869:, scientists simulated the evolution of DNA sequences to analyze the emergence of features that underly enhancer function. This allowed the design and production of a range of functioning synthetic enhancers for different cell types of the fruit fly brain. A second approach trained artificial intelligence models on single-cell DNA accessibility data and transferred the learned models towards the prediction of enhancers for selected tissues in the fruit fly embryo. These enhancer prediction models were used to design synthetic enhancers for the nervous system, brain, muscle, epidermis and gut.
234:
316:), with one member of the dimer anchored to its binding motif on the enhancer and the other member anchored to its binding motif on the promoter (represented by the red zigzags in the illustration). Several cell function specific transcription factors (there are about 1,600 transcription factors in a human cell) generally bind to specific motifs on an enhancer and a small combination of these enhancer-bound transcription factors, when brought close to a promoter by a DNA loop, govern level of transcription of the target gene.
40:
304:
enhancer DNA regions, for a particular type of tissue only specific enhancers are brought into proximity with the promoters that they regulate. In a study of brain cortical neurons, 24,937 loops were found, bringing enhancers to their target promoters. Multiple enhancers, each often at tens or hundreds of thousands of nucleotides distant from their target genes, loop to their target gene promoters and can coordinate with each other to control the expression of their common target gene.
696:
gene it regulates). On its own, each enhancer drives nearly identical patterns of gene expression. Are the two enhancers truly redundant? Recent work has shown that multiple enhancers allow fruit flies to survive environmental perturbations, such as an increase in temperature. When raised at an elevated temperature, a single enhancer sometimes fails to drive the complete pattern of expression, whereas the presence of both enhancers permits normal gene expression.
335:. An inactive enhancer may be bound by an inactive transcription factor. Phosphorylation of the transcription factor may activate it and that activated transcription factor may then activate the enhancer to which it is bound (see small red star representing phosphorylation of transcription factor bound to enhancer in the illustration). An activated enhancer begins transcription of its RNA before activating transcription of messenger RNA from its target gene.
562:, yet both species' sequences produce nearly identical patterns of reporter gene expression in zebrafish. Similarly, in highly diverged insects (separated by around 350 million years), similar gene expression patterns of several key genes was found to be regulated through similarly constituted CRMs although these CRMs do not show any appreciable sequence conservation detectable by standard sequence alignment methods such as
33:
645:). The PEE turns on Nodal expression in response to a combination of Wnt signaling plus a second, unknown signal; thus, a member of the LEF/TCF transcription factor family likely binds to a TCF binding site in the cells in the node. Diffusion of Nodal away from the node forms a gradient which then patterns the extending anterior-posterior axis of the embryo. The ASE is an intronic enhancer bound by the
198:, which recruits polymerase II and the general transcription factors which then begin transcribing the genes. Enhancers can also be found within introns. An enhancer's orientation may even be reversed without affecting its function; additionally, an enhancer may be excised and inserted elsewhere in the chromosome, and still affect gene transcription. That is one reason that introns
116:. They can be located up to 1 Mbp (1,000,000 bp) away from the gene, upstream or downstream from the start site. There are hundreds of thousands of enhancers in the human genome. They are found in both prokaryotes and eukaryotes. Active enhancers typically get transcribed as enhancer or regulatory non-coding RNA, whose expression levels correlate with mRNA levels of target genes.
513:, clustering of known or predicted TF-binding sites, and supervised machine-learning approaches trained on known CRMs. All of these methods have proven effective for CRM discovery, but each has its own considerations and limitations, and each is subject to a greater or lesser number of false-positive identifications. In the
596:
transcription factors, thus creating zones within which different combinations of transcription factors are expressed. The pair-rule genes are separated from one another by non-expressing cells. Moreover, the stripes of expression for different pair-rule genes are offset by a few cell diameters from
695:
Some genes involved in critical developmental processes contain multiple enhancers of overlapping function. Secondary enhancers, or "shadow enhancers", may be found many kilobases away from the primary enhancer ("primary" usually refers to the first enhancer discovered, which is often closer to the
303:
Enhancers are regions of the genome that are major gene-regulatory elements. Enhancers control cell-type-specific gene expression programs, most often by looping through long distances to come in physical proximity with the promoters of their target genes. While there are hundreds of thousands of
833:
facilitates remodeling of chromatin in a manner that selectively redistributes cofactors from high-occupancy enhancers, thereby repressing genes involved in maintaining cellular identify whose expression they enhance; at the same time, this F-κB-driven remodeling and redistribution activates other
418:
An enhancer near the gene GADD45g has been described that may regulate brain growth in chimpanzees and other mammals, but not in humans. The GADD45G regulator in mice and chimps is active in regions of the brain where cells that form the cortex, ventral forebrain, and thalamus are located and may
269:
shown by a small red star on a transcription factor on the enhancer) the enhancer is activated and can now activate its target promoter. The active enhancer is transcribed on each strand of DNA in opposite directions by bound RNAP IIs. Mediator (a complex consisting of about 26 proteins in an
478:. If the reporter gene integrates near an enhancer, its expression will reflect the expression pattern driven by that enhancer. Thus, staining the flies for LacZ expression or activity and cloning the sequence surrounding the integration site allows the identification of the enhancer sequence.
682:
expression is controlled in the early embryo by an intronic enhancer that binds another forkhead domain transcription factor, FoxA2. Initially the enhancer drives broad gene expression throughout the embryo, but the expression quickly becomes restricted to the endoderm, suggesting that other
299:
have a leading role in the regulation of gene expression. An enhancer localized in a DNA region distant from the promoter of a gene can have a very large effect on gene expression, with some genes undergoing up to 100-fold increased expression due to an activated enhancer.
834:
enhancers that guide changes in cellular function through inflammation. As a result, inflammation reprograms cells, altering their interactions with the rest of tissue and with the immune system. In cancer, proteins that control NF-κB activity are dysregulated, permitting
286:
of genes. Core promoters are sufficient to direct transcription initiation, but generally have low basal activity. Other important cis-regulatory modules are localized in DNA regions that are distant from the transcription start sites. These include enhancers,
792:
gene produce gene expression in precisely this pattern – the vein spot enhancer drives reporter gene expression in the 12 spots, and the intervein shade enhancer drives reporter expression in the 4 distinct patches. These two enhancers are responsive to the
597:
one another. Thus, unique combinations of pair-rule gene expression create spatial domains along the anterior-posterior axis to set up each of the 14 individual segments. The 480 bp enhancer responsible for driving the sharp stripe two of the pair-rule gene
127:, provided an explanation for the transcriptional activation of rearranged Vh gene promoters while unrearranged Vh promoters remained inactive. Lately, enhancers have been shown to be involved in certain medical conditions, for example,
554:. Although much evidence has pointed to sequence conservation for critical developmental enhancers, other work has shown that the function of enhancers can be conserved with little or no primary sequence conservation. For example, the
735:
gene involved in posterior limb development in vertebrates. Preliminary genetic analyses indicated that changes in the expression of this gene were responsible for pelvic reduction in sticklebacks. Fish expressing only the freshwater
320:(a complex usually consisting of about 26 proteins in an interacting structure) communicates regulatory signals from enhancer DNA-bound transcription factors directly to the RNA polymerase II (pol II) enzyme bound to the promoter.
307:
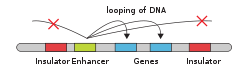
The schematic illustration in this section shows an enhancer looping around to come into close physical proximity with the promoter of a target gene. The loop is stabilized by a dimer of a connector protein (e.g. dimer of
850:
Synthetic regulatory elements such as enhancers promise to be a powerful tool to direct gene products to particular cell types in order to treat disease by activating beneficial genes or by halting aberrant cell states.
683:
repressors may be involved in its restriction. Late in development, the same enhancer restricts expression to the tissues that will become the stomach and pancreas. An additional enhancer is responsible for maintaining
175:, repress the transcription of the gene. Silencers and enhancers may be in close proximity to each other or may even be in the same region only differentiated by the transcription factor the region binds to.
260:
of the gene. The loop is stabilized by one architectural protein anchored to the enhancer and one anchored to the promoter and these proteins are joined to form a dimer (red zigzags). Specific regulatory
723:
fish. Sticklebacks exist in both marine and freshwater environments, but sticklebacks in many freshwater populations have completely lost their pelvic fins (appendages homologous to the posterior limb of
43:
Seen here is a four step diagram depicting the usage of an enhancer. Within this DNA sequence, protein(s) known as transcription factor(s) bind to the enhancer and increase the activity of the promoter.
525:
can be indicative of enhancers. Sequences from multiple species are aligned, and conserved regions are identified computationally. Identified sequences can then be attached to a reporter gene such as
744:
do not have pelvic spines, whereas fish expressing a marine allele retain pelvic spines. A more thorough characterization showed that a 500 base pair enhancer sequence is responsible for turning on
609:
is only expressed in a narrow stripe of cells that contain high concentrations of the activators and low concentration of the repressors for this enhancer sequence. Other enhancer regions drive
443:
and other DNA-binding proteins in a developing tissue controls which genes will be expressed in that tissue. Enhancers allow the same gene to be used in diverse processes in space and time.
825:". These enhancers contain a large number of binding sites for sequence-specific, inducible transcription factors, and regulate expression of genes involved in cell differentiation. During
1251:
Gillies SD, Morrison SL, Oi VT, Tonegawa S (July 1983). "A tissue-specific transcription enhancer element is located in the major intron of a rearranged immunoglobulin heavy chain gene".
605:) has been well-characterized. The enhancer contains 12 different binding sites for maternal and gap gene transcription factors. Activating and repressing sites overlap in sequence.
668:
is another critical step in animal development. Each of the three germ layers has unique patterns of gene expression that promote their differentiation and development. The
419:
suppress further neurogenesis. Loss of the GADD45G enhancer in humans may contribute to an increase of certain neuronal populations and to forebrain expansion in humans.
265:
bind to DNA sequence motifs on the enhancer. General transcription factors bind to the promoter. When a transcription factor is activated by a signal (here indicated as
1205:
Banerji J, Olson L, Schaffner W (July 1983). "A lymphocyte-specific cellular enhancer is located downstream of the joining region in immunoglobulin heavy chain genes".
708:("evo-devo") is investigating the role of enhancers and other cis-regulatory elements in producing morphological changes via developmental differences between species.
4107:
de
Almeida BP, Reiter F, Pagani M, Stark A (May 2022). "DeepSTARR predicts enhancer activity from DNA sequence and enables the de novo design of synthetic enhancers".
