252:. The term coenzyme refers specifically to enzymes and, as such, to the functional properties of a protein. On the other hand, "prosthetic group" emphasizes the nature of the binding of a cofactor to a protein (tight or covalent) and, thus, refers to a structural property. Different sources give slightly different definitions of coenzymes, cofactors, and prosthetic groups. Some consider tightly bound organic molecules as prosthetic groups and not as coenzymes, while others define all non-protein organic molecules needed for enzyme activity as coenzymes, and classify those that are tightly bound as coenzyme prosthetic groups. These terms are often used loosely.
31:
1642:
123:", which consists of a coenzyme that is tightly (or even covalently) and permanently bound to a protein. The second type of coenzymes are called "cosubstrates", and are transiently bound to the protein. Cosubstrates may be released from a protein at some point, and then rebind later. Both prosthetic groups and cosubstrates have the same function, which is to facilitate the reaction of enzymes and proteins. An inactive enzyme without the cofactor is called an
588:
1875:, methylation, or glycosylation in that the amino acids typically acquire new functions. This increases the functionality of the protein; unmodified amino acids are typically limited to acid-base reactions, and the alteration of resides can give the protein electrophilic sites or the ability to stabilize free radicals. Examples of cofactor production include
131:. (The International Union of Pure and Applied Chemistry (IUPAC) defines "coenzyme" a little differently, namely as a low-molecular-weight, non-protein organic compound that is loosely attached, participating in enzymatic reactions as a dissociable carrier of chemical groups or electrons; a prosthetic group is defined as a tightly bound,
648:, for instance, are tightly bound. Tightly bound cofactors are, in general, regenerated during the same reaction cycle, while loosely bound cofactors can be regenerated in a subsequent reaction catalyzed by a different enzyme. In the latter case, the cofactor can also be considered a substrate or cosubstrate.
1907:
are termed cofactors or coactivators, whereas molecules that inhibit receptor proteins are termed corepressors. One such example is the G protein-coupled receptor family of receptors, which are frequently found in sensory neurons. Ligand binding to the receptors activates the G protein, which then
1870:
In a number of enzymes, the moiety that acts as a cofactor is formed by post-translational modification of a part of the protein sequence. This often replaces the need for an external binding factor, such as a metal ion, for protein function. Potential modifications could be oxidation of aromatic
615:
Organic cofactors are small organic molecules (typically a molecular mass less than 1000 Da) that can be either loosely or tightly bound to the enzyme and directly participate in the reaction. In the latter case, when it is difficult to remove without denaturing the enzyme, it can be called a
1758:
evolving to bind a restricted set of nucleotides and related compounds. Adenosine-based cofactors are thought to have acted as interchangeable adaptors that allowed enzymes and ribozymes to bind new cofactors through small modifications in existing adenosine-binding
1785:. If enzymes require a co-enzyme, how does the coenzyme evolve? The most likely scenario is that enzymes can function initially without their coenzymes and later recruit the coenzyme, even if the catalyzed reaction may not be as efficient or as fast. Examples are
259:
noted the confusion in the literature and the essentially arbitrary distinction made between prosthetic groups and coenzymes group and proposed the following scheme. Here, cofactors were defined as an additional substance apart from protein and
1858:, who established the link between the oxidation of sugars and the generation of ATP. This confirmed the central role of ATP in energy transfer that had been proposed by Fritz Albert Lipmann in 1941. Later, in 1949, Morris Friedkin and
1700:
of 100 to 150 moles of ATP daily, which is around 50 to 75 kg. In typical situations, humans use up their body weight of ATP over the course of the day. This means that each ATP molecule is recycled 1000 to 1500 times daily.
1661:. This common chemistry allows cells to use a small set of metabolic intermediates to carry chemical groups between different reactions. These group-transfer intermediates are the loosely bound organic cofactors, often called
606:
Iron–sulfur clusters are complexes of iron and sulfur atoms held within proteins by cysteinyl residues. They play both structural and functional roles, including electron transfer, redox sensing, and as structural modules.
