266:
412:, play a crucial role in RNA processing, a fundamental step in gene expression. This process involves the precise cleavage of precursor RNA molecules, guided by endonucleases, to generate functional RNAs essential for various cellular functions. Endonucleases selectively cleave precursor RNAs at specific sites, defining the boundaries of functional RNA segments during RNA processing. The outcome of RNA processing is the production of functional RNA molecules, such as
3156:
298:, specifically, catalyzes the incision of DNA exclusively at AP sites, and therefore prepares DNA for subsequent excision, repair synthesis and DNA ligation. For example, when depurination occurs, this lesion leaves a deoxyribose sugar with a missing base. The AP endonuclease recognizes this sugar and essentially cuts the DNA at this site and then allows for DNA repair to continue.
1975:
Budde BS, Namavar Y, Barth PG, Poll-The BT, Nürnberg G, Becker C, van
Ruissen F, Weterman MA, Fluiter K, te Beek ET, Aronica E, van der Knaap MS, Höhne W, Toliat MR, Crow YJ, Steinling M, Voit T, Roelenso F, Brussel W, Brockmann K, Kyllerman M, Boltshauser E, Hammersen G, Willemsen M, Basel-Vanagaite
366:
that catalyzes the initial steps in the repair of these UV-induced thymine dimers. Endonuclease V first cleaves the glycosylic bond on the 5’ side of a pyrimidine dimer and then catalyzes cleavage of the DNA phospodiester bond that originally linked the two nucleotides of the dimer. Subsequent steps
273:
Restriction endonucleases come in several types. A restriction endonuclease typically requires a recognition site and a cleavage pattern (typically of nucleotide bases: A, C, G, T). If the recognition site is outside the region of the cleavage pattern, then the restriction endonuclease is referred to
379:
is activated to initiate controlled cellular disassembly. This disintegration is characterized by the cleavage of genomic DNA into specific fragments. The precise role of endonucleases in this context is to cleave the DNA at specific sites, generating fragments with defined lengths. These fragments
96:
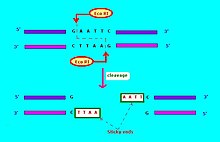
sequence about four to six nucleotides long. Most restriction endonucleases cleave the DNA strand unevenly, leaving complementary single-stranded ends. These ends can reconnect through hybridization and are termed "sticky ends". Once paired, the phosphodiester bonds of the fragments can be joined by
324:
requires multiple incisions in order to disengage the strands and remove the damage. Incisions are required on both sides of the crosslink and on both strands of the duplex DNA. In mouse embryonic stem cells, an intermediate stage of crosslink repair involves production of double-strand breaks.
333:
is a structure specific endonuclease involved in converting interstrand crosslinks to double-strand breaks in a DNA replication-dependent manner. After introduction of a double-strand break, further steps are required to complete the repair process. If a crosslink is not properly repaired it can
149:
activities. Type I can cleave at random sites of about 1000 base pairs or more from the recognition sequence and it requires ATP as source of energy. Type II behaves slightly differently and was first isolated by
Hamilton Smith in 1970. They are simpler versions of the endonucleases and require no
399:
fragment processing. Endonucleases are actively involved in processing these fragments by cleaving the phosphodiester bonds between them. This process is integral to the seamless synthesis and joining of
Okazaki fragments, contributing to the overall continuity of the newly replicated DNA strand.
844:
tRNA splicing endonuclease mutations cause pontocerebellar hypoplasia. Pontocerebellar hypoplasias (PCH) represent a group of neurodegenerative autosomal recessive disorders that is caused by mutations in three of the four different subunits of the tRNA-splicing endonuclease complex.
1679:
Fischer, Heinz; Szabo, Sandra; Scherz, Jennifer; Jaeger, Karin; Rossiter, Heidemarie; Buchberger, Maria; Ghannadan, Minoo; Hermann, Marcela; Theussl, Hans-Christian; Tobin, Desmond J.; Wagner, Erwin F.; Tschachler, Erwin; Eckhart, Leopold (June 2011).
1976:
L, Krägeloh-Mann I, de Vries LS, Sztriha L, Muntoni F, Ferrie CD, Battini R, Hennekam RC, Grillo E, Beemer FA, Stoets LM, Wollnik B, Nürnberg P, Baas F (September 2008). "tRNA splicing endonuclease mutations cause pontocerebellar hypoplasia".
477:
Restriction endonucleases may be found that cleave standard dsDNA (double-stranded DNA), or ssDNA (single-stranded DNA), or even RNA. This discussion is restricted to dsDNA; however, the discussion can be extended to the following:
258:(strain not specified), restriction endonucleases number II and number III, respectively. The restriction enzymes used in molecular biology usually recognize short target sequences of about 4 – 8 base pairs. For instance, the
101:. There are hundreds of restriction endonucleases known, each attacking a different restriction site. The DNA fragments cleaved by the same endonuclease can be joined regardless of the origin of the DNA. Such DNA is called
307:
840:
Sickle Cell anemia is a disease caused by a point mutation. The sequence altered by the mutation eliminates the recognition site for the restriction endonuclease MstII that recognizes the nucleotide sequence.
77:", however, are not limited to either nuclease function, displaying qualities that are both endo- and exo-like. Evidence suggests that endonuclease activity experiences a lag compared to exonuclease activity.
642:
Periplasmic location; average chain length of product is 7; inhibited by tRNA; produces double stranded DNA break; produces nick when complexed with tRNA; endo I mutants grow normally
926:"Differentiation between exonucleases and endonucleases and between haplotomic and diplotomic endonucleases using 3-h-dna-coated wells of plastic depression plates as substrate"
230:. Finally, when a particular type or strain has several different restriction endonucleases, these are identified by Roman numerals, thus, the restriction endonucleases from
524:
In addition, research is now underway to construct synthetic or artificial restriction endonucleases, especially with recognition sites that are unique within a genome.
2637:
431:(ELAC2), shape precursor tRNAs into mature, functional tRNAs, crucial for accurate translation during protein synthesis. In ribosome biogenesis, endonucleases from the
2583:
1430:"The 3′→5′ exonuclease of DNA polymerase δ can substitute for the 5′ flap endonuclease Rad27/Fen1 in processing Okazaki fragments and preventing genome instability"
367:
in the repair process involve removal of the dimer remnants and repair synthesis to fill in the resulting single-strand gap using the undamaged strand as template.
837:
is a rare, autosomal recessive disease caused by a defective UV-specific endonuclease. Patients with mutations are unable to repair DNA damage caused by sunlight.
