189:
76:
17:
201:
migration is optimal and the junction will be free to move up and down the strands. When the ions are present, they neutralize the negatively charged backbone. This allows the strands to move closer together and the junction adopts the stacked X structure. It is during this state that resolution will be optimal, allowing RuvC to bind to the junction.
101:
come together and form a complex that facilitates the process in a number of ways. RuvA is a tetramer and binds to the DNA at the
Holliday junction when it is in the open X form. The protein binds in a way that the DNA entering/departing the junction is still free to rotate and slide through. RuvA
200:
ions, specifically magnesium ions (Mg), present during recombination. The ions determine which structure the
Holliday junction will adopt, as they play a stabilizing role. When the ions are absent, the backbones repel each other and the junction takes on the open X structure. In this condition,
43:, following the exchange of two single strands of DNA between two homologous chromosomes. The process is random, and the branch point can be displaced in either direction on the strand, influencing the degree of which the genetic material is exchanged. Branch migration can also be seen in
20:
Diagram illustrating the movement of a branch point between two homologous pieces of DNA. Migration travels to the left and stops when it reaches the end of the homologous region. The second branch point on the right is free to move in either direction as
180:, for efficiently repairing DNA damage. Helicase Saci-0814 is classified as an aLhr1 (archaeal long helicase related 1) under superfamily 2 helicases, and its homologs are conserved among the archaea.
79:
Open X structure of a
Holliday junction. RuvA binds to the DNA and fits in between the double strands on all four sides. RuvA also has a domain that fits inside the centre of the junction.
168:
strain deleted for Saci-0814, the homologous recombination frequency was reduced five-fold compared to the parental strain indicating that Saci-0814 is involved in homologous recombination
121:
activity, and also binds the DNA. As ATP is hydrolyzed, RuvB rotates the recombined strands while pulling them out of the junction, but does not separate the strands as helicase would.
106:
residues that interfere with the base pairs in the centre of the junction. This forces the base pairs apart so that they can re-anneal with base pairs on the homologous strands.
314:
Yamada, Kazuhiro; Ariyoshi, Mariko; Morikawa, Kosuke (2004-04-01). "Three-dimensional structural views of branch migration and resolution in DNA homologous recombination".
132:
activity, and cleaves the strands at exactly the same time. The cleavage is symmetrical, and gives two recombined DNA molecules with single stranded breaks.
176:
and functions as a branch migration helicase. Homologous recombination appears to be an important adaptation in hyperthermophiles, such as
192:
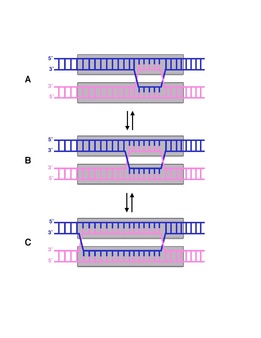
The
Holliday junction converts between the open X structure (top) and the stacked X structure (bottom) depending on the Mg concentration.
458:
Suzuki, Shoji; Kurosawa, Norio; Yamagami, Takeshi; Matsumoto, Shunsuke; Numata, Tomoyuki; Ishino, Sonoko; Ishino, Yoshizumi (2021).
140:
The eukaryotic mechanism is much more complex involving different and additional proteins, but follows the same general path.
460:"Genetic and Biochemical Characterizations of aLhr1 Helicase in the Thermophilic Crenarchaeon Sulfolobus acidocaldarius"
128:. The protein is a dimer, and will bind to the Holliday junction when it takes on the stacked X form. The protein has
400:"RAD54 N-terminal domain is a DNA sensor that couples ATP hydrolysis with branch migration of Holliday junctions"
51:, when filling in gaps in the sequence. It can also be seen when a foreign piece of DNA invades the strand.
540:
535:
283:
172:. Based on this evidence it appears that Saci-0814 is employed in homologous recombination in
110:
39:, moving the branch point up or down the DNA sequence. Branch migration is the second step of
40:
411:
8:
507:
415:
432:
399:
375:
350:
256:
29:
223:
Lilley, David M. J. (2000-05-01). "Structures of helical junctions in nucleic acids".
511:
437:
380:
331:
248:
240:
188:
145:
36:
260:
503:
471:
427:
419:
370:
362:
323:
232:
160:
dissociated DNA Holliday junction structures, and showed branch migration activity
85:
351:"Crystal structure of RuvC resolvase in complex with Holliday junction substrate"
124:
The final step in branch migration is called resolution and requires the protein
48:
423:
114:
327:
236:
156:
A helicase (designated Saci-0814) isolated from the thermophilic crenarchaeon
83:
The mechanism for prokaryotic branch migration has been studied many times in
529:
398:
Goyal N, Rossi MJ, Mazina OM, Chi Y, Moritz RL, Clurman BE, Mazin AV (2018).
244:
75:
494:
Clegg, R. M. (1993-01-01). "The
Structure of the Four-Way Junction in DNA".
441:
384:
335:
252:
129:
515:
476:
459:
366:
144:, a highly conserved eukaryotic protein, is reported to oligomerize on
117:
ATP, driving the movement of the branch point. RuvB is a hexamer with
109:
In order for migration to occur, RuvA must be associated with RuvB and
103:
60:
44:
64:
16:
197:
118:
457:
141:
125:
98:
94:
196:
The rate of branch migration is dependent on the amount of
32:
348:
496:
Annual Review of
Biophysics and Biomolecular Structure
313:
349:
Górecka, K. M.; Komorowska, W.; Nowotny, M. (2013).