1696:. This ATP is constantly being broken down into ADP, and then converted back into ATP. Thus, at any given time, the total amount of ATP + ADP remains fairly constant. The energy used by human cells requires the
1908:
activates an enzyme to activate the effector. In order to avoid confusion, it has been suggested that such proteins that have ligand-binding mediated activation or repression be referred to as coregulators.
1668:
Each class of group-transfer reaction is carried out by a particular cofactor, which is the substrate for a set of enzymes that produce it, and a set of enzymes that consume it. An example of this are the
714:
groups. Most of these cofactors are found in a huge variety of species, and some are universal to all forms of life. An exception to this wide distribution is a group of unique cofactors that evolved in
268:
attached to a single enzyme molecule. However, the author could not arrive at a single all-encompassing definition of a "coenzyme" and proposed that this term be dropped from use in the literature.
1895:
The term is used in other areas of biology to refer more broadly to non-protein (or even protein) molecules that either activate, inhibit, or are required for the protein to function. For example,
208:. It has been suggested that the AMP part of the molecule can be considered to be a kind of "handle" by which the enzyme can "grasp" the coenzyme to switch it between different catalytic centers.
4250:"The Ninth Sir Hans Krebs Lecture. Compartmentation and communication in living systems. Ligand conduction: a general catalytic principle in chemical, osmotic and chemiosmotic reaction systems"
1879:(TTQ), derived from two tryptophan side chains, and 4-methylidene-imidazole-5-one (MIO), derived from an Ala-Ser-Gly motif. Characterization of protein-derived cofactors is conducted using
620:. There is no sharp division between loosely and tightly bound cofactors. Many such as NAD can be tightly bound in some enzymes, while it is loosely bound in others. Another example is
1731:
indicates that these molecules evolved very early in the development of living things. At least some of the current set of cofactors may, therefore, have been present in the
5210:
Lodish, Harvey; Berk, Arnold; Zipursky, S. Lawrence; Matsudaira, Paul; Baltimore, David; Darnell, James (2000-01-01). "G Protein–Coupled
Receptors and Their Effectors".
1957:
1843:. Other cofactors were identified throughout the early 20th century, with ATP being isolated in 1929 by Karl Lohmann, and coenzyme A being discovered in 1945 by
2537:
Chan MK, Mukund S, Kletzin A, Adams MW, Rees DC (March 1995). "Structure of a hyperthermophilic tungstopterin enzyme, aldehyde ferredoxin oxidoreductase".
3599:"Structure of component B (7-mercaptoheptanoylthreonine phosphate) of the methylcoenzyme M methylreductase system of Methanobacterium thermoautotrophicum"
1871:
residues, binding between residues, cleavage or ring-forming. These alterations are distinct from other post-translation protein modifications, such as
2178:
Denessiouk KA, Rantanen VV, Johnson MS (August 2001). "Adenine recognition: a motif present in ATP-, CoA-, NAD-, NADP-, and FAD-dependent proteins".
353:
is another special case, in that it is required as a component of the human diet, and it is needed for the full activity of many enzymes, such as
2922:
Hanukoglu I (December 2017). "Conservation of the Enzyme–Coenzyme
Interfaces in FAD and NADP Binding Adrenodoxin Reductase-A Ubiquitous Enzyme".
1990:
2150:
1763:, which had originally evolved to bind a different cofactor. This process of adapting a pre-evolved structure for a novel use is known as
5857:
5720:
5716:
1802:
1547:
793:
1657:
Metabolism involves a vast array of chemical reactions, but most fall under a few basic types of reactions that involve the transfer of
5649:
5311:
2015:
1746:
is present in cofactors that catalyse many basic metabolic reactions such as methyl, acyl, and phosphoryl group transfer, as well as
5153:"A new member of the 4-methylideneimidazole-5-one-containing aminomutase family from the enediyne kedarcidin biosynthetic pathway"
4918:
Warburg O, Christian W (1936). "Pyridin, the hydrogen-transferring component of the fermentation enzymes (pyridine nucleotide)".