380:
are then packaged into apoptotic bodies, ensuring a neat and efficient removal of the dying cell without causing inflammation or damage to neighboring cells.
186:" are, in italics, the first letter of the genus and the first two letters of the species where this restriction endonuclease may be found, for example,
302:
cells contain two AP endonucleases: endonuclease IV (endoIV) and exonuclease III (exoIII) while in eukaryotes, there is only one AP endonuclease.
286:
Endonucleases play a role in many aspects of biological life. Below are a couple examples of processes where endonucleases play a crucial role.
2245:
2406:
141:
that relatively contribute to the cleavage of specific sequences. The types I and III are large multisubunit complexes that include both the
113:) are divided into three categories, Type I, Type II, and Type III, according to their mechanism of action. These enzymes are often used in
2063:
2502:
898:
1764:"Isolation and comparison of two molecular species of the BAL 31 nuclease from Alteromonas espejiana with distinct kinetic properties"
2213:
170:
III. Type III, however, cleaves the DNA at about 25 base pairs from the recognition sequence and also requires ATP in the process.
2230:
202:. This is followed by the optional, non-italicized symbol "y", which indicates the type or strain identification, for example,
2477:
811:
388:
1428:
Jin, Yong Hwan; Obert, Robyn; Burgers, Peter M. J.; Kunkel, Thomas A.; Resnick, Michael A.; Gordenin, Dmitry A. (2001-04-24).
88:
that recognize a specific DNA sequence. The nucleotide sequence recognized for cleavage by a restriction enzyme is called the
1959:
1934:
1746:
1277:"The structure-specific endonuclease Mus81-Eme1 promotes conversion of interstrand DNA crosslinks into double-strands breaks"
1215:
1190:
1165:
1030:
986:
461:
also contribute prominently to the removal of DNA during the formation of hair and nails. This process is essential for the
2391:
420:. Endonucleases contribute to the precision of this process, ensuring the formation of mature and functional RNA species.
2492:
2486:
2287:
2037:
1846:"Studies on a nuclease from Ustilago maydis. I. Purification, properties, and implication in recombination of the enzyme"
2335:
1055:
516:, refer to the research by S. Benner, and enlarging the amino acid set in polypeptides, thus enlarging the proteome or
2875:
2446:
2235:
1682:"Essential role of the keratinocyte-specific endonuclease DNase1L2 in the removal of nuclear DNA from hair and nails"
535:
531:
typically cleave in two ways: blunt-ended or sticky-ended patterns. An example of a Type I restriction endonuclease.
625:
Also an exonuclease; nibbles away 3' and 5' ends of duplex DNA. A mixture of at least two nucleases, fast and slow.
612:
Splits -TpC- sequence to yield 5'-dCMP- terminated oligonucleotides; chain length of product varies with conditions
2553:
2441:
2362:
2272:
2225:
274:
as Type I. If the recognition sequence overlaps with the cleavage sequence, then the restriction endonuclease
3031:
2784:
2725:
1488:
2829:
2396:
2386:
2789:
2681:
2306:
1077:
620:
662:
Produces 3'-P termini; requires Ca2+; also acts on RNA; prefers single stranded DNA and AT-rich regions
3146:
2375:
2371:
2367:
2283:
2116:
508:
Synthetic or artificial DNA (for example, containing bases other than A, C, G, T, refer to the work of
972:
306:
3132:
3119:
3106:
3093:
3080:
3067:
3054:
3016:
2543:
2099:
138:
57:
1534:
Hartmann, Roland K.; Gössringer, Markus; Späth, Bettina; Fischer, Susan; Marchfelder, Anita (2009).
1078:"A suggested nomenclature for bacterial host modification and restriction systems and their enzymes"
3026:
2980:
2923:
2291:
2140:
2054:
859:
462:
439:, play a role in processing precursor rRNAs, contributing to the assembly of functional ribosomes.
2928:
2160:
2030:
376:
1633:
1022:
1015:
55:, cut DNA relatively nonspecifically (without regard to sequence), while many, typically called
2716:
2355:
1887:"Purification and characterization of HeLa endonuclease R. A G-specific mammalian endonuclease"
150:
ATP in their degradation processes. Some examples of type II restriction endonucleases include
1375:"Endonuclease Activation and Chromosomal DNA Fragmentation during Apoptosis in Leukemia Cells"
1233:"Structure and function of nucleases in DNA repair: shape, grip and blade of the DNA scissors"
2949:
2868:
2651:
2548:
2340:
2301:
2165:
2087:
1504:
1275:
Hanada, K.; Budzowska, M.; Modesti, M.; Maas, A.; Wyman, C.; Essers, J.; Kanaar, R. (2006).
465:
of hair and nail structures and is crucial for the transformation of cells into durable and
449:
also from the RNase III family play a role in the processing pre-miRNA to functional miRNA.
3021:
2834:
2669:
2664:
2597:
2257:
2082:
1632:
Kuehbacher, Angelika; Urbich, Carmen; Zeiher, Andreas M.; Dimmeler, Stefanie (2007-07-06).
834:
647:
347:
8:
2985:
2689:
2659:
2463:
2458:
2382:
2318:
2104:
540:
528:
495:
321:
114:
52:
36:
679:
Cleavage next to AP site; also a 3'→5' exonuclease; phosphomonoesterase on 3'-P termini
2918:
2822:
2674:
2155:
2145:
2023:
2002:
1714:
1681:
1609:
1582:
1410:
1301:
1276:
780:
Average chain length of product is 4; produces double strand break in presence of Mn2+
743:
275:
110:
1903:
1886:
1862:
1845:
1821:
1804:
1780:
1551:
1350:
1325:
1133:
950:
925:
2623:
2568:
2536:
2411:
2130:
2006:
1994:
1955:
1930:
1908:
1867:
1826:
1785:
1742:
1719:
1701:
1661:
1653:
1614:
1563:
1555:
1516:
1508:
1469:
1464:
1451:
1429:
1402:
1394:
1355:
1306:
1254:
1211:
1186:
1161:
1138:
1097:
1093:
1051:
1026:
982:
973:
Stephen T. Kilpatrick; Jocelyn E. Krebs; Lewin, Benjamin; Goldstein, Elliott (2011).