1408:
de
Almeida BP, Reiter F, Pagani M, Stark A (May 2022). "DeepSTARR predicts enhancer activity from DNA sequence and enables the de novo design of synthetic enhancers".
649:
transcription factor Fox1. Early in development, Fox1-driven Nodal expression establishes the visceral endoderm. Later in development, Fox1 binding to the ASE drives
194:
as first shown by in vivo competition experiments. Subsequently, molecular studies showed direct interactions with transcription factors and cofactors, including the
3437:
Norris DP, Brennan J, Bikoff EK, Robertson EJ (July 2002). "The Foxh1-dependent autoregulatory enhancer controls the level of Nodal signals in the mouse embryo".
489:(NGS) methods now enable high-throughput functional CRM discovery assays, and the vastly increasing amounts of available data, including large-scale libraries of
357:
Flexible billboards – less integrative, multiple proteins independently regulate gene expression and their sum is read in by the basal transcriptional machinery.
1712:
Scholer H, Haslinger A, Heguy A, Holtgreve H, Karin M (April 1986). "In vivo competition between a metallothionein regulatory element and the SV40 enhancer".
455:
techniques using a reporter gene or by comparative sequence analysis and computational genomics. In genetically tractable models such as the fruit fly
633:
gene contains two enhancers: the
Proximal Epiblast Enhancer (PEE) and the Asymmetric Enhancer (ASE). The PEE is upstream of the Nodal gene and drives
2915:
862:
strategies have led to a better understanding of the features of regulatory DNA sequences, the prediction, and the design of synthetic enhancers.
323:
Enhancers, when active, are generally transcribed from both strands of DNA with RNA polymerases acting in two different directions, producing two
772:
Pigmentation patterns provide one of the most striking and easily scored differences between different species of animals. Pigmentation of the
1669:
Mercola M, Goverman J, Mirell C, Calame K (January 1985). "Immunoglobulin heavy-chain enhancer requires one or more tissue-specific factors".
760:
expression in the pelvic spines in isolated freshwater population, and without this enhancer, freshwater fish fail to develop pelvic spines.
629:
superfamily ligand, is a key gene involved in patterning both the anterior-posterior axis and the left-right axis of the early embryo. The
2012:
Spilianakis CG, Lalioti MD, Town T, Lee GR, Flavell RA (June 2005). "Interchromosomal associations between alternatively expressed loci".
821:
Each cell typically contains several hundred of a special class of enhancers that stretch over many kilobases long DNA sequences, called "
497:
data across many cell types, are making accurate computational CRM discovery an attainable goal. An example of NGS-based approach called
1162:
Mercola M, Wang XF, Olsen J, Calame K (August 1983). "Transcriptional enhancer elements in the mouse immunoglobulin heavy chain locus".
3482:"Direct transcriptional regulation of Gata4 during early endoderm specification is controlled by FoxA2 binding to an intronic enhancer"
501:
have enabled identification of nucleosome-depleted, or open chromatin regions, which can contain CRM. More recently techniques such as
505:
have been developed which require less starting material. Nucelosome depleted regions can be identified in vivo through expression of
3280:
3902:"Late-phase synthesis of IκBα insulates the TLR4-activated canonical NF-κB pathway from noncanonical NF-κB signaling in macrophages"
3643:
Werner T, Koshikawa S, Williams TM, Carroll SB (April 2010). "Generation of a novel wing colour pattern by the
Wingless morphogen".
3853:"Acute TNF-induced repression of cell identity genes is mediated by NFκB-directed redistribution of cofactors from super-enhancers"
4252:
4398:
171:
in the eukaryotic genome. Silencers are antagonists of enhancers that, when bound to its proper transcription factors called
3398:"Nodal cis-regulatory elements reveal epiblast and primitive endoderm heterogeneity in the peri-implantation mouse embryo"
865:
Building on work in cell culture, synthetic enhancers were successfully applied to entire living organisms in 2023. Using
776:
wing has proven to be a particularly amenable system for studying the development of complex pigmentation phenotypes. The
4630:
270:
interacting structure) communicates regulatory signals from the enhancer DNA-bound transcription factors to the promoter.
135:
to design synthetic enhancers and applied them in animal systems, first in a cell line, and one year later also in vivo.
705:
3349:"Asymmetric and node-specific nodal expression patterns are controlled by two distinct cis-acting regulatory elements"
780:
wing has 12 dark pigmentation spots and 4 lighter gray intervein patches. Pigment spots arise from expression of the
490:
4602:
4219:
2867:"Enhancer RNAs predict enhancer-gene regulatory links and are critical for enhancer function in neuronal systems"
2720:"The degree of enhancer or promoter activity is reflected by the levels and directionality of eRNA transcription"
626:
588:
transcription factors are responsible for activating and repressing a number of segmentation genes, such as the
4655:
2769:"A comparison of experimental assays and analytical methods for genome-wide identification of active enhancers"
190:
upstream or downstream of the start site. Enhancers do not act on the promoter region itself, but are bound by
4597:
3745:"Chromatin stretch enhancer states drive cell-specific gene regulation and harbor human disease risk variants"
546:, which provides a more direct measure of enhancer activity, since it is not subjected to the complexities of
4650:
179:
4618:
4592:
3900:
Chatterjee B, Banoth B, Mukherjee T, Taye N, Vijayaragavan B, Chattopadhyay S, et al. (December 2016).
2477:
Schoenfelder S, Fraser P (August 2019). "Long-range enhancer-promoter contacts in gene expression control".
509:, allowing for greater control of cell-type specific enhancer identification. Computational methods include
752:—a sequence of DNA that is likely to be broken and thus more likely to be mutated as a result of imprecise
160:
3281:"Evidence for Deep Regulatory Similarities in Early Developmental Programs across Highly Diverged Insects"
481:
The development of genomic and epigenomic technologies, however, has dramatically changed the outlook for
4245:
3038:"Regulation of eukaryotic gene expression by the untranslated gene regions and other non-coding elements"
2954:
1103:
Burren OS, Rubio García A, Javierre BM, Rainbow DB, Cairns J, Cooper NJ, et al. (4 September 2017).
427:
The development, differentiation and growth of cells and tissues require precisely regulated patterns of
398:
on two legs". Evidence to date shows that of the 110,000 gene enhancer sequences identified in the human
218:
155:
DNA, so although the enhancer DNA may be far from the gene in a linear way, it is spatially close to the
3085:
Hartenstein V, Jan YN (June 1992). "Studying
Drosophila embryogenesis with P-lacZ enhancer trap lines".
4278:
4194:
3130:"CATaDa reveals global remodelling of chromatin accessibility during stem cell differentiation in vivo"
2818:"MAP kinase phosphorylation-dependent activation of Elk-1 leads to activation of the co-activator p300"
2427:"Three-dimensional genome restructuring across timescales of activity-induced neuronal gene expression"
120:
2865:
Carullo NV, Phillips Iii RA, Simon RC, Soto SA, Hinds JE, Salisbury AJ, et al. (September 2020).
1506:
Taskiran II, Spanier KI, Dickmänken H, Kempynck N, Pančíková A, Ekşi EC, et al. (February 2024).
592:. The gap genes are expressed in blocks along the anterior-posterior axis of the fly along with other
4475:
4411:
4381:
4359:
2718:
Mikhaylichenko O, Bondarenko V, Harnett D, Schor IE, Males M, Viales RR, et al. (January 2018).
657:, thus establishing left-right asymmetry necessary for asymmetric organ development in the mesoderm.
621:
Establishing body axes is a critical step in animal development. During mouse embryonic development,
526:
376:
343:
As of 2005, there are two different theories on the information processing that occurs on enhancers:
17:
756:. This fragile site has caused repeated, independent losses of the enhancer responsible for driving
4695:
4269:
4261:
4198:
3396:
Granier C, Gurchenkov V, Perea-Gomez A, Camus A, Ott S, Papanayotou C, et al. (January 2011).
749:
436:
283:
183:
151:
complex of DNA is folded in a way that functionally mimics the supercoiled state characteristic of
98:
3588:"Adaptive evolution of pelvic reduction in sticklebacks by recurrent deletion of a Pitx1 enhancer"
3300:"Dissecting the regulatory switches of development: lessons from enhancer evolution in Drosophila"
186:
initiation site to affect transcription, as some have been found located several hundred thousand
4576:
4265:
855:
801:
expression at all of the pigmented locations. Thus, in the evolution of the complex pigmentation
580:
457:
257:
132:
128:
3696:"Master transcription factors and mediator establish super-enhancers at key cell identity genes"
898:
Blackwood EM, Kadonaga JT (July 1998). "Going the distance: a current view of enhancer action".
4485:
4238:
3743:
Parker SC, Stitzel ML, Taylor DL, Orozco JM, Erdos MR, Akiyama JA, et al. (October 2013).
2065:"Genome-wide mapping of HATs and HDACs reveals distinct functions in active and inactive genes"
1359:
Zhigulev A, Norberg Z, Cordier J, Spalinskas R, Bassereh H, Björn N, et al. (March 2024).
584:
embryos are among the best characterized developmental enhancers. In the early fly embryo, the
575:
432:
317:
275:
199:
195:
110:
2979:
McLean CY, Reno PL, Pollen AA, Bassan AI, Capellini TD, Guenther C, et al. (March 2011).
4669:
4664:
4532:
4434:
4142:
Avsec Ž, Weilert M, Shrikumar A, Krueger S, Alexandari A, Dalal K, et al. (March 2021).
4006:
Vlahopoulos SA, Cen O, Hengen N, Agan J, Moschovi M, Critselis E, et al. (August 2015).
3586:
Chan YF, Marks ME, Jones FC, Villarreal G, Shapiro MD, Brady SD, et al. (January 2010).
2520:
Weintraub AS, Li CH, Zamudio AV, Sigova AA, Hannett NM, Day DS, et al. (December 2017).
794:
563:
547:
543:
539:
482:
440:
296:
295:
and tethering elements. Among this constellation of elements, enhancers and their associated
203:
2114:"Histone modifications at human enhancers reflect global cell-type-specific gene expression"
1865:"Exonic remnants of whole-genome duplication reveal cis-regulatory function of coding exons"
1814:
Ramasamy S, Aljahani A, Karpinska MA, Cao TB, Velychko T, Cruz JN, et al. (July 2023).
