagnosis
414:Detection
371:Mechanism
315:Guide RNA
307:wild type
171:catalytic
64:deletions
2422:Mutation
2416:Category
2270:Deletion
2251:Mutation
2196:Archived
2033:18978469
1984:46682245
1976:21262074
1941:20932317
1847:24 March
1786:18639722
1729:11385577
1667:24 March
1643:23434521
1558:16801344
1509:10373526
1441:20616022
1382:22006311
1322:19679224
1273:22322200
1219:19155277
1170:14993592
1047:15269381
1006:16162506
806:Mutation
795:See also
775:(SCID),
696:syndrome
649:HIV/AIDS
616:Cytosine
536:Diseases
513:sequence
390:Mutation
356:sequence
284:bacteria
272:prophase
241:ribosome
179:organism
157:In 1956
120:esterase
108:nylonase
2179:introns
2171:protein
2101:8982463
2024:2686128
1932:2964533
1890:9090523
1827:1990834
1818:1683026
1777:2810958
1699:Bibcode
1549:2563184
1432:2906584
1409:Bibcode
1373:3207658
1350:Bibcode
1313:2725244
1264:3334572
1210:2655659
1138:Bibcode
1111:8028006
965:9132277
956:2230106
826:protein
769:mitotic
577:cancers
488:cloning
404:enzymes
400:E. coli
280:viruses
227:process
167:protein
50:) is a
2157:(MeSH)
2134:2 June
2116:
2099:
2092:239420
2089:
2031:
2021:
1982:
1974:
1939:
1929:
1888:
1825:
1815:
1784:
1774:
1735:
1727:
1691:Nature
1641:
1605:
1556:
1546:
1507:
1497:
1439:
1429:
1380:
1370:
1320:
1310:
1271:
1261:
1217:
1207:
1168:
1161:373478
1158:
1109:
1072:
1045:
1004:
963:
953:
918:17 May
913:Nature
891:
779:, and
567:cancer
561:Cancer
484:patent
456:indels
183:genome
112:et al.
76:codons
56:indels
2184:FastY
2167:Wise2
2050:et al
1980:S2CID
1733:S2CID
1687:(PDF)
1500:84275
1459:et al
821:codon
739:Cures
732:TNNC1
637:lungs
597:tumor
579:with
482:A US
344:codon
337:codon
288:yeast
265:EF-Tu
142:model
46:or a
2214:HGMD
2193:Path
2177:and
2136:2009
2114:ISBN
2097:PMID
2029:PMID
1972:PMID
1937:PMID
1886:PMID
1849:2013
1823:PMID
1782:PMID
1725:PMID
1669:2013
1639:PMID
1623:Gene
1603:PMID
1554:PMID
1505:PMID
1437:PMID
1378:PMID
1318:PMID
1269:PMID
1215:PMID
1166:PMID
1107:PMID
1070:ISBN
1043:PMID
1002:PMID
961:PMID
920:2019
889:ISBN
702:RAI1
675:womb
654:CCR5
504:and
461:exon
450:and
286:and
245:rRNA
237:mRNA
223:The
213:and
193:and
175:cell
138:The
2087:PMC
2079:doi
2019:PMC
2011:doi
1964:doi
1927:PMC
1917:doi
1876:doi
1813:PMC
1772:PMC
1764:doi
1760:153
1715:hdl
1707:doi
1695:411
1631:doi
1627:519
1593:hdl
1585:doi
1544:PMC
1536:doi
1495:PMC
1487:doi
1427:PMC
1417:doi
1405:107
1368:PMC
1358:doi
1346:108
1308:PMC
1300:doi
1259:PMC
1249:doi
1205:PMC
1197:doi
1156:PMC
1146:doi
1134:101
1099:doi
1095:240
1033:doi
992:doi
988:280
951:PMC
659:ORF
643:HIV
261:GTP
163:DNA
74:by
62:or
2418::
2095:.
2085:.
2075:30
2073:.
2069:.
2027:.
2017:.
2005:.
2001:.
1978:.
1970:.
1960:21
1958:.
1935:.
1925:.
1913:11
1911:.
1907:.
1884:.
1870:.
1866:.
1821:.
1809:48
1807:.
1803:.
1780:.
1770:.
1758:.
1754:.
1731:.
1723:.
1713:.
1705:.
1693:.
1689:.
1637:.
1625:.
1607:10
1601:.
1591:.
1581:24
1579:.
1575:.
1552:.
1542:.
1532:43
1530:.
1526:.
1503:.
1493:.
1483:19
1481:.
1477:.
1465:^
1435:.
1425:.
1415:.
1403:.
1399:.
1376:.
1366:.
1356:.
1344:.
1340:.
1316:.
1306:.
1296:85
1294:.
1290:.
1267:.
1257:.
1245:13
1243:.
1239:.
1227:^
1213:.
1203:.
1193:37
1191:.
1187:.
1164:.
1154:.
1144:.
1132:.
1128:.
1105:.
1093:.
1041:.
1029:85
1027:.
1023:.
1000:.
986:.
982:.
959:.
945:.
941:.
928:^
911:.
859:^
767:,
515:.
463:.
396:UV
342:A
201:.
38:A
2243:e
2236:t
2229:v
2138:.
2122:.
2103:.
2081::
2035:.
2013::
2007:8
1986:.
1966::
1943:.
1919::
1892:.
1878::
1872:9
1851:.
1829:.
1788:.
1766::
1739:.
1717::
1709::
1701::
1671:.
1645:.
1633::
1609:.
1595::
1587::
1560:.
1538::
1511:.
1489::
1443:.
1419::
1411::
1384:.
1360::
1352::
1324:.
1302::
1275:.
1251::
1221:.
1199::
1172:.
1148::
1140::
1113:.
1101::
1078:.
1049:.
1035::
1008:.
994::
967:.
947:3
922:.
897:.
699:(
443:.
58:(
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.