955:
820:
490:
396:
395:
on the lagging strand, participating in crucial processes such as primer removal and
122:
20:
1414:
578:
Partially ATP dependent; also an exonuclease; functions in recombination and repair
2964:
2959:
2933:
2861:
2699:
2279:
2184:
2179:
2135:
1986:
1898:
1857:
1816:
1775:
1709:
1693:
1645:
1604:
1594:
1547:
1500:
1459:
1441:
1386:
1345:
1337:
1296:
1288:
1244:
1128:
1089:
975:
945:
937:
595:
Essential for replication; preference for single stranded over double stranded DNA
505:
Double-stranded hybrids of DNA and RNA (one strand is DNA, the other strand is RNA)
432:
409:
355:
1649:
1599:
3176:
3011:
2995:
2908:
2801:
2615:
2531:
2526:
2521:
2434:
2429:
2189:
2067:
874:
785:
392:
335:
295:
118:
102:
2015:
3160:
3049:
2990:
2774:
2769:
2764:
2150:
2125:
2121:
2094:
2077:
1763:
1634:"Role of Dicer and Drosha for Endothelial MicroRNA Expression and Angiogenesis"
1434:
Proceedings of the
National Academy of Sciences of the United States of America
1341:
428:
424:
363:
65:, cleave only at very specific nucleotide sequences. Endonucleases differ from
40:
1181:
Ellenberger T, Friedberg EC, Walker GS, Wolfram S, Wood RJ, Schultz R (2006).
1116:
3170:
2954:
2913:
2515:
2424:
2174:
1705:
1657:
1559:
1512:
1455:
1398:
1292:
417:
142:
32:
1535:
265:
2903:
2573:
2296:
2217:
1998:
1723:
1665:
1618:
1567:
1520:
1473:
1446:
1406:
1310:
1258:
1249:
1232:
869:
509:
499:
413:
1912:
1871:
1830:
1789:
1359:
1101:
941:
3127:
3062:
2898:
2817:
2588:
2482:
2345:
2311:
2249:
1978:
1142:
959:
854:
770:
728:
711:
706:
669:
652:
632:
602:
585:
351:
178:
The commonly used notation for restriction endonucleases is of the form "
93:
66:
1697:
69:, which cleave the ends of recognition sequences instead of the middle (
2794:
1390:
517:
317:
121:
for introduction into bacterial, plant, or animal cells, as well as in
98:
81:
1581:
Lejars, Maxence; Kobayashi, Asaki; Hajnsdorf, Eliane (December 2021).
3101:
3075:
2707:
2506:
2419:
2046:
1374:
801:
552:
Below are tables of common prokaryotic and eukaryotic endonucleases.
146:
3155:
2752:
2747:
2742:
2563:
2198:
2050:
1990:
1762:
Wei, CF; Alianell, GA; Bencen, GH; Gray HB, Jr (25 November 1983).
864:
469:
structures, ensuring the strength and integrity of hair and nails.
458:
1954:. Philadelphia: Wolters Kluwer/Lippincott Williams & Wilkins.
2759:
2737:
2732:
2350:
1180:
1156:
Losick R, Watson JD, Baker TA, Bell S, Gann S, Levine MW (2008).
879:
466:
85:
262:
RI enzyme recognizes and cleaves the sequence 5' – GAATTC – 3'.
3114:
2884:
2779:
2712:
2605:
2203:
2111:
569:
513:
446:
436:
28:
1736:
1631:
1533:
3088:
2721:
2472:
2468:
442:
326:
1489:"Flap Endonuclease 1: A Central Component of DNA Metabolism"
1373:
Yoshida, Akira; Pommier, Yves; Ueda, Takanori (2006-02-01).
105:; DNA formed by the joining of genes into new combinations.
2453:
2330:
2323:
2267:
2262:
1008:
1006:
1004:
1002:
1000:
998:
330:
210:
strains bearing the drug resistance transfer factor RTF-1,
126:
2853:
1274:
1071:
1069:
1067:
1974:
48:
44:
1805:"An endonuclease from mitochondria of Neurospora crassa"
1761:
1678:
1580:
995:
320:
in which the two complementary strands are joined by an
281:
1540:
Progress in
Molecular Biology and Translational Science
1487:
Liu, Yuan; Kao, Hui-I; Bambara, Robert A. (June 2004).
1064:
3144:
1536:"The making of tRNAs and more - RNase P and tRNase Z"
1427:
1155:
1949:
1372:
1014:
1012:
974:
2341:Fructose 6-P,2-kinase:fructose 2,6-bisphosphatase
2045:
1943:
1919:
1199:
3168:
1884:
1843:
1174:
1149:
1048:Emergent computation: Emphasizing Bioinformatics
452:
1326:"Deoxyribonucleic acid repair in bacteriophage"
1230:
1844:Holloman, WK; Holliday, R (10 December 1973).
899:"Properties of Exonucleases and Endonucleases"
2869:
2031:
1486:
1270:
1268:
793:Involved in DNA Base Excision Repair pathway
1968:
1224:
1160:. San Francisco: Pearson/Benjamin Cummings.
1114:
1075:
1583:"RNase III, Ribosome Biogenesis and Beyond"
80:Restriction enzymes are endonucleases from
2876:
2862:
2038:
2024:
1802:
1265:
1076:Smith, HO; Nathans, D (15 December 1973).
137:Ultimately, there are three categories of
125:. One of the more famous endonucleases is
92:. Typically, a restriction site will be a
1950:Ferrier DR, Champe PC, Harvey RP (2008).
1902:
1861:
1820:
1779:
1737:Tania A. Baker; Kornberg, Arthur (2005).
1713:
1608:
1598:
1463:
1445:
1349:
1323:
1300:
1248:
1205:
1132:
1115:Rubin, RA; Modrich, P (25 October 1977).
1039:
949:
375:During apoptosis, Apoptotic endonuclease
294:Endonucleases play a role in DNA repair.
1885:Gottlieb, J; Muzyczka, N (5 July 1990).
1686:The Journal of Investigative Dermatology
1505:10.1146/annurev.biochem.73.012803.092453
1021:. San Francisco: W.H. Freeman. pp.
264:
2231:Ubiquitin carboxy-terminal hydrolase L1
1231:Nishino T, Morikawa K (December 2002).
1045:
1013:Cox M, Nelson DR, Lehninger AL (2005).
547:
341:
312:
16:Enzymes which cleave a nucleotide chain
3169:
544:, which act on dsDNA, ssDNA, and RNA.
391:and Dna2 endonuclease are integral to
2857:
2811:either deoxy- or ribo-
2019:
472:
282:Processes involved with endonucleases
2392:Protein serine/threonine phosphatase
1803:Linn, S; Lehman, IR (10 June 1966).
1017:Lehninger principles of biochemistry
923:
2493:Cyclic nucleotide phosphodiesterase
2487:Clostridium perfringens alpha toxin
2288:Tartrate-resistant acid phosphatase
1891:The Journal of Biological Chemistry
1850:The Journal of Biological Chemistry
1809:The Journal of Biological Chemistry
1768:The Journal of Biological Chemistry
1379:International Journal of Hematology
1121:The Journal of Biological Chemistry
1050:. New York: Springer. p. 437.