116:
in small amounts. (Some scientists limit the use of the term "cofactor" for inorganic substances; both types are included here.)
4111:
Salisbury SA, Forrest HS, Cruse WB, Kennard O (August 1979). "A novel coenzyme from bacterial primary alcohol dehydrogenases".
5292:
4756:
4719:
3837:
3810:
3532:
2865:
2840:
2811:
2134:
2089:
5023:"Esterification of inorganic phosphate coupled to electron transport between dihydrodiphosphopyridine nucleotide and oxygen"
3187:
3747:
4818:
2475:
Niki I, Yokokura H, Sudo T, Kato M, Hidaka H (October 1996). "Ca2+ signaling and intracellular Ca2+ binding proteins".
2049:
403:
1775:. A computational method, IPRO, recently predicted mutations that experimentally switched the cofactor specificity of
5712:
5708:
5358:
5005:
4694:
4409:
3238:
Jitrapakdee S, Wallace JC (2003). "The biotin enzyme family: conserved structural motifs and domain rearrangements".
3150:
Eliot AC, Kirsch JF (2004). "Pyridoxal phosphate enzymes: mechanistic, structural, and evolutionary considerations".
1790:
1674:
1650:
789:
699:
197:
163:
1854:
identified the function of NAD in hydride transfer. This discovery was followed in the early 1940s by the work of
435:
In many cases, the cofactor includes both an inorganic and organic component. One diverse set of examples is the
591:
A simple cluster containing two iron atoms and two sulfur atoms, coordinated by four protein cysteine residues.
5776:
5642:
66:
5514:
5106:
4291:
4213:
3855:
1835:. Through a long and difficult purification from yeast extracts, this heat-stable factor was identified as a
1677:(NAD) as a cofactor. Here, hundreds of separate types of enzymes remove electrons from their substrates and
5065:
Davidson VL (2007). "Protein-Derived
Cofactors. Expanding the Scope of Post-Translational Modifications†".
1876:
5698:
1932:
1176:
703:
637:
193:
159:
30:
3353:"Microbial ubiquinones: multiple roles in respiration, gene regulation and oxidative stress management"
338:
5635:
5615:
5602:
5589:
5576:
5563:
5550:
5537:
5499:
2248:
428:
3930:"The active species of 'CO2' utilized by formylmethanofuran dehydrogenase from methanogenic Archaea"
1994:
264:
that is required for enzyme activity and a prosthetic group as a substance that undergoes its whole
5929:
5509:
5463:
5406:
5325:
4072:
Negishi M, Pedersen LG, Petrotchenko E, Shevtsov S, Gorokhov A, Kakuta Y, Pedersen LC (June 2001).
1927:
1572:
17:
1862:
proved that NAD linked metabolic pathways such as the citric acid cycle and the synthesis of ATP.
1750:
reactions. This ubiquitous chemical scaffold has, therefore, been proposed to be a remnant of the
5942:
4736:
1904:
1814:
1732:
1728:
1618:
601:
236:
181:
138:
Some enzymes or enzyme complexes require several cofactors. For example, the multienzyme complex
43:
35:
694:). However, vitamins do have other functions in the body. Many organic cofactors also contain a
5842:
5684:
3524:
3518:
1887:; structural data is necessary because sequencing does not readily identify the altered sites.
1840:
1828:
1716:
1268:
751:
633:
629:
621:
294:
185:
151:
139:
2272:
Aggett PJ (August 1985). "Physiology and metabolism of essential trace elements: an outline".