379:
2) is a gene enhancer "that may have contributed to the evolution of the uniquely opposable
4566:
4480:
4446:
4340:
4008:"Dynamic aberrant NF-κB spurs tumorigenesis: a new model encompassing the microenvironment"
3962:
3802:
Brown JD, Lin CY, Duan Q, Griffin G, Federation A, Paranal RM, et al. (October 2014).
3756:
3652:
3599:
3542:
2992:
2623:
2379:(September 2012). "Transcription factors: from enhancer binding to developmental control".
2182:
2125:
2021:
1768:
1721:
1678:
1625:
1519:
1459:
1311:
1171:
907:
514:
510:
475:
439:
in specific cells and/or repressing it in other cells. Thus, the particular combination of
292:
262:
191:
106:
94:
4057:"Brd4 maintains constitutively active NF-κB in cancer cells by binding to acetylated RelA"
3284:
2112:
Heintzman ND, Hon GC, Hawkins RD, Kheradpour P, Stark A, Harp LF, et al. (May 2009).
1612:
Smemo S, Tena JJ, Kim KH, Gamazon ER, Sakabe NJ, Gómez-Marín C, et al. (March 2014).
1581:
1564:
1023:
1006:
719:
Recent work has investigated the role of enhancers in morphological changes in threespine
8:
4517:
4403:
4377:
2425:
Beagan JA, Pastuzyn ED, Fernandez LR, Guo MH, Feng K, Titus KR, et al. (June 2020).
1614:"Obesity-associated variants within FTO form long-range functional connections with IRX3"
839:
288:
279:
241:
233:
4230:
3966:
3760:
3656:
3603:
3546:
2996:
2627:
2569:
Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, et al. (February 2018).
2186:
2129:
2025:
1840:
1815:
1772:
1725:
1682:
1629:
1540:
1523:
1507:
1480:
1463:
1447:
1385:
1360:
1175:
911:
182:
of the gene it regulates. Furthermore, an enhancer does not need to be located near the
4318:
4168:
4143:
4081:
4056:
4032:
4007:
3983:
3950:
3949:
Vahedi G, Kanno Y, Furumoto Y, Jiang K, Parker SC, Erdos MR, et al. (April 2015).
3926:
3901:
3877:
3852:
3828:
3803:
3779:
3744:
3720:
3695:
3694:
Whyte WA, Orlando DA, Hnisz D, Abraham BJ, Lin CY, Kagey MH, et al. (April 2013).
3676:
3620:
3587:
3563:
3530:
3506:
3481:
3462:
3324:
3299:
3257:
3232:
3205:
3180:
3156:
3129:
3110:
3062:
3037:
3013:
2980:
2946:
2891:
2866:
2793:
2768:
2744:
2719:
2695:
2670:
2646:
2611:
2546:
2521:
2502:
2451:
2426:
2404:
2352:
2325:
2301:
2276:
2252:
2227:
2226:
Blow MJ, McCulley DJ, Li Z, Zhang T, Akiyama JA, Holt A, et al. (September 2010).
2203:
2170:
2146:
2113:
2089:
2064:
2045:
1989:
1962:
1938:
1913:
1912:
Birnbaum RY, Clowney EJ, Agamy O, Kim MJ, Zhao J, Yamanaka T, et al. (June 2012).
1889:
1864:
1646:
1613:
1594:
1336:
1295:
1276:
1230:
1139:
1104:
1080:
1055:
1036:
982:
957:
931:
518:
156:
3373:
3348:
2842:
2817:
1791:
1756:
1448:"Targeted design of synthetic enhancers for selected tissues in the Drosophila embryo"
1105:"Chromosome contacts in activated T cells identify autoimmune disease candidate genes"
4637:
4173:
4124:
4086:
4037:
3988:
3931:
3882:
3833:
3784:
3725:
3668:
3625:
3568:
3511:
3454:
3419:
3378:
3329:
3262:
3210:
3161:
3102:
3067:
3018:
2938:
2896:
2847:
2798:
2749:
2700:
2651:
2592:
2551:
2506:
2494:
2456:
2408:
2396:
2357:
2306:
2257:
2208:
2151:
2094:
2037:
1994:
1943:
1894:
1845:
1796:
1737:
1694:
1651:
1586:
1545:
1485:
1425:
1390:
1341:
1268:
1264:
1222:
1218:
1187:
1144:
1126:
1085:
1028:
987:
923:
859:
253:
164:
3466:
3114:
2950:
1598:
1280:
1234:
1040:
935:
838:
to decrease their dependence on interactions with local tissue, and hindering their
4559:
4542:
4224:
4163:
4155:
4116:
4076:
4068:
4027:
4019:
3978:
3970:
3921:
3913:
3872:
3864:
3823:
3815:
3774:
3764:
3715:
3707:
3680:
3660:
3615:
3607:
3558:
3550:
3501:
3493:
3446:
3409:
3368:
3360:
3319:
3311:
3252:
3244:
3200:
3192:
3151:
3141:
3094:
3057:
3049:
3008:
3000:
2930:
2886:
2878:
2837:
2829:
2788:
2780:
2739:
2731:
2690:
2682:
2641:
2631:
2612:"Positional specificity of different transcription factor classes within enhancers"
2582:
2541:
2533:
2486:
2446:
2438:
2388:
2347:
2337:
2296:
2288:
2247:
2239:
2198:
2190:
2141:
2133:
2084:
2076:
2063:
Wang Z, Zang C, Cui K, Schones DE, Barski A, Peng W, et al. (September 2009).
2049:
2029:
1984:
1974:
1933:
1925:
1884:
1876:
1835:
1827:
1786:
1776:
1729:
1686:
1641:
1633:
1576:
1535:
1527:
1475:
1467:
1446:
de
Almeida BP, Schaub C, Pagani M, Secchia S, Furlong EE, Stark A (February 2024).
1417:
1380:
1372:
1361:"Enhancer mutations modulate the severity of chemotherapy-induced myelosuppression"
1331:
1321:
1260:
1214:
1179:
1134:
1116:
1075:
1067:
1054:
Kulaeva OI, Nizovtseva EV, Polikanov YS, Ulianov SV, Studitsky VM (December 2012).
1018:
977:
969:
915:
646:
638:
27:
DNA sequence that binds activators to increase the likelihood of gene transcription
4023:
3804:"NF-κB directs dynamic super enhancer formation in inflammation and atherogenesis"
2981:"Human-specific loss of regulatory DNA and the evolution of human-specific traits"
2816:
Li QJ, Yang SH, Maeda Y, Sladek FM, Sharrocks AD, Martins-Green M (January 2003).
2169:
Visel A, Blow MJ, Li Z, Zhang T, Akiyama JA, Holt A, et al. (February 2009).
1326:
533:
pattern of gene expression produced by the enhancer when injected into an embryo.
4423:
4144:"Base-resolution models of transcription-factor binding reveal soft motif syntax"
3851:
Schmidt SF, Larsen BD, Loft A, Nielsen R, Madsen JG, Mandrup S (September 2015).
3819:
3248:
1979:
1863:
Dong X, Navratilova P, Fredman D, Drivenes Ø, Becker TS, Lenhard B (March 2010).
878:
593:
589:
551:
428:
351:
266:
222:
3497:
3414:
3397:
2277:"Eukaryotic core promoters and the functional basis of transcription initiation"
919:
4633:
4621:
4571:
4204:
4159:
4120:
3749:
Proceedings of the
National Academy of Sciences of the United States of America
3711:
2784:
2616:
Proceedings of the
National Academy of Sciences of the United States of America
2587:
2570:
2537:
2376:
2080:
1831:
1761:
Proceedings of the
National Academy of Sciences of the United States of America
1531:
1471:
1421:
835:
822:
642:
486:
350:– rely on highly cooperative, coordinated action and can be disabled by single
332:
240:. An active enhancer regulatory region of DNA is enabled to interact with the
168:
39:
3917:
3554:
3053:
2916:"Transcriptional enhancers: Intelligent enhanceosomes or flexible billboards?"
2610:
Grossman SR, Engreitz J, Ray JP, Nguyen TH, Hacohen N, Lander ES (July 2018).
2490:
2442:
2342:
2292:
1316:
1121:
687:
expression in the endoderm during the intermediate stages of gut development.
4689:
4335:
1130:
866:
534:
522:
452:
328:
249:
3769:
3611:
3450:
2636:
1781:
1733:
1690:
1183:
4177:
4128:
4090:
4041:
3992:
3935:
3886:
3837:
3788:
3729:
3672:
3629:
3572:
3515:
3458:
3423:
3382:
3364:
3333:
3266:
3214:
3165:
3128:
Aughey GN, Estacio Gomez A, Thomson J, Yin H, Southall TD (February 2018).
3106:
3071:
3022:
2942:
2900:
2851:
2833:
2802:
2753:
2735:
2704:
2655:
2596:
2555:
2498:
2460:
2400:
2361:
2310:
2261:
2212:
2155:
2098:
2041:
1998:
1947:
1898:
1849:
1800:
1655:
1590:
1549:
1489:
1429:
1394:
1376:
1345:
1148:
1089:
1032:
991:
826:
665:
347:
324:
3951:"Super-enhancers delineate disease-associated regulatory nodes in T cells"
3868:
2882:
1929:
1880:
1741:
1698:
1272:
1226:
1191:
927:
435:
to mediate both spatial and temporal control of development by turning on
4522:
4490:
4393:
4072:
1071:
720:
661:
407:
152:
4055:
Zou Z, Huang B, Wu X, Zhang H, Qi J, Bradner J, et al. (May 2014).
3974:
3664:
3146:
3004:
2326:"The Why of YY1: Mechanisms of Transcriptional Regulation by Yin Yang 1"
2194:
2137:
2033:
1637:
956:
Pennacchio LA, Bickmore W, Dean A, Nobrega MA, Bejerano G (April 2013).
4674:
4368:
4323:
4313:
4308:
4303:
4298:
3315:
3098:
753:
558:
enhancers in humans have very little sequence conservation to those in
494:
211:
187:
144:
4102:
4100:
3181:"Identifying transcriptional cis-regulatory modules in animal genomes"
2934:
690:
4330:
3196:
1246:
1244:
809:
pigment gene evolved enhancers responsive to the wingless signal and
802:
748:
expression in the posterior fin bud. This enhancer is located near a
559:
498:
472:
403:
172:
148:
82:
2717:
2686:
2392:
2171:"ChIP-seq accurately predicts tissue-specific activity of enhancers"
1914:"Coding exons function as tissue-specific enhancers of nearby genes"
1757:"Association of the Mediator complex with enhancers of active genes"
1056:"Distant activation of transcription: mechanisms of enhancer action"
973:
813:
expression evolved at new locations to produce novel wing patterns.