667:Endonuclease II (endo VI, exo III;
13:
2336:Pyruvate dehydrogenase phosphatase
538:, such as those that are found in
536:DNA/RNA non-specific endonucleases
383:
73:) portion. Some enzymes known as "
14:
3188:
2236:4-hydroxybenzoyl-CoA thioesterase
823:fragments during DNA replication
520:, see the research by P. Schultz.
408:Endonucleases, more specifically
403:
3154:
1927:Medical Biochemistry at a Glance
358:in the phage DNA. The phage T4
305:
2554:N-acetylglucosamine-6-sulfatase
2442:Sphingomyelin phosphodiesterase
1878:
1837:
1796:
1755:
1730:
1672:
1625:
1574:
1527:
1480:
1421:
1366:
1317:
924:Slor, Hanoch (April 14, 1975).
698:Neurospora crassa, mitochondria
2363:Inositol-phosphate phosphatase
2226:Palmitoyl protein thioesterase
1185:. Washington, D.C: ASM Press.
1108:
981:. Boston: Jones and Bartlett.
966:
917:
891:
322:interstrand covalent crosslink
1:
2726:RNA-induced silencing complex
1904:10.1016/S0021-9258(19)38522-9
1863:10.1016/S0021-9258(19)43199-2
1822:10.1016/S0021-9258(18)96595-6
1781:10.1016/S0021-9258(17)43942-1
1650:10.1161/CIRCRESAHA.107.153916
1600:10.3390/microorganisms9122608
1552:10.1016/S0079-6603(08)00808-8
1493:Annual Review of Biochemistry
1210:. New York: Garland Science.
1208:Molecular biology of the cell
1158:Molecular biology of the gene
1134:10.1016/S0021-9258(19)66964-4
885:
527:Restriction endonucleases or
453:Maturation of Nails and Hairs
289:
246:dIII, etc. Another example: "
132:
2830:Serratia marcescens nuclease
2397:Dual-specificity phosphatase
2387:Protein tyrosine phosphatase
1094:10.1016/0022-2836(73)90152-6
1082:Journal of Molecular Biology
829:
370:
173:
7:
2883:
2307:Fructose 1,6-bisphosphatase
848:
819:Responsible for processing
757:Ustilago nuclease (Dnase I)
512:). Research with synthetic
10:
3193:
1342:10.1128/mr.45.1.72-98.1981
1183:DNA repair and mutagenesis
498:, quadruple-stranded DNA (
389:Flap endonuclease 1 (FEN1)
3040:
3032:Michaelis–Menten kinetics
3004:
2973:
2942:
2891:
2810:
2698:
2650:
2636:
2614:
2596:
2582:
2562:
2544:Galactosamine-6 sulfatase
2501:
2405:
2244:
2212:
2100:6-phosphogluconolactonase
2062:
1929:. New York: Wiley. 2012.
534:Furthermore, there exist
269:Restriction enzyme Eco RI
139:restriction endonucleases
107:Restriction endonucleases
58:restriction endonucleases
2924:Diffusion-limited enzyme
2292:Purple acid phosphatases
1293:10.1038/sj.emboj.7601344
860:Restriction endonuclease
630:Endonuclease I (endo I;
348:bacteriophage (phage) T4
254:III" refer to bacterium
1330:Microbiological Reviews
695:Neurospora endonuclease
2717:Microprocessor complex
2356:Beta-propeller phytase
1741:. University Science.
1447:10.1073/pnas.091095198
1324:Bernstein, C. (1981).
1250:10.1038/sj.onc.1206135
930:Nucleic Acids Research
806:Specific for GC sites
418:ribosomal RNAs (rRNAs)
270:
196:Haemophilus influenzae
3017:Eadie–Hofstee diagram
2950:Allosteric regulation
2652:Endodeoxyribonuclease
2549:Iduronate-2-sulfatase
2302:Glucose 6-phosphatase
2088:Butyrylcholinesterase
790:Nucleus, mitochondria
414:transfer RNAs (tRNAs)
268:
256:Haemophilus aegyptius
3027:Lineweaver–Burk plot
2835:Micrococcal nuclease
2670:Deoxyribonuclease IV
2665:Deoxyribonuclease II
2598:Exodeoxyribonuclease
2258:Alkaline phosphatase
2083:Acetylcholinesterase
1638:Circulation Research
835:Xeroderma pigmentosa
735:Penicillium citrinum
648:Micrococcal nuclease
600:T4 endonuclease II (
548:Common endonucleases
354:irradiation induces
342:Thymine dimer repair
313:DNA crosslink repair
2690:UvrABC endonuclease
2660:Deoxyribonuclease I
2383:Protein phosphatase
2319:Protein phosphatase
2117:Bile salt-dependent
2105:PAF acetylhydrolase
1698:10.1038/jid.2011.13
942:10.1093/nar/2.6.897
903:New England BioLabs
617:Bal 31 endonuclease
541:Serratia marcescens
529:restriction enzymes
496:Triple-stranded DNA
423:Endonucleases like
234:strain d are named
115:genetic engineering
111:restriction enzymes
63:restriction enzymes
53:deoxyribonuclease I
37:phosphodiester bond
2986:Enzyme superfamily
2919:Enzyme promiscuity
2823:Mung bean nuclease
2682:Restriction enzyme
2675:Restriction enzyme
1391:10.1007/BF03342699
1206:Alberts B (2002).
744:Mung bean nuclease
718:Aspergillus oryzae
701:Also acts on RNA.
558:Prokaryotic Enzyme
491:Holliday junctions
473:Further discussion
276:restriction enzyme
271:
3142:
3141:
2851:
2850:
2847:
2846:
2843:
2842:
2632:
2631:
2624:Oligonucleotidase
2569:deoxyribonuclease
2537:Steroid sulfatase
2412:Phosphodiesterase
2141:Hormone-sensitive
1961:978-0-7817-6960-0
1936:978-0-470-65451-4
1748:978-1-891389-44-3
1287:(20): 4921–4932.