1831:
in unboiled yeast extracts. They called the unidentified factor responsible for this effect a
85:
transformations. The rates at which these happen are characterized in an area of study called
5852:
5847:
5694:
5432:
5351:
5102:"Posttranslational biosynthesis of the protein-derived cofactor tryptophan tryptophylquinone"
3827:
3208:
3164:
1896:
1880:
1851:
1798:
1794:
1786:
1739:
1377:
1293:
1138:
571:
507:
366:
354:
90:
5321:
4629:"Computational design of Candida boidinii xylose reductase for altered cofactor specificity"
2755:
Meyer J (February 2008). "Iron-sulfur protein folds, iron-sulfur chemistry, and evolution".
5760:
5504:
5164:
4851:
4499:
4446:
4120:
3610:
2931:
2660:
2601:
2546:
2106:
1937:
1844:
1751:
1022:
205:
5308:
5119:
3548:
Chiang PK, Gordon RK, Tal J, Zeng GC, Doctor BP, Pardhasaradhi K, McCann PP (March 1996).
2488:
2019:
8:
5934:
5740:
5468:
3196:
3152:
3103:"The power to reduce: pyridine nucleotides—small molecules with a multitude of functions"
2391:
Cavalieri RR (April 1997). "Iodine metabolism and thyroid physiology: current concepts".
1922:
1859:
1589:
831:
411:
345:
is also an essential trace element, but this element is used as part of the structure of
5331:
5168:
4855:
4503:
4450:
4349:
Chen X, Li N, Ellington AD (2007). "Ribozyme catalysis of metabolism in the RNA world".
4304:
4226:
4124:
3868:
3614:
3474:
Mack M, Grill S (2006). "Riboflavin analogs and inhibitors of riboflavin biosynthesis".
3352:
3067:
3042:
2935:
2664:
2605:
2550:
6054:
5956:
5401:
5281:
5253:
5228:
5187:
5152:
5128:
5101:
4997:
4970:
4867:
4653:
4628:
4523:
4472:
4374:
4327:
4266:
4249:
4188:
4163:
4144:
3775:
3699:"Specificity and biological distribution of coenzyme M (2-mercaptoethanesulfonic acid)"
3579:
3499:
3333:
3220:
3127:
3102:
3080:
2955:
2800:
2780:
2732:
2705:
2686:
2570:
2457:
2332:
2203:
2078:
575:
466:
420:
113:
5039:
4903:
4886:
4748:
4605:
4569:
4542:
4401:
4289:
Wimmer MJ, Rose IA (1978). "Mechanisms of enzyme-catalyzed group transfer reactions".
3723:
3698:
3674:
3657:
3633:
3598:
3018:
3001:
2899:
2285:
6027:
5786:
5768:
5288:
5258:
5192:
5133:
5082:
5044:
5001:
4962:
4762:
4752:
4715:
4690:
4658:
4609:
4574:
4515:
4464:
4459:
4434:
4415:
4405:
4366:
4308:
4271:
4230:
4193:
4136:
4093:
4054:
4027:
3992:
3951:
3946:
3929:
3907:
3872:
3833:
3806:
3799:
3767:
3728:
3679:
3638:
3571:
3528:
3491:
3456:
3418:
3413:
3396:
3377:
3337:
3325:
3290:
3255:
3212:
3168:
3132:
3072:
3023:
2982:
2947:
2904:
2861:
2836:
2829:
2807:
2772:
2737:
2678:
2629:
2624:
2589:
2562:
2519:
2492:
2449:
2408:
2373:
2324:
2289:
2234:
2195:
2130:
2085:
2045:
1976:
1884:
798:
683:
539:
389:
147:
78:
62:
4974:
4871:
4627:
Khoury GA, Fazelinia H, Chin JW, Pantazes RJ, Cirino PC, Maranas CD (October 2009).
4527:
4476:
4378:
4328:"Estimating ATP resynthesis during a marathon run: a method to introduce metabolism"
3779:
3583:
3503:
3224:
3084:
2784:
2690:
2574:
2461:
2336:
2207:
6049:
5447:
5442:
5416:
5344:
5248:
5240:
5182:
5172:
5123:
5115:
5074:
5034:
4993:
4988:
Lipmann F (1941). "Metabolic generation and utilization of phosphate bond energy".