4537:
4527:
4097:
2323:
2243:
1294:
Hauptman G, Reichert MC, Abdal Rhida MA, Evans TA (December 2022).
732:
669:
654:
585:
502:
74:
3480:
Rojas A, Schachterle W, Xu SM, Martín F, Black BL (October 2010).
1241:
830:
4497:
4364:
3395:
1505:
1102:
1053:
785:
678:
expression, and Gata4 goes on to direct gut morphogenesis later.
210:
region of an unrelated gene and they may act on genes on another
90:
1358:
1293:
32:
4293:
3531:"Shadow enhancers foster robustness of Drosophila gastrulation"
1816:"The Mediator complex regulates enhancer-promoter interactions"
737:
641:
that will differentiate into the node (also referred to as the
399:
372:
124:
4214:
3642:
3529:
Perry MW, Boettiger AN, Bothma JP, Levine M (September 2010).
1862:
1711:
4554:
4406:
4209:
3899:
3127:
2671:"The Mediator complex: a central integrator of transcription"
2228:"ChIP-Seq identification of weakly conserved heart enhancers"
2111:
1445:
955:
727:
674:
622:
506:
387:
383:
380:
354:
that move or remove the binding sites of individual proteins.
4141:
3436:
2864:
2767:
Yao L, Liang J, Ozer A, Leung AK, Lis JT, Yu H (July 2022).
2324:
Verheul TC, van Hijfte L, Perenthaler E, Barakat TS (2020).
1813:
4625:
4455:
4386:
4106:
1668:
1407:
845:
468:
464:
395:
391:
309:
245:
207:
102:
3528:
2522:"YY1 Is a Structural Regulator of Enhancer-Promoter Loops"
2424:
493:, collections of annotated, validated CRMs, and extensive
248:
by the formation of a chromosome loop. This can initiate
4260:
3742:
2609:
2568:
2011:
1565:"Transcriptional regulatory elements in the human genome"
1250:
1007:"Transcriptional regulatory elements in the human genome"
313:
86:
4005:
3948:
3850:
3693:
3585:
3479:
2519:
699:
119:
The first discovery of a eukaryotic enhancer was in the
1911:
763:
3233:"Enhancer identification through comparative genomics"
3230:
3185:
2978:
1754:
1161:
446:
105:
will occur. These proteins are usually referred to as
3298:
Borok MJ, Tran DA, Ho MC, Drewell RA (January 2010).
3035:
1204:
406:
of humans following the split with the ancestors of
3801:
3231:Visel A, Bristow J, Pennacchio LA (February 2007).
3036:Barrett LW, Fletcher S, Wilton SD (November 2012).
2225:
1960:
1905:
1755:Kuras L, Borggrefe T, Kornberg RD (November 2003).
1611:
691:
Multiple enhancers promote developmental robustness
471:can be randomly integrated into the genome using a
3297:
2476:
2062:
1508:"Cell-type-directed design of synthetic enhancers"
788:. Recent work has shown that two enhancers in the
402:, HACNS1 has undergone the most change during the
123:gene in 1983. This enhancer, located in the large
2815:
2766:
897:
653:expression on the left side of the lateral plate
4687:
3178:
2168:
1563:Maston GA, Evans SK, Green MR (1 January 2006).
1562:
1004:
537:expression of the reporter can be visualized by
274:Gene expression in mammals is regulated by many
4054:
3346:
2913:
2472:
2470:
1501:
1499:
1441:
1439:
491:transcription factor-binding site (TFBS) motifs
361:
3283:. Genome Biology and Evolution. Archived from
3084:
462:for example, a reporter construct such as the
159:and gene. This allows it to interact with the
4246:
1963:"De novo genesis of enhancers in vertebrates"
816:
613:expression in 6 other stripes in the embryo.
569:
331:, these eRNAs are usually protected by their
280:core promoters and promoter-proximal elements
3237:Seminars in Cell & Developmental Biology
3226:
3224:
2858:
2809:
2711:
2668:
2662:
2603:
2562:
2513:
2467:
2374:
2368:
2317:
2274:
2268:
1961:Eichenlaub MP, Ettwiller L (November 2011).
1569:Annual Review of Genomics and Human Genetics
1496:
1436:
1401:
1011:Annual Review of Genomics and Human Genetics
951:
949:
947:
945:
451:Traditionally, enhancers were identified by
2330:Frontiers in Cell and Developmental Biology
1705:
1296:"Characterization of enhancer fragments in
616:
422:
327:(eRNAs) as illustrated in the Figure. Like
4253:
4239:
4222:Enhancer discovery and characterization.
228:
4197:at the U.S. National Library of Medicine
4167:
4080:
4031:
3982:
3925:
3876:
3827:
3778:
3768:
3719:
3619:
3562:
3505:
3413:
3372:
3323:
3256:
3221:
3204:
3155:
3145:
3061:
3012:
2890:
2841:
2792:
2743:
2694:
2645:
2635:
2586:
2545:
2450:
2420:
2418:
2351:
2341:
2300:
2251:
2202:
2145:
2088:
1988:
1978:
1937:
1888:
1839:
1820:Nature Structural & Molecular Biology
1790:
1780:
1645:
1580:
1539:
1479:
1384:
1335:
1325:
1315:
1138:
1120:
1079:
1022:
981:
942:
386:, and possibly also modifications in the
3087:Roux's Archives of Developmental Biology
846:Designing enhancers in synthetic biology
232:
38:
31:
256:(RNAP II) bound to the promoter at the
221:and their location can be predicted by
202:may have effects although they are not
14:
4688:
2914:Arnosti DN, Kulkarni MM (April 2005).
2675:Nature Reviews. Molecular Cell Biology
2415:
2281:Nature Reviews. Molecular Cell Biology
1005:Maston GA, Evans SK, Green MR (2006).
238:Regulation of transcription in mammals
4399:Histone acetylation and deacetylation
4234:
3347:Norris DP, Robertson EJ (June 1999).
1582:10.1146/annurev.genom.7.080505.115623
1024:10.1146/annurev.genom.7.080505.115623
958:"Enhancers: five essential questions"
700:Evolution of developmental mechanisms
672:is specified early in development by
225:against this family of coactivators.
206:. Enhancers can also be found at the
4012:Cytokine & Growth Factor Reviews
3042:Cellular and Molecular Life Sciences
2005:
711:
167:. The same mechanism holds true for
2907:
2669:Allen BL, Taatjes DJ (March 2015).
2275:Haberle V, Stark A (October 2018).
784:gene, whose product produces black
447:Identification and characterization
131:. Since 2022, scientists have used
24:
706:evolutionary developmental biology
97:) to increase the likelihood that
25:
4707:
4188:
2571:"The Human Transcription Factors"
840:surveillance by the immune system
637:expression in the portion of the
3179:Suryamohan K, Halfon MS (2014).
2923:Journal of Cellular Biochemistry
574:The enhancers determining early
4603:Archaeal transcription factor B
4135:
4048:
3999:
3942:
3893:
3844:
3795:
3736:
3687:
3636:
3579:
3522:
3473:
3430:
3389:
3340:
3291:
3273:
3172:
3121:
3078:
3029:
2972:
2760:
2219:
2162:
2105:
2056:
1954:
1856:
1807:
1748:
1662:
1605:
1556:
1352:
627:transforming growth factor-beta
60:Transcription Activator Protein
1287:
1198:
1155:
1096:
1060:Molecular and Cellular Biology
1047:
998:
891:
13:
1:
4024:10.1016/j.cytogfr.2015.06.001
1327:10.1080/19336934.2022.2126259
884:
161:general transcription factors
3820:10.1016/j.molcel.2014.08.024
3249:10.1016/j.semcdb.2006.12.014
1980:10.1371/journal.pbio.1001188
1265:10.1016/0092-8674(83)90014-4
1219:10.1016/0092-8674(83)90015-6
362:Examples in the human genome
138:
7:
3498:10.1016/j.ydbio.2010.07.032
3415:10.1016/j.ydbio.2010.10.036
920:10.1126/science.281.5373.60
872:
829:, the transcription factor
338:
178:An enhancer may be located
147:cells the structure of the
10:
4712:
4279:Transcriptional regulation
4160:10.1038/s41588-021-00782-6
4121:10.1038/s41588-022-01048-5
3712:10.1016/j.cell.2013.03.035
2785:10.1038/s41587-022-01211-7
2588:10.1016/j.cell.2018.01.029
2538:10.1016/j.cell.2017.11.008
2081:10.1016/j.cell.2009.06.049
1832:10.1038/s41594-023-01027-2
1532:10.1038/s41586-023-06936-2
1472:10.1038/s41586-023-06905-9
1422:10.1038/s41588-022-01048-5
817:In inflammation and cancer
570:In segmentation of insects
487:Next-generation sequencing
413:
282:that are located near the
121:immunoglobulin heavy chain
4646:
4611:
4585:
4510:
4476:Transcription coregulator
4468:
4445:
4422:
4412:Histone acetyltransferase
4382:Histone methyltransferase
4360:Histone-modifying enzymes
4358:
4351:
4286:
4277:
4195:Enhancer+Elements,Genetic
3918:10.1126/scisignal.aaf1129
3555:10.1016/j.cub.2010.07.043
3054:10.1007/s00018-012-0990-9
2491:10.1038/s41576-019-0128-0
2443:10.1038/s41593-020-0634-6
2343:10.3389/fcell.2020.592164
2293:10.1038/s41580-018-0028-8
1317:10.1101/2022.08.01.502399
1122:10.1186/s13059-017-1285-0
704:One theme of research in
529:or lacZ to determine the
527:green fluorescent protein
366:
284:transcription start sites
244:DNA region of its target
4199:Medical Subject Headings
2479:Nature Reviews. Genetics
2381:Nature Reviews. Genetics
962:Nature Reviews. Genetics
797:, which is activated by
750:chromosomal fragile site
617:In vertebrate patterning
423:In developmental biology
377:Human Accelerated Region
258:transcription start site
4577:Internal control region
4220:ENCODE threads explorer
3770:10.1073/pnas.1317023110
3612:10.1126/science.1182213
3451:10.1242/dev.129.14.3455
3353:Genes & Development
2724:Genes & Development
2637:10.1073/pnas.1804663115
1782:10.1073/pnas.2036346100
1734:10.1126/science.3006253
1691:10.1126/science.3917575
1184:10.1126/science.6306772
856:artificial intelligence
581:Drosophila melanogaster
458:Drosophila melanogaster
433:cis-regulatory elements
276:cis-regulatory elements
229:Role in gene expression
217:Enhancers are bound by
133:artificial intelligence
3365:10.1101/gad.13.12.1575
2871:Nucleic Acids Research
2736:10.1101/gad.308619.117
1869:Nucleic Acids Research
1377:10.26508/lsa.202302244
768:wing pattern evolution
483:cis-regulatory modules
371:HACNS1 (also known as
271:
180:upstream or downstream
70:
36:
4670:Intrinsic termination
4435:DNA methyltransferase
3869:10.1101/gr.188300.114
3486:Developmental Biology
3402:Developmental Biology
1930:10.1101/gr.133546.111
1365:Life Science Alliance
795:Wnt signaling pathway
519:sequence conservation
441:transcription factors
394:that allow humans to
297:transcription factors
263:transcription factors
236:
107:transcription factors
89:that can be bound by
42:
35:
4447:Chromatin remodeling
4073:10.1038/onc.2013.179
2834:10.1093/emboj/cdg028
2773:Nature Biotechnology
2532:(7): 1573–1588.e28.