1217:978-0-8153-3218-3
1192:978-1-55581-319-2
1167:978-0-8053-9592-1
1117:"EcoRI methylase"
1032:978-0-7167-4339-2
988:978-0-7637-6632-0
827:
826:
763:Also acts on RNA
752:Also acts on RNA
749:mung bean sprouts
738:Also acts on RNA
721:Also acts on RNA
684:Eukaryotic Enzyme
592:phage T7 (gene 3)
583:T7 endonuclease (
457:The endonuclease
123:synthetic biology
75:exo-endonucleases
51:). Some, such as
21:molecular biology
3184:
3159:
3158:
3150:
3022:Hanes–Woolf plot
2965:Enzyme activator
2960:Enzyme inhibitor
2934:Enzyme catalysis
2878:
2871:
2864:
2855:
2854:
2700:Endoribonuclease
2686:
2680:
2648:
2647:
2594:
2593:
2580:
2579:
2280:Acid phosphatase
2161:Monoacylglycerol
2071:ester hydrolases
2040:
2033:
2026:
2017:
2016:
2011:
2010:
1972:
1966:
1965:
1947:
1941:
1940:
1923:
1917:
1916:
1906:
1897:(19): 10836–41.
1882:
1876:
1875:
1865:
1841:
1835:
1834:
1824:
1800:
1794:
1793:
1783:
1774:(22): 13506–12.
1759:
1753:
1752:
1734:
1728:
1727:
1717:
1692:(6): 1208–1215.
1676:
1670:
1669:
1629:
1623:
1622:
1612:
1602:
1578:
1572:
1571:
1531:
1525:
1524:
1484:
1478:
1477:
1467:
1449:
1440:(9): 5122–5127.
1425:
1419:
1418:
1370:
1364:
1363:
1353:
1321:
1315:
1314:
1304:
1281:The EMBO Journal
1272:
1263:
1262:
1252:
1228:
1222:
1221:
1203:
1197:
1196:
1178:
1172:
1171:
1153:
1147:
1146:
1136:
1112:
1106:
1105:
1073:
1062:
1061:
1046:Simon M (2010).
1043:
1037:
1036:
1020:
1010:
993:
992:
980:
970:
964:
963:
953:
921:
915:
914:
912:
910:
895:
773:
731:
714:
672:
655:
635:
605:
588:
555:
554:
485:Non-standard DNA
433:RNase III family
410:endoribonuclease
309:
188:Escherichia coli
90:restriction site
3192:
3191:
3187:
3186:
3185:
3183:
3182:
3181:
3167:
3166:
3165:
3153:
3145:
3143:
3138:
3050:Oxidoreductases
3036:
3012:Enzyme kinetics
3000:
2996:List of enzymes
2969:
2938:
2909:Catalytic triad
2887:
2882:
2852:
2839:
2806:
2694:
2684:
2678:
2641:
2628:
2616:Exoribonuclease
2610:
2587:
2571:
2567:
2558:
2532:Arylsulfatase L
2527:Arylsulfatase B
2522:Arylsulfatase A
2497:
2410:
2401:
2240:
2208:
2070:
2058:
2044:
2014:
1973:
1969:
1962:
1948:
1944:
1937:
1925:
1924:
1920:
1883:
1879:
1856:(23): 8107–13.
1842:
1838:
1801:
1797:
1760:
1756:
1749:
1739:DNA replication
1735:
1731:
1677:
1673:
1630:
1626:
1579:
1575:
1532:
1528:
1485:
1481:
1426:
1422:
1371:
1367:
1322:
1318:
1273:
1266:
1243:(58): 9022–32.
1229:
1225:
1218:
1204:
1200:
1193:
1179:
1175:
1168:
1154:
1150:
1127:(20): 7265–72.
1113:
1109:
1074:
1065:
1058:
1044:
1040:
1033:
1011:
996:
989:
977:Lewin's genes X
971:
967:
922:
918:
908:
906:
897:
896:
892:
888:
875:AP endonuclease
851:
832:
786:AP endonuclease
777:Bovine pancreas
769:
760:Ustilago maydis
727:
710:
668:
651:
631:
609:phage T4 (denA)
601:
584:
550:
475:
455:
406:
393:DNA replication
386:
384:DNA Replication
373:
344:
336:DNA replication
315:
296:AP endonuclease
292:
284:
176:
135:
119:recombinant DNA
103:recombinant DNA
17:
12:
11:
5:
3190:
3180:
3179:
3164:
3163:
3140:
3139:
3137:
3136:
3123:
3110:
3097:
3084:
3071:
3058:
3044:
3042:
3038:
3037:
3035:
3034:
3029:
3024:
3019:
3014:
3008:
3006:
3002:
3001:
2999:
2998:
2993:
2988:
2983:
2977:
2975:
2974:Classification
2971:
2970:
2968:
2967:
2962:
2957:
2952:
2946:
2944:
2940:
2939:
2937:
2936:
2931:
2926:
2921:
2916:
2911:
2906:
2901:
2895:
2893:
2889:
2888:
2881:
2880:
2873:
2866:
2858:
2849:
2848:
2845:
2844:
2841:
2840:
2838:
2837:
2832:
2827:
2826:
2825:
2814:
2812:
2808:
2807:
2805:
2804:
2799:
2798:
2797:
2792:
2787:
2782:
2772:
2767:
2762:
2757:
2756:
2755:
2750:
2745:
2740:
2730:
2729:
2728:
2719:
2704:
2702:
2696:
2695:
2693:
2692:
2687:
2672:
2667:
2662:
2656:
2654:
2645:
2634:
2633:
2630:
2629:
2627:
2626:
2620:
2618:
2612:
2611:
2609:
2608:
2602:
2600:
2591:
2577:
2560:
2559:
2557:
2556:
2551:
2546:
2541:
2540:
2539:
2534:
2529:
2524:
2511:
2509:
2499:
2498:
2496:
2495:
2490:
2480:
2475:
2466:
2461:
2456:
2451:
2450:
2449:
2439:
2438:
2437:
2432:
2422:
2416:
2414:
2403:
2402:
2400:
2399:
2394:
2389:
2380:
2379:
2378:
2360:
2359:
2358:
2348:
2343:
2338:
2333:
2328:
2327:
2326:
2316:
2315:
2314:
2304:
2299:
2294:
2277:
2276:
2275:
2270:
2265:
2254:
2252:
2242:
2241:
2239:
2238:
2233:
2228:
2222:
2220:
2210:
2209:
2207:
2206:
2201:
2195:
2194:
2193:
2192:
2187:
2182:
2171:
2170:
2169:
2168:
2166:Diacylglycerol
2163:
2158:
2153:
2148:
2143:
2138:
2133:
2128:
2119:
2108:
2107:
2102:
2097:
2095:Pectinesterase
2092:
2091:
2090:
2085:
2078:Cholinesterase
2074:
2072:
2060:
2059:
2043:
2042:
2035:
2028:
2020:
2013:
2012:
1991:10.1038/ng.204
1967:
1960:
1942:
1935:
1918:
1877:
1836:
1815:(11): 2694–9.