397:
284:"Genetic Recombination | Learn Science at Scitable"
59:The mechanism for branch migration differs between
527:
453:
451:
391:
475:
448:
431:
374:
35:strands are consecutively exchanged at a
187:
74:
15:
528:
222:
28:is the process by which base pairs on
493:
316:Current Opinion in Structural Biology
489:
487:
309:
307:
305:
303:
278:
276:
274:
272:
270:
218:
216:
214:
508:10.1146/annurev.bb.22.060193.001503
13:
14:
552:
484:
300:
267:
211:
225:Quarterly Reviews of Biophysics
342:
70:
1:
204:
148:to promote branch migration.
135:
54:
7:
113:. RuvB has the ability to
10:
557:
424:10.1038/s41467-017-02497-x
183:
151:
328:10.1016/j.sbi.2004.03.005
237:10.1017/s0033583500003590
158:Sulfolobus acidocaldarius
102:has a domain with acidic
355:Nucleic Acids Research
193:
80:
22:
477:10.3390/catal12010034
191:
78:
41:genetic recombination
19:
416:2018NatCo...9...34G
541:Molecular genetics
536:Cellular processes
367:10.1093/nar/gkt769
194:
146:Holliday junctions
81:
23:
361:(21): 9945–9955.
178:S. acidocaldarius
174:S. acidocaldarius
166:S. acidocaldarius
37:Holliday junction
548:
520:
519:
491:
482:
481:
479:
455:
446:
445:
435:
395:
389:
388:
378:
346:
340:
339:
311:
298:
297:
295:
294:
280:
265:
264:
220:
86:Escherichia coli
26:Branch migration
556:
555:
551:
550:
549:
547:
546:
545:
526:
525:
524:
523:
492:
485:
456:
449:
396:
392:
347:
343:
312:
301:
292:
290:
282:
281:
268:
221:
212:
207:
186:
154:
138:
73:
57:
12:
11:
5:
554:
544:
543:
538:
522:
521:
502:(1): 299–328.
483:
447:
390:
341:
322:(2): 130–137.
299:
288:www.nature.com
266:
231:(2): 109–159.
209:
208:
206:
203:
185:
182:
153:
150:
137:
134:
72:
69:
56:
53:
9:
6:
4:
3:
2:
553:
542:
539:
537:
534:
533:
531:
517:
513:
509:
505:
501:
497:
490:
488:
478:
473:
469:
465:
461:
454:
452:
443:
439:
434:
429:
425:
421:
417:
413:
409:
405:
401:
394:
386:
382:
377:
372:
368:
364:
360:
356:
352:
345:
337:
333:
329:
325:
321:
317:
310:
308:
306:
304:
289:
285:
279:
277:
275:
273:
271:
262:
258:
254:
250:
246:
242:
238:
234:
230:
226:
219:
217:
215:
210:
202:
199:
190:
181:
179:
175:
171:
167:
163:
159:
149:
147:
143:
133:
131:
127:
122:
120:
116:
112:
107:
105:
100:
96:
93:the proteins
92:
88:
87:
77:
68:
66:
62:
52:
50:
46:
42:
38:
34:
31:
27:
18:
499:
495:
467:
463:
407:
403:
393:
358:
354:
344:
319:
315:
291:. Retrieved
287:
228:
224:
195:
177:
173:
169:
165:
161:
157:
155:
139:
130:endonuclease
123:
108:
90:
84:
82:
58:
25:
24:
71:Prokaryotes
61:prokaryotes
49:replication
530:Categories
404:Nat Commun
293:2015-11-13
205:References
136:Eukaryotes
104:amino acid
65:eukaryotes
45:DNA repair
30:homologous
464:Catalysts
410:(1): 34.
245:1469-8994
115:hydrolyze
55:Mechanism
442:29295984
385:23980027
336:15093826
261:40501795
253:11131562
198:divalent
164:. In a
162:in vitro
119:helicase
91:E. coli,
516:8347993
433:5750232
412:Bibcode
376:3834835
184:Control
170:in vivo
152:Archaea
514:
470:: 34.
440:
430:
383:
373:
334:
259:
251:
243:
257:S2CID
142:Rad54
89:. In
21:well.
512:PMID
438:PMID
381:PMID
332:PMID
249:PMID
241:ISSN
126:RuvC
99:RuvB
97:and
95:RuvA
63:and
47:and
504:doi
472:doi
428:PMC
420:doi
371:PMC
363:doi
324:doi
233:doi
111:ATP
33:DNA
532::
510:.
500:22
498:.
486:^
468:12
466:.
462:.
450:^
436:.
426:.
418:.
406:.
402:.
379:.
369:.
359:41
357:.
353:.
330:.
320:14
318:.
302:^
286:.
269:^
255:.
247:.
239:.
229:33
227:.
213:^
67:.
518:.
506::
480:.
474::
444:.
422::
414::
408:9
387:.
365::
338:.
326::
296:.
263:.
235::
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.