1072:10.1128/MCB.01127-12
867:deep neural networks
778:Drosophila guttifera
515:comparative genomics
511:comparative genomics
431:. Enhancers work as
252:(mRNA) synthesis by
81:is a short (50–1500
4404:Histone deacetylase
4394:Histone demethylase
4378:Histone methylation
3975:10.1038/nature14154
3967:2015Natur.520..558V
3761:2013PNAS..11017921P
3755:(44): 17921–17926.
3665:10.1038/nature08896
3657:2010Natur.464.1143W
3651:(7292): 1143–1148.
3604:2010Sci...327..302C
3547:2010CBio...20.1562P
3147:10.7554/eLife.32341
3005:10.1038/nature09774
2997:2011Natur.471..216M
2883:10.1093/nar/gkaa671
2628:2018PNAS..115E7222G
2622:(30): E7222–E7230.
2431:Nature Neuroscience
2195:10.1038/nature07730
2187:2009Natur.457..854V
2138:10.1038/nature07829
2130:2009Natur.459..108H
2034:10.1038/nature03574
2026:2005Natur.435..637S
1881:10.1093/nar/gkp1124
1773:2003PNAS..10013887K
1767:(24): 13887–13891.
1726:1986Sci...232...76S
1683:1985Sci...227..266M
1638:10.1038/nature13138
1630:2014Natur.507..371S
1524:2024Natur.626..212T
1464:2024Natur.626..207D
1176:1983Sci...221..663M
912:1998Sci...281...60.
660:Establishing three
375:and located in the
3316:10.1242/dev.036160
3099:10.1007/BF00188752
523:non-coding regions
272:
192:activator proteins
71:
37:
4683:
4682:
4638:RNA polymerase II
4506:
4505:
4464:
4463:
4067:(18): 2395–2404.
3961:(7548): 558–562.
3906:Science Signaling
3598:(5963): 302–305.
3541:(17): 1562–1567.
3445:(14): 3455–3468.
3359:(12): 1575–1588.
3048:(21): 3613–3634.
2991:(7337): 216–219.
2935:10.1002/jcb.20352
2877:(17): 9550–9570.
2181:(7231): 854–858.
2124:(7243): 108–112.
2020:(7042): 637–645.
1677:(4684): 266–270.
1624:(7492): 371–375.
1518:(7997): 212–220.
1458:(7997): 207–211.
1371:(3): e202302244.
1170:(4611): 663–665.
1066:(24): 4892–4897.
860:transfer learning
485:(CRM) discovery.
254:RNA polymerase II
165:RNA polymerase II
16:(Redirected from
4703:
4560:Response element
4543:Response element
4356:
4355:
4284:
4283:
4255:
4248:
4241:
4232:
4231:
4182:
4181:
4171:
4139:
4133:
4132:
4104:
4095:
4094:
4084:
4052:
4046:
4045:
4035:
4003:
3997:
3996:
3986:
3946:
3940:
3939:
3929:
3897:
3891:
3890:
3880:
3863:(9): 1281–1294.
3848:
3842:
3841:
3831:
3799:
3793:
3792:
3782:
3772:
3740:
3734:
3733:
3723:
3691:
3685:
3684:
3640:
3634:
3633:
3623:
3583:
3577:
3576:
3566:
3526:
3520:
3519:
3509:
3477:
3471:
3470:
3434:
3428:
3427:
3417:
3393:
3387:
3386:
3376:
3344:
3338:
3337:
3327:
3295:
3289:
3288:
3287:on 10 July 2015.
3277:
3271:
3270:
3260:
3228:
3219:
3218:
3208:
3197:10.1002/wdev.168
3176:
3170:
3169:
3159:
3149:
3125:
3119:
3118:
3082:
3076:
3075:
3065:
3033:
3027:
3026:
3016:
2976:
2970:
2969:
2967:
2965:
2959:
2953:. Archived from
2920:
2911:
2905:
2904:
2894:
2862:
2856:
2855:
2845:
2822:The EMBO Journal
2813:
2807:
2806:
2796:
2779:(7): 1056–1065.
2764:
2758:
2757:
2747:
2715:
2709:
2708:
2698:
2666:
2660:
2659:
2649:
2639:
2607:
2601:
2600:
2590:
2566:
2560:
2559:
2549:
2517:
2511:
2510:
2474:
2465:
2464:
2454:
2422:
2413:
2412:
2372:
2366:
2365:
2355:
2345:
2321:
2315:
2314:
2304:
2272:
2266:
2265:
2255:
2223:
2217:
2216:
2206:
2166:
2160:
2159:
2149:
2109:
2103:
2102:
2092:
2075:(5): 1019–1031.
2060:
2054:
2053:
2009:
2003:
2002:
1992:
1982:
1973:(11): e1001188.
1958:
1952:
1951:
1941:
1924:(6): 1059–1068.
1909:
1903:
1902:
1892:
1875:(4): 1071–1085.
1860:
1854:
1853:
1843:
1811:
1805:
1804:
1794:
1784:
1752:
1746:
1745:
1709:
1703:
1702:
1666:
1660:
1659:
1649:
1609:
1603:
1602:
1584:
1560:
1554:
1553:
1543:
1503:
1494:
1493:
1483:
1443:
1434:
1433:
1405:
1399:
1398:
1388:
1356:
1350:
1349:
1339:
1329:
1319:
1298:Drosophila robo2
1291:
1285:
1284:
1248:
1239:
1238:
1202:
1196:
1195:
1159:
1153:
1152:
1142:
1124:
1100:
1094:
1093:
1083:
1051:
1045:
1044:
1026:
1002:
996:
995:
985:
953:
940:
939:
895:
879:Shadow enhancers
647:fork head domain
639:primitive streak
196:mediator complex
129:myelosuppression
109:. Enhancers are
101:of a particular
63:Mediator Protein
21:
4711:
4710:
4706:
4705:
4704:
4702:
4701:
4700:
4696:Gene expression
4686:
4685:
4684:
4679:
4654:
4648:
4642:
4607:
4581:
4502:
4460:
4441:
4424:DNA methylation
4418:
4362:
4347:
4273:
4259:
4191:
4186:
4185:
4148:Nature Genetics
4140:
4136:
4109:Nature Genetics
4105:
4098:
4053:
4049:
4004:
4000:
3947:
3943:
3898:
3894:
3857:Genome Research
3849:
3845:
3800:
3796:
3741:
3737:
3692:
3688:
3641:
3637:
3584:
3580:
3535:Current Biology
3527:
3523:
3478:
3474:
3435:
3431:
3394:
3390:
3345:
3341:
3296:
3292:
3279:
3278:
3274:
3229:
3222:
3177:
3173:
3126:
3122:
3083:
3079:
3034:
3030:
2977:
2973:
2963:
2961:
2960:on 21 July 2006
2957:
2918:
2912:
2908:
2863:
2859:
2814:
2810:
2765:
2761:
2716:
2712:
2687:10.1038/nrm3951
2667:
2663:
2608:
2604:
2567:
2563:
2518:
2514:
2475:
2468:
2423:
2416:
2393:10.1038/nrg3207
2373:
2369:
2322:
2318:
2287:(10): 621–637.
2273:
2269:
2232:Nature Genetics
2224:
2220:
2167:
2163:
2110:
2106:
2061:
2057:
2010:
2006:
1959:
1955:
1918:Genome Research
1910:
1906:
1861:
1857:
1826:(7): 991–1000.
1812:
1808:
1753:
1749:
1720:(4746): 76–80.
1710:
1706:
1667:
1663:
1610:
1606:
1561:
1557:
1504:
1497:
1444:
1437:
1410:Nature Genetics
1406:
1402:
1357:
1353:
1292:
1288:
1249:
1242:
1203:
1199:
1160:
1156:
1101:
1097:
1052:
1048:
1003:
999:
974:10.1038/nrg3458
954:
943:
906:(5373): 60–63.
896:
892:
887:
875:
848:
836:malignant cells
823:super-enhancers
819:
770:
725:
717:
702:
693:
619:
594:maternal effect
590:pair rule genes
572:
552:protein folding
449:
429:gene expression
425:
416:
369:
364:
352:point mutations
341:
267:phosphorylation
231:
141:
69:
28:
23:
22:
15:
12:
11:
5:
4709:
4699:
4698:
4681:
4680:
4678:
4677:
4672:
4667:
4661:
4659:
4644:
4643:
4641:
4640:
4634:RNA polymerase
4628:
4622:RNA polymerase
4615:
4613:
4609:
4608:
4606:
4605:
4600:
4595:
4589:
4587:
4583:
4582:
4580:
4579:
4574:
4569:
4564:
4563:
4562:
4557:
4547:
4546:
4545:
4540:
4535:
4530:
4525:
4514:
4512:
4508:
4507:
4504:
4503:
4501:
4500:
4495:
4494:
4493:
4488:
4483:
4472:
4470:
4466:
4465:
4462:
4461:
4459:
4458:
4452:
4450:
4443:
4442:
4440:
4439:
4438:
4437:
4429:
4427:
4420:
4419:
4417:
4416:
4415:
4414:
4409:
4396:
4391:
4390:
4389:
4374:
4372:
4353:
4349:
4348:
4346:
4345:
4344:
4343:
4338:
4328:
4327:
4326:
4321:
4316:
4311:
4306:
4301:
4290:
4288:
4281:
4275:
4274:
4258:
4257:
4250:
4243:
4235:
4229:
4228:
4217:
4212:
4207:
4202:
4190:
4189:External links
4187:
4184:
4183:
4154:(3): 354–366.