1795:
1754:
1747:
1729:
1671:
1624:
1587:Microorganisms
1573:
1526:
1499:(1): 589–615.
1479:
1420:
1365:
1316:
1264:
1223:
1216:
1198:
1191:
1173:
1166:
1148:
1107:
1063:
1057:978-1441919632
1056:
1038:
1031:
994:
987:
965:
936:(6): 897–903.
916:
889:
887:
884:
883:
882:
877:
872:
867:
862:
857:
850:
847:
831:
828:
825:
824:
817:
814:
808:
807:
804:
799:
795:
794:
791:
788:
782:
781:
778:
775:
765:
764:
761:
758:
754:
753:
750:
747:
740:
739:
736:
733:
723:
722:
719:
716:
703:
702:
699:
696:
692:
691:
688:
685:
681:
680:
677:
676:E. coli (xthA)
674:
664:
663:
660:
659:Staphylococcus
657:
644:
643:
640:
639:E. coli (endA)
637:
627:
626:
623:
618:
614:
613:
610:
607:
597:
596:
593:
590:
580:
579:
576:
573:
566:
565:
562:
559:
549:
546:
522:
521:
506:
503:
493:
487:
486:
483:
482:Standard dsDNA
474:
471:
454:
451:
405:
404:RNA Processing
402:
385:
382:
372:
369:
364:endonuclease V
356:thymine dimers
343:
340:
314:
311:
291:
288:
283:
280:
218:strain B, and
175:
172:
134:
131:
43:chain (namely
41:polynucleotide
15:
9:
6:
4:
3:
2:
3189:
3178:
3175:
3174:
3172:
3162:
3157:
3152:
3151:
3148:
3134:
3130:
3129:
3124:
3121:
3117:
3116:
3111:
3108:
3104:
3103:
3098:
3095:
3091:
3090:
3085:
3082:
3078:
3077:
3072:
3069:
3065:
3064:
3059:
3056:
3052:
3051:
3046:
3045:
3043:
3039:
3033:
3030:
3028:
3025:
3023:
3020:
3018:
3015:
3013:
3010:
3009:
3007:
3003:
2997:
2994:
2992:
2991:Enzyme family
2989:
2987:
2984:
2982:
2979:
2978:
2976:
2972:
2966:
2963:
2961:
2958:
2956:
2955:Cooperativity
2953:
2951:
2948:
2947:
2945:
2941:
2935:
2932:
2930:
2927:
2925:
2922:
2920:
2917:
2915:
2914:Oxyanion hole
2912:
2910:
2907:
2905:
2902:
2900:
2897:
2896:
2894:
2890:
2886:
2879:
2874:
2872:
2867:
2865:
2860:
2859:
2856:
2836:
2833:
2831:
2828:
2824:
2821:
2820:
2819:
2816:
2815:
2813:
2809:
2803:
2800:
2796:
2793:
2791:
2788:
2786:
2783:
2781:
2778:
2777:
2776:
2773:
2771:
2768:
2766:
2763:
2761:
2758:
2754:
2751:
2749:
2746:
2744:
2741:
2739:
2736:
2735:
2734:
2731:
2727:
2723:
2720:
2718:
2714:
2711:
2710:
2709:
2706:
2705:
2703:
2701:
2697:
2691:
2688:
2683:
2676:
2673:
2671:
2668:
2666:
2663:
2661:
2658:
2657:
2655:
2653:
2649:
2646:
2644:
2639:
2635:
2625:
2622:
2621:
2619:
2617:
2613:
2607:
2604:
2603:
2601:
2599:
2595:
2592:
2590:
2585:
2581:
2578:
2575:
2570:
2565:
2561:
2555:
2552:
2550:
2547:
2545:
2542:
2538:
2535:
2533:
2530:
2528:
2525:
2523:
2520:
2519:
2518:
2517:
2516:arylsulfatase
2513:
2512:
2510:
2508:
2504:
2500:
2494:
2491:
2488:
2484:
2481:
2479:
2476:
2474:
2470:
2467:
2465:
2462:
2460:
2457:
2455:
2452:
2448:
2445:
2444:
2443:
2440:
2436:
2433:
2431:
2428:
2427:
2426:
2425:Phospholipase
2423:
2421:
2418:
2417:
2415:
2413:
2408:
2404:
2398:
2395:
2393:
2390:
2388:
2384:
2381:
2377:
2373:
2369:
2366:
2365:
2364:
2361:
2357:
2354:
2353:
2352:
2349:
2347:
2344:
2342:
2339:
2337:
2334:
2332:
2329:
2325:
2322:
2321:
2320:
2317:
2313:
2310:
2309:
2308:
2305:
2303:
2300:
2298:
2295:
2293:
2289:
2285:
2281:
2278:
2274:
2271:
2269:
2266:
2264:
2261:
2260:
2259:
2256:
2255:
2253:
2251:
2247:
2243:
2237:
2234:
2232:
2229:
2227:
2224:
2223:
2221:
2219:
2215:
2211:
2205:
2202:
2200:
2197:
2196:
2191:
2188:
2186:
2183:
2181:
2178:
2177:
2176:
2175:Phospholipase
2173:
2172:
2167:
2164:
2162:
2159:
2157:
2154:
2152:
2149:
2147:
2144:
2142:
2139:
2137:
2134:
2132:
2129:
2127:
2123:
2120:
2118:
2115:
2114:
2113:
2110:
2109:
2106:
2103:
2101:
2098:
2096:
2093:
2089:
2086:
2084:
2081:
2080:
2079:
2076:
2075:
2073:
2069:
2065:
2061:
2056:
2052:
2048:
2041:
2036:
2034:
2029:
2027:
2022:
2021:
2018:
2008:
2004:
2000:
1996:
1992:
1988:
1985:(9): 1113–8.