4954:
4927:
4898:
4859:
4798:
4744:
4682:
4648:
4640:
4601:
4564:
4554:
4507:
4454:
4397:
4358:
4300:
4261:
4222:
4183:
4175:
4148:
4128:
4085:
4019:
3982:
3941:
3899:
3890:
Wijayanti N, Katz N, Immenschuh S (2004). "Biology of heme in health and disease".
3864:
3763:
3759:
3718:
3710:
3669:
3628:
3618:
3561:
3483:
3448:
3408:
3367:
3317:
3282:
3247:
3204:
3160:
3122:
3114:
3062:
3054:
3013:
2959:
2939:
2894:
2764:
2727:
2717:
2668:
2619:
2609:
2554:
2484:
2439:
2400:
2363:
2316:
2281:
2230:
2187:
1972:
1917:
1658:
1522:
985:
617:
547:
362:
261:
248:
120:
101:
3714:
3286:
5790:
5494:
5478:
5391:
5315:
4676:
4023:
3273:
Leonardi R, Zhang YM, Rock CO, Jackowski S (2005). "Coenzyme A: back in action".
1872:
1051:
867:
346:
265:
86:
4945:
Kalckar HM (November 1974). "Origins of the concept oxidative phosphorylation".
4179:
3783:
3566:
3549:
3372:
5806:
5532:
5473:
5211:
5157:
Proceedings of the
National Academy of Sciences of the United States of America
3603:
Proceedings of the
National Academy of Sciences of the United States of America
3452:
2594:
Proceedings of the
National Academy of Sciences of the United States of America
2444:
2427:
1855:
1760:
1692:. As an example, the total quantity of ATP in the human body is about 0.1
1527:
1382:
579:
515:
461:
374:
282:
132:
97:
4931:
4819:"Fermentation of sugars and fermentative enzymes: Nobel Lecture, May 23, 1930"
4211:
DiMarco AA, Bobik TA, Wolfe RS (1990). "Unusual coenzymes of methanogenesis".
3487:
3058:
2943:
2768:
2706:"Structural analysis of heme proteins: implications for design and prediction"
6043:
5919:
5872:
5824:
5437:
5396:
4490:
White HB (March 1976). "Coenzymes as fossils of an earlier metabolic state".
3925:
3903:
3251:
1820:
1819:
The first organic cofactor to be discovered was NAD, which was identified by
1670:
1496:
1214:
1107:
1037:
766:
625:
377:
molecule, and not usually considered a cofactor of the enzymes it regulates.
302:
5177:
3623:
2722:
2558:
2320:
200:. This common structure may reflect a common evolutionary origin as part of
5924:
5914:
5882:
5386:
5262:
5196:
5137:
5086:
5048:
5022:
4803:
4786:
4662:
4613:
4578:
4559:
4370:
4362:
4197:
4097:
4089:
4045:
Ginsburg V (1978). "Comparative biochemistry of nucleotide-linked sugars".
4031:
3996:
3911:
3771:
3495:
3460:
3422:
3381:
3329:
3321:
3294:
3259:
3216:
3172:
3136:
3076:
2986:
2951:
2883:"Studies on the nature of the binding of thiamine pyrophosphate to enzymes"
2776:
2741:
2682:
2633:
2614:
2510:
Eady RR (July 1988). "The vanadium-containing nitrogenase of
Azotobacter".
2453:
2404:
2377:
2368:
2351:
2328:
2199:
1623:
1479:
1340:
1298:
293:
are common cofactors. The study of these cofactors falls under the area of
224:
82:
39:
4966:
4766:
4519:
4468:
4419:
4234:
3955:
3876:
3683:
3642:
3575:
3043:"Structure, mechanism and catalytic duality of thiamine-dependent enzymes"
3027:
2908:
2566:
2523:
2496:
2412:
2293:
1641:
341:, no human enzyme that uses this metal as a cofactor has been identified.