4134:
4115:(5): 613–624.
4096:
4047:
4018:(4): 389–403.
3998:
3941:
3912:(457): ra120.
3892:
3843:
3814:(2): 219–231.
3808:Molecular Cell
3794:
3735:
3706:(2): 307–319.
3686:
3635:
3578:
3521:
3492:(2): 346–355.
3472:
3429:
3408:(2): 350–362.
3388:
3339:
3290:
3272:
3243:(1): 140–152.
3220:
3171:
3120:
3093:(4): 194–220.
3077:
3028:
2971:
2929:(5): 890–898.
2906:
2857:
2828:(2): 281–291.
2808:
2759:
2710:
2681:(3): 155–166.
2661:
2602:
2581:(4): 650–665.
2561:
2512:
2485:(8): 437–455.
2466:
2437:(6): 707–717.
2414:
2387:(9): 613–626.
2367:
2316:
2267:
2244:10.1038/ng.650
2238:(9): 806–810.
2218:
2161:
2104:
2055:
2004:
1953:
1904:
1855:
1806:
1747:
1704:
1661:
1604:
1555:
1495:
1435:
1416:(5): 613–624.
1400:
1351:
1310:(1): 312–346.
1286:
1259:(3): 717–728.
1240:
1213:(3): 729–740.
1197:
1154:
1109:Genome Biology
1095:
1046:
997:
968:(4): 288–295.
941:
889:
888:
886:
883:
882:
881:
874:
871:
847:
844:
818:
815:
769:
762:
716:
710:
701:
698:
692:
689:
643:primitive node
618:
615:
571:
568:
448:
445:
424:
421:
415:
412:
368:
365:
363:
360:
359:
358:
355:
340:
337:
230:
227:
140:
137:
68:
67:
66:RNA Polymerase
64:
61:
58:
55:
52:
49:
45:
26:
9:
6:
4:
3:
2:
4708:
4697:
4694:
4693:
4691:
4676:
4673:
4671:
4668:
4666:
4663:
4662:
4660:
4657:
4652:
4645:
4639:
4635:
4632:
4629:
4627:
4623:
4620:
4617:
4616:
4614:
4610:
4604:
4601:
4599:
4596:
4594:
4591:
4590:
4588:
4584:
4578:
4575:
4573:
4570:
4568:
4565:
4561:
4558:
4556:
4553:
4552:
4551:
4548:
4544:
4541:
4539:
4536:
4534:
4531:
4529:
4526:
4524:
4521:
4520:
4519:
4516:
4515:
4513:
4509:
4499:
4496:
4492:
4489:
4487:
4484:
4482:
4479:
4478:
4477:
4474:
4473:
4471:
4467:
4457:
4454:
4453:
4451:
4448:
4444:
4436:
4433:
4432:
4431:
4430:
4428:
4425:
4421:
4413:
4410:
4408:
4405:
4402:
4401:
4400:
4397:
4395:
4392:
4388:
4385:
4384:
4383:
4379:
4376:
4375:
4373:
4370:
4366:
4361:
4357:
4354:
4350:
4342:
4341:trp repressor
4339:
4337:
4336:lac repressor
4334:
4333:
4332:
4329:
4325:
4322:
4320:
4317:
4315:
4312:
4310:
4307:
4305:
4302:
4300:
4297:
4296:
4295:
4292:
4291:
4289:
4285:
4282:
4280:
4276:
4271:
4267:
4263:
4262:Transcription
4256:
4251:
4249:
4244:
4242:
4237:
4236:
4233:
4227:
4226:
4221:
4218:
4216:
4213:
4211:
4208:
4206:
4203:
4200:
4196:
4193:
4192:
4179:
4175:
4170:
4165:
4161:
4157:
4153:
4149:
4145:
4138:
4130:
4126:
4122:
4118:
4114:
4110:
4103:
4101:
4092:
4088:
4083:
4078:
4074:
4070:
4066:
4062:
4058:
4051:
4043:
4039:
4034:
4029:
4025:
4021:
4017:
4013:
4009:
4002:
3994:
3990:
3985:
3980:
3976:
3972:
3968:
3964:
3960:
3956:
3952:
3945:
3937:
3933:
3928:
3923:
3919:
3915:
3911:
3907:
3903:
3896:
3888:
3884:
3879:
3874:
3870:
3866:
3862:
3858:
3854:
3847:
3839:
3835:
3830:
3825:
3821:
3817:
3813:
3809:
3805:
3798:
3790:
3786:
3781:
3776:
3771:
3766:
3762:
3758:
3754:
3750:
3746:
3739:
3731:
3727:
3722:
3717:
3713:
3709:
3705:
3701:
3697:
3690:
3682:
3678:
3674:
3670:
3666:
3662:
3658:
3654:
3650:
3646:
3639:
3631:
3627:
3622:
3617:
3613:
3609:
3605:
3601:
3597:
3593:
3589:
3582:
3574:
3570:
3565:
3560:
3556:
3552:
3548:
3544:
3540:
3536:
3532:
3525:
3517:
3513:
3508:
3503:
3499:
3495:
3491:
3487:
3483:
3476:
3468:
3464:
3460:
3456:
3452:
3448:
3444:
3440:
3433:
3425:
3421:
3416:
3411:
3407:
3403:
3399:
3392:
3384:
3380:
3375:
3370:
3366:
3362:
3358:
3354:
3350:
3343:
3335:
3331:
3326:
3321:
3317:
3313:
3309:
3305:
3301:
3294:
3286:
3282:
3276:
3268:
3264:
3259:
3254:
3250:
3246:
3242:
3238:
3234:
3227:
3225:
3216:
3212:
3207:
3202:
3198:
3194:
3190:
3186:
3182:
3175:
3167:
3163:
3158:
3153:
3148:
3143:
3139:
3135:
3131:
3124:
3116:
3112:
3108:
3104:
3100:
3096:
3092:
3088:
3081:
3073:
3069:
3064:
3059:
3055:
3051:
3047:
3043:
3039:
3032:
3024:
3020:
3015:
3010:
3006:
3002:
2998:
2994:
2990:
2986:
2982:
2975:
2956:
2952:
2948:
2944:
2940:
2936:
2932:
2928:
2924:
2917:
2910:
2902:
2898:
2893:
2888:
2884:
2880:
2876:
2872:
2868:
2861:
2853:
2849:
2844:
2839:
2835:
2831:
2827:
2823:
2819:
2812:
2804:
2800:
2795:
2790:
2786:
2782:
2778:
2774:
2770:
2763:
2755:
2751:
2746:
2741:
2737:
2733:
2729:
2725:
2721:
2714:
2706:
2702:
2697:
2692:
2688:
2684:
2680:
2676:
2672:
2665:
2657:
2653:
2648:
2643:
2638:
2633:
2629:
2625:
2621:
2617:
2613:
2606:
2598:
2594:
2589:
2584:
2580:
2576:
2572:
2565:
2557:
2553:
2548:
2543:
2539:
2535:
2531:
2527:
2523:
2516:
2508:
2504:
2500:
2496:
2492:
2488:
2484:
2480:
2473:
2471:
2462:
2458:
2453:
2448:
2444:
2440:
2436:
2432:
2428:
2421:
2419:
2410:
2406:
2402:
2398:
2394:
2390:
2386:
2382:
2378:
2371:
2363:
2359:
2354:
2349:
2344:
2339:
2335:
2331:
2327:
2320:
2312:
2308:
2303:
2298:
2294:
2290:
2286:
2282:
2278:
2271:
2263:
2259:
2254:
2249:
2245:
2241:
2237:
2233:
2229:
2222:
2214:
2210:
2205:
2200:
2196:
2192:
2188:
2184:
2180:
2176:
2172:
2165:
2157:
2153:
2148:
2143:
2139:
2135:
2131:
2127:
2123:
2119:
2115:
2108:
2100:
2096:
2091:
2086:
2082:
2078:
2074:
2070:
2066:
2059:
2051:
2047:
2043:
2039:
2035:
2031:
2027:
2023:
2019:
2015:
2008:
2000:
1996:
1991:
1986:
1981:
1976:
1972:
1968:
1964:
1957:
1949:
1945:
1940:
1935:
1931:
1927:
1923:
1919:
1915:
1908:
1900:
1896:
1891:
1886:
1882:
1878:
1874:
1870:
1866:
1859:
1851:
1847:
1842:
1837:
1833:
1829:
1825:
1821:
1817:
1810:
1802:
1798:
1793:
1788:
1783:
1778:
1774:
1770:
1766:
1762:
1758:
1751:
1743:
1739:
1735:
1731:
1727:
1723:
1719:
1715:
1708:
1700:
1696:
1692:
1688:
1684:
1680:
1676:
1672:
1665:
1657:
1653:
1648:
1643:
1639:
1635:
1631:
1627:
1623:
1619:
1615:
1608:
1600:
1596:
1592:
1588:
1583:
1578:
1574:
1570:
1566:
1559:
1551:
1547:
1542:
1537:
1533:
1529:
1525:
1521:
1517:
1513:
1509:
1502:
1500:
1491:
1487:
1482:
1477:
1473:
1469:
1465:
1461:
1457:
1453:
1449:
1442:
1440:
1431:
1427:
1423:
1419:
1415:
1411:
1404:
1396:
1392:
1387:
1382:
1378:
1374:
1370:
1366:
1362:
1355:
1347:
1343:
1338:
1333:
1328:
1323:
1318:
1313:
1309:
1305:
1301:
1299:
1290:
1282:
1278:
1274:
1270:
1266:
1262:
1258:
1254:
1247:
1245:
1236:
1232:
1228:
1224:
1220:
1216:
1212:
1208:
1201:
1193:
1189:
1185:
1181:
1177:
1173:
1169:
1165:
1158:
1150:
1146:
1141:
1136:
1132:
1128:
1123:
1118:
1114:
1110:
1106:
1099:
1091:
1087:
1082:
1077:
1073:
1069:
1065:
1061:
1057:
1050:
1042:
1038:
1034:
1030:
1025:
1020:
1016:
1012:
1008:
1001:
993:
989:
984:
979:
975:
971:
967:
963:
959:
952:
950:
948:
946:
937:
933:
929:
925:
921:
917:
913:
909:
905:
901:
894:
890:
880:
877:
876:
870:
868:
863:
861:
857:
852:
843:
841:
837:
832:
828:
824:
814:
812:
808:
804:
800:
796:
791:
787:
783:
779:
775:
767:
761:
759:
755:
751:
747:
743:
739:
734:
730:
729:
722:
715:
709:
707:
697:
688:
686:
681:
677:
676:
671:
667:
663:
658:
656:
652:
648:
644:
640:
636:
632:
628:
624:
614:
612:
608:
604:
600:
595:
591:
587:
583:
582:
577:
567:
565:
561:
557:
553:
549:
545:
544:hybridization
542:
541:
536:
532:
528:
524:
520:
516:
512:
508:
507:Dam methylase
504:
500:
496:
492:
488:
484:
479:
477:
474:
470:
467:
466:
461:
459:
454:
453:enhancer trap
444:
442:
438:
437:transcription
434:
430:
420:
411:
409:
405:
401:
397:
393:
389:
385:
382:
378:
374:
356:
353:
349:
348:Enhanceosomes
346:
345:
344:
336:
334:
330:
326:
325:Enhancer RNAs
321:
319:
315:
311:
305:
301:
298:
294:
290:
285:
281:
277:
268:
264:
259:
255:
251:
250:messenger RNA
247:
243:
239:
235:
226:
224:
220:
215:
213:
209:
205:
201:
200:polymorphisms
197:
193:
189:
185:
184:transcription
181:
176:
174:
170:
166:
162:
158:
154:
150:
146:
136:
134:
130:
126:
122:
117:
115:
113:
108:
104:
100:
99:transcription
96:
92:
88:
84:
80:
76:
65:
62:
59:
56:
53:
50:
47:
46:
41:
34:
30:
19:
4549:
4223:
4151:
4147:
4137:
4112:
4108:
4064:
4060:
4050:
4015:
4011:
4001:
3958:
3954:
3944:
3909:
3905:
3895:
3860:
3856:
3846:
3811:
3807:
3797:
3752:
3748:
3738:
3703:
3699:
3689:
3648:
3644:
3638:
3595:
3591:
3581:
3538:
3534:
3524:
3489:
3485:
3475:
3442:
3438:
3432:
3405:
3401:
3391:
3356:
3352:
3342:
3307:
3303:
3293:
3285:the original
3275:
3240:
3236:
3191:(2): 59–84.