1984:
1981:
1980:
1971:
1963:
1957:
1953:
1946:
1938:
1932:
1928:
1922:
1914:
1910:
1905:
1900:
1896:
1892:
1888:
1881:
1873:
1869:
1864:
1859:
1855:
1851:
1847:
1840:
1832:
1828:
1823:
1818:
1814:
1810:
1806:
1799:
1791:
1787:
1782:
1777:
1773:
1769:
1765:
1758:
1750:
1744:
1740:
1733:
1725:
1721:
1716:
1711:
1707:
1703:
1699:
1695:
1691:
1687:
1683:
1675:
1667:
1663:
1659:
1655:
1651:
1647:
1643:
1639:
1635:
1628:
1620:
1616:
1611:
1606:
1601:
1596:
1592:
1588:
1584:
1577:
1569:
1565:
1561:
1557:
1553:
1549:
1545:
1541:
1537:
1530:
1522:
1518:
1514:
1510:
1506:
1502:
1498:
1494:
1490:
1483:
1475:
1471:
1466:
1461:
1457:
1453:
1448:
1443:
1439:
1435:
1431:
1424:
1416:
1412:
1408:
1404:
1400:
1396:
1392:
1388:
1384:
1380:
1376:
1369:
1361:
1357:
1352:
1347:
1343:
1339:
1335:
1331:
1327:
1320:
1312:
1308:
1303:
1298:
1294:
1290:
1286:
1282:
1278:
1271:
1269:
1260:
1256:
1251:
1246:
1242:
1238:
1234:
1227:
1219:
1213:
1209:
1202:
1194:
1188:
1184:
1177:
1169:
1163:
1159:
1152:
1144:
1140:
1135:
1130:
1126:
1122:
1118:
1111:
1103:
1099:
1095:
1091:
1088:(3): 419–23.
1087:
1083:
1079:
1072:
1070:
1068:
1059:
1053:
1049:
1042:
1034:
1028:
1024:
1019:
1018:
1009:
1007:
1005:
1003:
1001:
999:
990:
984:
979:
978:
969:
961:
957:
952:
947:
943:
939:
935:
931:
927:
920:
904:
900:
894:
890:
881:
878:
876:
873:
871:
868:
866:
863:
861:
858:
856:
853:
852:
846:
842:
838:
836:
822:
818:
815:
813:
810:
809:
805:
803:
800:
797:
796:
792:
789:
787:
784:
783:
779:
776:
772:
767:
766:
762:
759:
756:
755:
751:
748:
745:
742:
741:
737:
734:
730:
726:P1-nuclease (
725:
724:
720:
717:
713:
708:
705:
704:
700:
697:
694:
693:
689:
686:
683:
682:
678:
675:
671:
666:
665:
661:
658:
654:
649:
646:
645:
641:
638:
634:
629:
628:
624:
622:
619:
616:
615:
611:
608:
604:
599:
598:
594:
591:
587:
582:
581:
577:
574:
571:
568:
567:
563:
560:
557:
556:
553:
545:
543:
542:
537:
532:
530:
525:
519:
515:
511:
507:
504:
501:
497:
494:
492:
489:
488:
484:
481:
480:
479:
470:
468:
464:
460:
450:
448:
444:
440:
438:
434:
430:
426:
421:
419:
415:
411:
401:
398:
394:
390:
381:
378:
368:
365:
362:gene encodes
361:
357:
353:
349:
339:
337:
332:
328:
323:
319:
318:Repair of DNA
310:
308:
303:
301:
297:
287:
279:
277:
267:
263:
261:
257:
253:
249:
245:
241:
237:
233:
232:H. influenzae
229:
225:
224:H. influenzae
221:
217:
213:
209:
205:
201:
197:
193:
189:
185:
181:
171:
169:
165:
161:
157:
153:
148:
144:
143:endonucleases
140:
130:
128:
124:
120:
116:
112:
108:
104:
100:
95:
91:
87:
83:
78:
76:
72:
68:
64:
60:
59:
54:
50:
46:
42:
38:
34:
30:
26:
25:endonucleases
22:
3128:Translocases
3125:
3112:
3099:
3086:
3073:
3063:Transferases
3060:
3047:
2904:Binding site
2685:}}
2679:{{
2643:Endonuclease
2642:
2574:ribonuclease
2514:
2297:Nucleotidase
2218:Thioesterase
1982:
1977:
1970:
1952:Biochemistry
1951:
1945:
1926:
1921:
1894:
1890:
1880:
1853:
1849:
1839:
1812:
1808:
1798:
1771:
1767:
1757:
1738:
1732:
1689:
1685:
1674:
1644:(1): 59–68.
1641:
1637:
1627:
1593:(12): 2608.
1590:
1586:
1576:
1543:
1539:
1529:
1496:
1492:
1482:
1437:
1433:
1423:
1385:(1): 31–37.
1382:
1378:
1368:
1336:(1): 72–98.
1333:
1329:
1319:
1284:
1280:
1240:
1236:
1226:
1207:
1201:
1182:
1176:
1157:
1151:
1124:
1120:
1110:
1085:
1081:
1047:
1041:
1016:
976:
968:
933:
929:
919:
907:. Retrieved
902:
893:
870:Ribonuclease
843:
839:
833:
621:P. espejiana
551:
539:
533:
526:
523:
510:Eric T. Kool
500:G-quadruplex
476:
456:
441:
422:
407:
387:
374:
359:
346:Exposure of
345:
316:
304:
299:
293:
285:
278:is Type II.
272:
259:
255:
251:
247:
243:
239:
235:
231:
227:
223:
219:
215:
211:
207:
203:
199:
195:
191:
187:
183:
182:yZ", where "
179:
177:
167:
163:
159:
155:
151:
136:
106:
89:
79:
74:
70:
67:exonucleases
62:
56:
24:
18:
2899:Active site
2818:Nuclease S1
2589:Exonuclease
2483:Lecithinase
2312:Calcineurin
2250:Phosphatase
2156:Lipoprotein
2146:Endothelial
1979:Nat. Genet.
1546:: 319–368.