5909:
5862:
5610:
5545:
5381:
5244:
4312:
4275:
4140:
4058:
3987:
3970:
3732:
2882:
1850:
The functions of these molecules were at first mysterious, but, in 1936,
1710:
1693:
1662:
1403:
872:
676:
543:
495:
491:
394:
385:
358:
105:
2647:
Lane TW, Saito MA, George GN, Pickering IJ, Prince RC, Morel FM (2005).
1958:"Coenzyme, Cofactor and Prosthetic Group — Ambiguous Biochemical Jargon"
305:
reflects their role as cofactors. In humans this list commonly includes
5989:
5877:
5867:
5730:
4958:
4863:
4543:"The tyranny of adenosine recognition among RNA aptamers to coenzyme A"
4511:
3118:
1836:
1765:
1724:
1697:
1689:
1628:
1489:
1352:
1345:
1335:
1328:
1318:
1237:
1219:
1181:
1143:
1027:
980:
908:
903:
836:
716:
707:
695:
687:
669:
662:
655:
534:
511:
482:
380:
Other organisms require additional metals as enzyme cofactors, such as
370:
330:
189:
178:
167:
143:
128:
5078:
2191:
1681:
NAD to NADH. This reduced cofactor is then a substrate for any of the
5984:
5979:
5814:
5800:
5584:
5558:
4132:
1823:
and
William Young 1906. They noticed that adding boiled and filtered
1743:
1682:
1611:
1565:
1540:
1515:
1472:
1455:
1450:
1443:
1418:
1396:
1370:
1311:
1286:
1273:
1207:
1169:
1131:
1100:
1072:
1065:
1015:
973:
938:
896:
860:
824:
782:
691:
645:
522:
502:
440:
314:
310:
298:
155:
124:
119:
Coenzymes are further divided into two types. The first is called a "
74:
4842:
4686:
4644:
4073:
2673:
2648:
5627:
4396:. Advances in Microbial Physiology. Vol. 40. pp. 353–99.
4071:
2044:(Fifth ed.). New York: W.H. Freeman and Company. p. 184.
1900:
1755:
1603:
1598:
1582:
1577:
1557:
1532:
1507:
1484:
1464:
1435:
1430:
1414:
1408:
1388:
1362:
1357:
1323:
1303:
1278:
1241:
1232:
1199:
1194:
1161:
1156:
1123:
1118:
1092:
1087:
1083:
1057:
1047:
1007:
965:
930:
921:
888:
852:
816:
811:
774:
756:
527:
478:
399:
381:
334:
201:
135:
unit in a protein that is regenerated in each enzymatic turnover.)
3829:
Significance of glutathione in plant adaptation to the environment
654:
can serve as precursors to many organic cofactors (e.g., vitamins
6020:
5964:
5676:
2249:"Biochemistry: Enzymes: Classification and catalysis (Cofactors)"
1723:, are present in all known forms of life and form a core part of
1607:
1561:
1552:
1536:
1511:
1468:
1439:
1392:
1366:
1307:
1282:
1203:
1165:
1127:
1096:
1061:
1011:
969:
934:
892:
856:
820:
778:
720:
651:
416:
407:
350:
174:
150:
requires five organic cofactors and one metal ion: loosely bound
109:
81:). Cofactors can be considered "helper molecules" that assist in
59:
4887:"Acetylation of sulfanilamide by liver homogenates and extracts"
1688:
Therefore, these cofactors are continuously recycled as part of
5994:
5969:
5903:
5899:
5895:
5891:
5750:
5658:
5597:
5367:
3397:"Vitamin C. Biosynthesis, recycling and degradation in mammals"
1685:
in the cell that require electrons to reduce their substrates.