3188:
3184:
3174:
3137:
3133:
3123:
3090:
3086:
3080:
3045:
3041:
3031:
2988:
2984:
2974:
2962:. Retrieved
2955:the original
2926:
2922:
2909:
2874:
2870:
2860:
2825:
2821:
2811:
2776:
2772:
2762:
2730:(1): 42–57.
2727:
2723:
2713:
2678:
2674:
2664:
2619:
2615:
2605:
2578:
2574:
2564:
2529:
2525:
2515:
2482:
2478:
2434:
2430:
2384:
2380:
2370:
2333:
2329:
2319:
2284:
2280:
2270:
2235:
2231:
2221:
2178:
2174:
2164:
2121:
2117:
2107:
2072:
2068:
2058:
2017:
2013:
2007:
1970:
1967:PLOS Biology
1966:
1956:
1921:
1917:
1907:
1872:
1868:
1858:
1823:
1819:
1809:
1764:
1760:
1750:
1717:
1713:
1707:
1674:
1670:
1664:
1621:
1617:
1607:
1575:(1): 29–59.
1572:
1568:
1558:
1515:
1511:
1455:
1451:
1413:
1409:
1403:
1368:
1364:
1354:
1307:
1303:
1297:
1289:
1256:
1252:
1210:
1206:
1200:
1167:
1163:
1157:
1112:
1108:
1098:
1063:
1059:
1049:
1014:
1010:
1000:
965:
961:
903:
899:
893:
864:
854:Since 2022,
853:
849:
827:inflammation
820:
810:
806:
798:
789:
781:
777:
773:
771:
765:
757:
745:
741:
726:
718:
713:
712:Stickleback
703:
694:
684:
679:
673:
666:gastrulation
659:
650:
634:
630:
620:
610:
606:
602:
599:even-skipped
598:
579:
576:segmentation
573:
555:
538:
530:
480:
463:
456:
450:
426:
417:
370:
342:
322:
306:
302:
278:, including
273:
237:
216:
177:
142:
118:
111:
85:) region of
78:
72:
29:
4647:Termination
4523:Pribnow box
4491:Corepressor
4486:Coactivator
4287:prokaryotic
3439:Development
3310:(1): 5–13.
3304:Development
724:tetrapods).
721:stickleback
662:germ layers
548:translation
408:chimpanzees
153:prokaryotic
4675:Rho factor
4665:Terminator
4656:eukaryotic
4631:eukaryotic
4612:Elongation
4598:Eukaryotic
4586:Initiation
4369:nucleosome
4352:eukaryotic
4324:gal operon
4319:ara operon
4314:Gua Operon
4309:gab operon
4304:trp operon
4299:lac operon
4270:Eukaryotic
2377:Furlong EE
2336:: 592164.
1115:(1): 165.
885:References
774:Drosophila
766:Drosophila
754:DNA repair
517:approach,
495:epigenetic
476:transposon
293:insulators
212:chromosome
204:translated
188:base pairs
173:repressors
145:eukaryotic
95:activators
4651:bacterial
4619:bacterial
4593:Bacterial
4567:Insulator
4511:Promotion
4481:Activator
4331:Repressor
4266:Bacterial
2507:152283312
2409:205485256
2375:Spitz F,
1131:1474-760X
1017:: 29–59.
803:phenotype
560:zebrafish
499:DNase-seq
473:P element
404:evolution
289:silencers
169:silencers
149:chromatin
139:Locations
18:Enhancers
4690:Category
4572:Silencer
4550:Enhancer
4538:CAAT box
4528:TATA box
4518:Promoter
4205:TFSEARCH
4178:33603233
4129:35551305
4091:23686307
4061:Oncogene
4042:26119834
3993:25686607
3936:27923915
3887:26113076
3838:25263595
3789:24127591
3730:23582322
3673:20376004
3630:20007865
3573:20797865
3516:20692247
3467:24426844
3459:12091315
3424:21047506
3383:10385626
3334:20023155
3267:17276707
3215:25704908
3166:29481322
3115:25759655
3107:28305845
3072:22538991
3023:21390129
2964:8 August
2951:32405464
2943:15696541
2901:32810208
2852:12514134
2803:35177836
2754:29378788
2705:25693131
2656:29987030
2597:29425488
2556:29224777
2499:31086298
2461:32451484
2401:22868264
2362:33102493
2311:29946135
2262:20729851
2213:19212405
2156:19295514
2099:19698979
2042:15880101
1999:22069375
1948:22442009
1899:19969543
1850:37430065
1841:10352134
1801:14623974
1656:24646999
1599:12346247
1591:16719718
1550:38086419
1541:10830415
1490:38086418
1481:10830412
1430:35551305
1395:38228368
1386:10796589
1346:36217698
1281:40313833
1235:23981549
1149:28870212
1090:23045397
1041:12346247
1033:16719718
992:23503198
936:11666739
873:See also
811:wingless
799:wingless
733:homeobox
670:endoderm
655:mesoderm
586:gap gene
503:ATAC-seq
339:Theories
318:Mediator
242:promoter
223:ChIP-seq
219:p300-CBP
157:promoter
91:proteins
79:enhancer
75:genetics
54:Promoter
51:Enhancer
4498:Inducer
4365:histone
4169:8812996
4082:3913736
4033:4526340
3984:4409450
3963:Bibcode
3927:5260935
3878:4561488
3829:4224636
3780:3816444
3757:Bibcode
3721:3653129
3681:4407744
3653:Bibcode
3621:3109066
3600:Bibcode
3592:Science
3564:4257487
3543:Bibcode
3507:2945415
3325:2796927
3258:1855162
3206:4339228
3157:5826290
3063:3474909
3014:3071156
2993:Bibcode
2892:7515708
2794:9288987
2745:5828394
2696:4963239
2647:6065035
2624:Bibcode
2547:5785279
2452:7558717
2353:7554316
2302:6205604
2253:3138496
2204:2745234
2183:Bibcode
2147:2910248
2126:Bibcode
2090:2750862
2050:1755326
2022:Bibcode
1990:3206014
1939:3371700
1890:2831330
1769:Bibcode
1742:3006253
1722:Bibcode
1714:Science
1699:3917575
1679:Bibcode
1671:Science
1647:4113484
1626:Bibcode
1520:Bibcode
1460:Bibcode
1337:9559326
1312:bioRxiv
1273:6409417
1227:6409418
1192:6306772
1172:Bibcode
1164:Science
1140:5584004
1081:3510544
983:4445073
928:9679020
908:Bibcode
900:Science
786:melanin
664:during
540:in situ
531:in vivo
414:GADD45G
114:-acting
4294:Operon
4225:Nature
4210:JASPAR
4201:(MeSH)
4176:
4166:
4127:
4089:
4079:
4040:
4030:
3991:
3981:
3955:Nature
3934:
3924:
3885:
3875:
3836:
3826:
3787:
3777:
3728:
3718:
3679:
3671:
3645:Nature
3628:
3618:
3571:
3561:
3514:
3504:
3465:
3457:
3422:
3381:
3374:316799
3371:
3332:
3322:
3265:
3255:
3213:
3203:
3164:
3154:
3113:
3105:
3070:
3060:
3021:
3011:
2985:Nature
2949:
2941:
2899:
2889:
2850:
2843:140103
2840:
2801:
2791:
2752:
2742:
2703:
2693:
2654:
2644:
2595:
2554:
2544:
2505:
2497:
2459:
2449:
2407:
2399:
2360:
2350:
2309:
2299:
2260:
2250:
2211:
2201:
2175:Nature
2154:
2144:
2118:Nature
2097:
2087:
2048:
2040:
2014:Nature
1997:
1987:
1946:
1936:
1897:
1887:
1848:
1838:
1799:
1792:283516
1789:
1740:
1697:
1654:
1644:
1618:Nature
1597:
1589:
1548:
1538:
1512:Nature
1488:
1478:
1452:Nature
1428:
1393:
1383:
1344:
1334:
1314:
1279:
1271:
1233:
1225:
1190:
1147:
1137:
1129:
1088:
1078:
1039:
1031:
990:
980:
934:
926:
807:yellow
805:, the
790:yellow
782:yellow
738:allele
400:genome
373:CENTG2
367:HACNS1
333:5′ cap
208:exonic
125:intron
4555:E-box
4407:HDAC1
4215:ReMap
3677:S2CID
3463:S2CID
3134:eLife
3111:S2CID
2958:(PDF)
2947:S2CID
2919:(PDF)
2503:S2CID
2405:S2CID