855:Exonuclease
707:S1 nuclease
572:enonuclease
467:keratinized
352:ultraviolet
94:palindromic
3102:Isomerases
3076:Hydrolases
2943:Regulation
2131:Pancreatic
2068:Carboxylic
886:References
802:HeLa cells
518:proteomics
463:maturation
290:DNA repair
166:dIII, and
133:Categories
99:DNA ligase
82:eubacteria
2981:EC number
2708:RNase III
2566:(includes
2507:Sulfatase
2420:Autotaxin
2284:Prostatic
2136:Lysosomal
2051:esterases
2047:Hydrolase
2007:205345070
1706:0022-202X
1658:0009-7330
1560:1877-1173
1513:0066-4154
1456:0027-8424
1399:1865-3774
830:Mutations
768:Dnase I (
690:Comments
564:Comments
371:Apoptosis
250:II" and "
174:Notations
147:methylase
39:within a
3171:Category
3005:Kinetics
2929:Cofactor
2892:Activity
2802:RNase T1
2564:Nuclease
2199:Cutinase
1999:18711368
1724:21307874
1666:17540974
1619:34946208
1568:19215776
1521:15189154
1474:11309502
1415:25475000
1407:16867899
1311:17036055
1259:12483517
1237:Oncogene
865:Nuclease
849:See also
816:Nucleus
459:DNase1L2
429:tRNase Z
117:to make
3161:Biology
3115:Ligases
2885:Enzymes
2775:RNase E
2770:RNase Z
2765:RNase A
2760:RNase P
2733:RNase H
2351:Phytase
2151:Hepatic
2126:Lingual
2122:Gastric
1913:2358441
1872:4201782
1831:4287861
1790:6643438
1715:3185332
1610:8708148
1360:6261109
1302:1618088
1102:4588280
909:May 21,
880:HUH-tag
821:Okazaki
575:E. coli
435:, like
425:RNase P
397:Okazaki
300:E. coli
226:strain
216:E. coli
208:E. coli
86:archaea
29:enzymes
3177:EC 3.1
3147:Portal
3089:Lyases
2713:Drosha
2638:3.1.21
2606:RecBCD
2584:3.1.11
2204:PETase
2112:Lipase
2005:
1997:
1958:
1933:
1911:
1870:
1829:
1788:
1745:
1722:
1712:
1704:
1664:
1656:
1617:
1607:
1566:
1558:
1519:
1511:
1472:
1462:
1454:
1413:
1405:
1397:
1358:
1351:281499
1348:
1309:
1299:
1257:
1214:
1189:
1164:
1143:332688
1141:
1100:
1054:
1029:
985:
960:167356
958:
951:343476
948:
905:. 2017
798:Endo R
771:P00639
729:P24289
712:P24021
687:Source
670:P09030
653:P00644
633:P25736
603:P07059
586:P00641
570:RecBCD
561:Source
514:codons
447:DROSHA
437:DROSHA
334:block
222:d for
214:B for
206:R for
194:, and
33:cleave
3041:Types
2722:Dicer
2677:;see
2503:3.1.6
2473:PDE4B
2469:PDE4A
2407:3.1.4
2376:IMPA3
2372:IMPA2
2368:IMPA1
2246:3.1.3
2214:3.1.2
2064:3.1.1
2003:S2CID
1465:33174
1411:S2CID
812:FLAP1
443:DICER
377:DFF40
327:MUS81
242:dII,
31:that
3133:list
3126:EC7
3120:list
3113:EC6
3107:list
3100:EC5
3094:list
3087:EC4
3081:list
3074:EC3
3068:list
3061:EC2
3055:list
3048:EC1
2640:-31:
2586:-16:
2572:and
2478:PDE5
2464:PDE3
2459:PDE2
2454:PDE1
2346:PTEN
2331:OCRL
2324:PP2A
2273:ALPP
2268:ALPL
2263:ALPI
2057:3.1)
1995:PMID
1956:ISBN
1931:ISBN
1909:PMID
1868:PMID
1827:PMID
1786:PMID
1743:ISBN
1720:PMID
1702:ISSN
1662:PMID
1654:ISSN
1615:PMID
1564:PMID
1556:ISSN
1517:PMID
1509:ISSN
1470:PMID
1452:ISSN
1403:PMID
1395:ISSN
1356:PMID
1307:PMID
1255:PMID
1212:ISBN
1187:ISBN
1162:ISBN
1139:PMID
1098:PMID
1052:ISBN
1027:ISBN
983:ISBN
956:PMID
911:2017
445:and
427:and
416:and
360:denV
331:EME1
238:dI,
162:RV,
158:RI,
154:HI,
145:and
127:Cas9
84:and
71:endo
35:the
27:are
2795:4/5
1987:doi
1899:doi
1895:265
1858:doi
1854:248
1817:doi
1813:241
1776:doi
1772:258
1710:PMC
1694:doi
1690:131
1646:doi
1642:101
1605:PMC
1595:doi
1548:doi
1501:doi
1460:PMC
1442:doi
1387:doi
1346:PMC
1338:doi
1297:PMC
1289:doi
1245:doi
1129:doi
1125:252
1090:doi
1023:952
946:PMC
938:doi
350:to
260:Eco
252:Hae
248:Hae
244:Hin
240:Hin
236:Hin
220:Hin
212:Eco
204:Eco
200:Hin
192:Eco
184:Vwx
180:Vwx
168:Hae
164:Hin
160:Eco
156:Eco
152:Bam
61:or
49:RNA
47:or
45:DNA
19:In
3173::
2753:2C
2748:2B
2743:2A
2724::
2715::
2505::
2385::
2374:,
2370:,
2286:)/
2248::
2216::
2185:A2
2180:A1
2066::
2055:EC
2049::
2001:.
1993:.
1983:40
1907:.
1893:.
1889:.
1866:.
1852:.
1848:.
1825:.
1811:.
1807:.
1784:.
1770:.
1766:.
1718:.
1708:.
1700:.
1688:.
1684:.
1660:.
1652:.
1640:.
1636:.
1613:.
1603:.
1589:.
1585:.
1562:.
1554:.
1544:85
1542:.
1538:.
1515:.
1507:.
1497:73
1495:.
1491:.
1468:.
1458:.
1450:.
1438:98
1436:.
1432:.
1409:.
1401:.
1393:.
1383:84
1381:.
1377:.
1354:.
1344:.
1334:45
1332:.
1328:.
1305:.
1295:.
1285:25
1283:.
1279:.
1267:^
1253:.
1241:21
1239:.
1235:.
1137:.
1123:.
1119:.
1096:.
1086:81
1084:.
1080:.
1066:^
1025:.
997:^
954:.
944:.
932:.
928:.
901:.
338:.
198:,
190:,
129:.
23:,
3149::
3135:)
3131:(
3122:)
3118:(
3109:)
3105:(
3096:)
3092:(
3083:)
3079:(
3070:)
3066:(
3057:)
3053:(
2877:e
2870:t
2863:v
2790:3
2785:2
2780:1
2738:1
2576:)
2489:)
2485:(
2471:/
2447:1
2435:D
2430:C
2409::
2290:/
2282:(
2190:B
2124:/
2053:(
2039:e
2032:t
2025:v
2009:.
1989::
1964:.
1939:.
1915:.
1901::
1874:.
1860::
1833:.
1819::
1792:.
1778::
1751:.
1726:.
1696::
1668:.
1648::
1621:.
1597::
1591:9
1570:.
1550::
1523:.
1503::
1476:.
1444::
1417:.
1389::
1362:.
1340::
1313:.
1291::
1261:.
1247::
1220:.
1195:.
1170:.
1145:.
1131::
1104:.
1092::
1060:.
1035:.
991:.
962:.
940::
934:2
913:.
774:)
746:I
732:)
715:)
709:(
673:)
656:)
650:(
636:)
606:)
589:)
502:)
329:/
228:d
109:(
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.