1594:
1501:
1043:
998:
950:
945:
641:
559:
554:
424:
342:
322:
318:
70:
4164:"Tetrahydrobiopterin biosynthesis, regeneration and functions"
5571:
2221:
Bryce (March 1979). "SAM – semantics and misunderstandings".
1824:
1747:
1678:
1646:
925:
287:
93:
in that they often derive their function by remaining bound.
5209:
4714:(4th ed.). Belmont, CA: Brooks/Cole, Cengage Learning.
4110:
1738:
Organic cofactors may have been present even earlier in the
5999:
5974:
5887:
4626:
4010:
Mendel RR, Bittner F (2006). "Cell biology of molybdenum".
2307:
Stearns DM (2000). "Is chromium a trace essential metal?".
2127:
Thiamine: catalytic mechanisms in normal and disease states
1720:
1459:
1425:
1002:
883:
711:
566:
486:
473:
444:
436:
326:
306:
228:
47:
5336:
3272:
2177:
290:
4435:"The emergence of major cellular processes in evolution"
2646:
587:
77:(a catalyst is a substance that increases the rate of a
4791:
Proceedings of the Royal
Society B: Biological Sciences
4592:
Jadhav VR, Yarus M (2002). "Coenzymes as coribozymes".
3889:
3597:
Noll KM, Rinehart KL, Tanner RS, Wolfe RS (June 1986).
3596:
1636:
3805:(1st ed.). American society of plant physiology.
3658:"Structure and methylation of coenzyme M(HSCH2CH2SO3)"
3394:
2973:
Bolander FF (2006). "Vitamins: not just for enzymes".
2536:
127:, while the complete enzyme with cofactor is called a
5060:
5058:
3796:
2649:"Biochemistry: a cadmium enzyme from a marine diatom"
2590:"A biological function for cadmium in marine diatoms"
242:
Organic cofactors are sometimes further divided into
5150:
2474:
2084:(2nd ed.). San Diego: Harcourt/Academic Press.
2080:
Biochemistry: the chemical reactions of living cells
5226:
3547:
2075:
369:, often binding to these enzymes in a complex with
5280:
5229:"Coactivators and corepressors: what's in a name?"
5055:
3971:"Molybdoenzymes and molybdenum cofactor in plants"
3798:
3237:
3100:
2915:
2828:
2826:
2799:
2077:
5151:Huang SX, Lohman JR, Huang T, Shen B (May 2013).
5020:
4734:
4210:
4161:
3790:
3439:Joosten V, van Berkel WJ (2007). "Flavoenzymes".
3438:
6041:
5283:An introduction to enzyme and coenzyme chemistry
4917:
4432:
3825:
3520:An introduction to enzyme and coenzyme chemistry
3310:Critical Reviews in Clinical Laboratory Sciences
3185:
3040:
2108:GLOSSARY OF TERMS USED IN BIOINORGANIC CHEMISTRY
216:Cofactors can be divided into two major groups:
4540:
4348:
4325:
3852:
2855:
5099:
4709:
3853:Meister A, Anderson ME (1983). "Glutathione".
3801:Biochemistry & molecular biology of plants
2703:
5643:
5352:
4992:. Adv Enzymol. Vol. 1. pp. 99–162.
4743:, vol. 113, Elsevier, pp. 484–490,
4074:"Structure and function of sulfotransferases"
4009:
3924:
2999:
2129:. New York, N.Y: Marcel Dekker. p. 588.
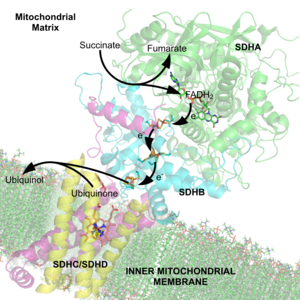
38:complex showing several cofactors, including
27:Non-protein chemical compound or metallic ion
4047:Progress in Clinical and Biological Research
3968:
3752:Journal of the American College of Nutrition
3350:
2860:(6th ed.). Pacific Grove: Brooks Cole.