2046:S2CID
1595:S2CID
1277:S2CID
1231:S2CID
1037:S2CID
932:S2CID
831:NF-κB
758:Pitx1
746:Pitx1
742:Pitx1
731:is a
728:Pitx1
714:Pitx1
685:Gata4
680:Gata4
675:Gata4
651:Nodal
635:Nodal
631:Nodal
623:Nodal
564:BLAST
388:ankle
384:thumb
381:human
329:mRNAs
77:, an
4626:rpoB
4469:both
4456:CHD7
4387:EZH2
4174:PMID
4125:PMID
4087:PMID
4038:PMID
3989:PMID
3932:PMID
3883:PMID
3834:PMID
3785:PMID
3726:PMID
3700:Cell
3669:PMID
3626:PMID
3569:PMID
3512:PMID
3455:PMID
3420:PMID
3379:PMID
3330:PMID
3263:PMID
3211:PMID
3162:PMID
3103:PMID
3068:PMID
3019:PMID
2966:2019
2939:PMID
2897:PMID
2848:PMID
2799:PMID
2750:PMID
2701:PMID
2652:PMID
2593:PMID
2575:Cell
2552:PMID
2526:Cell
2495:PMID
2457:PMID
2397:PMID
2358:PMID
2307:PMID
2258:PMID
2209:PMID
2152:PMID
2095:PMID
2069:Cell
2038:PMID
1995:PMID
1944:PMID
1895:PMID
1846:PMID
1797:PMID
1738:PMID
1695:PMID
1652:PMID
1587:PMID
1546:PMID
1486:PMID
1426:PMID
1391:PMID
1342:PMID
1269:PMID
1253:Cell
1223:PMID
1207:Cell
1188:PMID
1145:PMID
1127:ISSN
1086:PMID
1029:PMID
988:PMID
924:PMID
858:and
625:, a
550:and
535:mRNA
469:gene
465:lacZ
396:walk
392:foot
310:CTCF
246:gene
163:and
103:gene
57:Gene
4533:BRE
4164:PMC
4156:doi
4117:doi
4077:PMC
4069:doi
4028:PMC
4020:doi
3979:PMC
3971:doi
3959:520
3922:PMC
3914:doi
3873:PMC
3865:doi
3824:PMC
3816:doi
3775:PMC
3765:doi
3753:110
3716:PMC
3708:doi
3704:153
3661:doi
3649:464
3616:PMC
3608:doi
3596:327
3559:PMC
3551:doi
3502:PMC
3494:doi
3490:346
3447:doi
3443:129
3410:doi
3406:349
3369:PMC
3361:doi
3320:PMC
3312:doi
3308:137
3253:PMC
3245:doi
3201:PMC
3193:doi
3152:PMC
3142:doi
3095:doi
3091:201
3058:PMC
3050:doi
3009:PMC
3001:doi
2989:471
2931:doi
2887:PMC
2879:doi
2838:PMC
2830:doi
2789:PMC
2781:doi
2740:PMC
2732:doi
2691:PMC
2683:doi
2642:PMC
2632:doi
2620:115
2583:doi
2579:172
2542:PMC
2534:doi
2530:171
2487:doi
2447:PMC
2439:doi
2389:doi
2348:PMC
2338:doi
2297:PMC
2289:doi
2248:PMC
2240:doi
2199:PMC
2191:doi
2179:457
2142:PMC
2134:doi
2122:459
2085:PMC
2077:doi
2073:138
2030:doi
2018:435
1985:PMC
1975:doi
1934:PMC
1926:doi
1885:PMC
1877:doi
1836:PMC
1828:doi
1787:PMC
1777:doi
1765:100
1730:doi
1718:232
1687:doi
1675:227
1642:PMC
1634:doi
1622:507
1577:doi
1536:PMC
1528:doi
1516:626
1476:PMC
1468:doi
1456:626
1418:doi
1381:PMC
1373:doi
1332:PMC
1322:doi
1304:Fly
1261:doi
1215:doi
1180:doi
1168:221
1135:PMC
1117:doi
1076:PMC
1068:doi
1019:doi
978:PMC
970:doi
916:doi
904:281
764:In
740:of
611:eve
607:Eve
603:eve
578:in
556:RET
521:of
390:or
314:YY1
312:or
143:In
112:cis
87:DNA
73:In
48:DNA
4692::
4636::
4624::
4371:):
4268:,
4172:.
4162:.
4152:53
4150:.
4146:.
4123:.
4113:54
4111:.
4099:^
4085:.
4075:.
4065:33
4063:.
4059:.
4036:.
4026:.
4016:26
4014:.
4010:.
3987:.
3977:.
3969:.
3957:.
3953:.
3930:.
3920:.
3908:.
3904:.
3881:.
3871:.
3861:25
3859:.
3855:.
3832:.
3822:.
3812:56
3810:.
3806:.
3783:.
3773:.
3763:.
3751:.
3747:.
3724:.
3714:.
3702:.
3698:.
3675:.
3667:.
3659:.
3647:.
3624:.
3614:.
3606:.
3594:.
3590:.
3567:.
3557:.
3549:.
3539:20
3537:.
3533:.
3510:.
3500:.
3488:.
3484:.
3461:.
3453:.
3441:.
3418:.
3404:.
3400:.
3377:.
3367:.
3357:13
3355:.
3351:.
3328:.
3318:.
3306:.
3302:.
3261:.
3251:.
3241:18
3239:.
3235:.
3223:^
3209:.
3199:.
3187:.
3183:.
3160:.
3150:.
3140:.
3136:.
3132:.
3109:.
3101:.
3089:.
3066:.
3056:.
3046:69
3044:.
3040:.
3017:.
3007:.
2999:.
2987:.
2983:.
2945:.
2937:.
2927:94
2925:.
2921:.
2895:.
2885:.
2875:48
2873:.
2869:.
2846:.
2836:.
2826:22
2824:.
2820:.
2797:.
2787:.
2777:40
2775:.
2771:.
2748:.
2738:.
2728:32
2726:.
2722:.
2699:.
2689:.
2679:16
2677:.
2673:.
2650:.
2640:.
2630:.
2618:.
2614:.
2591:.
2577:.
2573:.
2550:.
2540:.
2528:.
2524:.
2501:.
2493:.
2483:20
2481:.
2469:^
2455:.
2445:.
2435:23
2433:.
2429:.
2417:^
2403:.
2395:.
2385:13
2383:.
2356:.
2346:.
2332:.
2328:.
2305:.
2295:.
2285:19
2283:.
2279:.
2256:.
2246:.
2236:42
2234:.
2230:.
2207:.
2197:.
2189:.
2177:.
2173:.
2150:.
2140:.
2132:.
2120:.
2116:.
2093:.
2083:.
2071:.
2067:.
2044:.
2036:.
2028:.
2016:.
1993:.
1983:.
1969:.
1965:.
1942:.
1932:.
1922:22
1920:.
1916:.
1893:.
1883:.
1873:38
1871:.
1867:.
1844:.
1834:.
1824:30
1822:.
1818:.
1795:.
1785:.
1775:.
1763:.
1759:.
1736:.
1728:.
1716:.
1693:.
1685:.
1673:.
1650:.
1640:.
1632:.
1620:.
1616:.
1593:.
1585:.
1571:.
1567:.
1544:.
1534:.
1526:.
1514:.
1510:.
1498:^
1484:.
1474:.
1466:.
1454:.
1450:.
1438:^
1424:.
1414:54
1412:.
1389:.
1379:.
1367:.
1363:.
1340:.
1330:.
1320:.
1308:16
1306:.
1302:.
1275:.
1267:.
1257:33
1255:.
1243:^
1229:.
1221:.
1211:33
1209:.
1186:.
1178:.
1166:.
1143:.
1133:.
1125:.
1113:18
1111:.
1107:.
1084:.
1074:.
1064:32
1062:.
1058:.
1035:.
1027:.
1013:.
1009:.
986:.
976:.
966:14
964:.
960:.
944:^
930:.
922:.
914:.
902:.
842:.
566:.
410:.
291:,
214:.
83:bp
4658:)
4653:,
4649:(
4449::
4426::
4380:/
4367:/
4363:(
4272:)
4264:(
4254:e
4247:t
4240:v
4180:.
4158::
4131:.
4119::
4093:.
4071::
4044:.
4022::
3995:.
3973::
3965::
3938:.
3916::
3910:9
3889:.
3867::
3840:.
3818::
3791:.
3767::
3759::
3732:.
3710::
3683:.
3663::
3655::
3632:.
3610::
3602::
3575:.
3553::
3545::
3518:.
3496::
3469:.
3449::
3426:.
3412::
3385:.
3363::
3336:.
3314::
3269:.
3247::
3217:.
3195::
3189:4
3168:.
3144::
3138:7
3117:.
3097::
3074:.
3052::
3025:.
3003::
2995::
2968:.
2933::
2903:.
2881::
2854:.
2832::
2805:.
2783::
2756:.
2734::
2707:.
2685::
2658:.
2634::
2626::
2599:.
2585::
2558:.
2536::
2509:.
2489::
2463:.
2441::
2411:.
2391::
2364:.
2340::
2334:8
2313:.
2291::
2264:.
2242::
2215:.
2193::
2185::
2158:.
2136::
2128::
2101:.
2079::
2052:.
2032::
2024::
2001:.
1977::
1971:9
1950:.
1928::
1901:.
1879::
1852:.
1830::
1803:.
1779::
1771::
1744:.
1732::
1724::
1701:.
1689::
1681::
1658:.
1636::
1628::
1601:.
1579::
1573:7
1552:.
1530::
1522::
1492:.
1470::
1462::
1432:.
1420::
1397:.
1375::
1369:7
1348:.
1324::
1300:"
1283:.
1263::
1237:.
1217::
1194:.
1182::
1174::
1151:.
1119::
1092:.
1070::
1043:.
1021::
1015:7
994:.
972::
938:.
918::
910::
601:(
460:,
93:(
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.