2835:(3rd ed.). New York: Worth Publishers.
2173:
2171:
1865:
96:Cofactors can be classified into two types:
4784:
4591:
3655:
3192:: catalysis by cobalamin-dependent enzymes"
3149:
2124:
726:
184:(AMP) as part of their structures, such as
5650:
5636:
5359:
5345:
4735:Carlberg, Inger; Mannervik, Bengt (1985),
4288:
4162:Thöny B, Auerbach G, Blau N (April 2000).
3797:Buchanan BB, Gruissem W, Jones RL (2000).
3696:
3434:
3432:
2104:
1955:
1890:
5324:at the U.S. National Library of Medicine
5252:
5186:
5176:
5127:
5038:
4987:
4902:
4884:
4841:
4802:
4710:Garrett, R.; Grisham, Charles M. (2010).
4652:
4568:
4558:
4458:
4265:
4187:
3986:
3945:
3722:
3673:
3632:
3622:
3565:
3473:
3412:
3371:
3126:
3096:
3094:
3066:
3017:
2921:
2898:
2880:
2731:
2721:
2704:Li T, Bonkovsky HL, Guo JT (March 2011).
2672:
2623:
2613:
2587:
2443:
2390:
2367:
2168:
2076:Sauke DJ, Metzler DE, Metzler CM (2001).
2071:
2069:
2067:
2065:
2063:
2061:
2039:
1735:, which lived about 4 billion years ago.
365:, but calcium activates these enzymes in
5227:O'Malley BW, McKenna NJ (October 2008).
5064:
4319:
4247:
4044:
4038:
3307:
3209:10.1146/annurev.biochem.72.121801.161828
3165:10.1146/annurev.biochem.73.011303.074021
2972:
1956:Hasim, Onn H.; Adnan, Nor Azila (2010).
1640:
719:, which are restricted to this group of
586:
456:Examples of enzymes containing this ion
235:, such as the metal ions Mg, Cu, Mn and
177:or made from vitamins. Many contain the
29:
4944:
4078:Archives of Biochemistry and Biophysics
3748:"Biochemical functions of coenzyme Q10"
3429:
3344:
3308:Donnelly JG (June 2001). "Folic acid".
2827:Cox M, Lehninger AL, Nelson DR (2000).
2425:
2349:
2306:
2274:Clinics in Endocrinology and Metabolism
595:
14:
6042:
4787:"The Alcoholic Ferment of Yeast-Juice"
4785:Harden A, Young WJ (24 October 1906).
3550:"S-Adenosylmethionine and methylation"
3523:. Oxford: Blackwell Science. pp.
3395:Linster CL, Van Schaftingen E (2007).
3091:
3041:Frank RA, Leeper FJ, Luisi BF (2007).
3002:"Novel biochemistry of methanogenesis"
2797:
2271:
2058:
1783:Evolution of enzymes without coenzymes
271:
5631:
5340:
5120:10.1146/annurev-biochem-051110-133601
4990:A Source Book in Chemistry, 1900-1950
4489:
3745:
3101:Pollak N, Dölle C, Ziegler M (2007).
2754:
2489:10.1093/oxfordjournals.jbchem.a021466
2220:
1779:xylose reductase from NADPH to NADH.
1548:3'-Phosphoadenosine-5'-phosphosulfate
5657:
5278:
4737:"[59] Glutathione reductase"
4433:Ouzounis C, Kyrpides N (July 1996).
4391:
3826:Grill D, Tausz T, De Kok LJ (2001).
3516:
2831:Lehninger principles of biochemistry
2509:
2042:Lehninger Principles of Biochemistry
1637:Cofactors as metabolic intermediates
690:) or as coenzymes themselves (e.g.,
632:, while it is less tightly bound in
108:. Coenzymes are mostly derived from
4947:Molecular and Cellular Biochemistry
4541:Saran D, Frank J, Burke DH (2003).
4305:10.1146/annurev.bi.47.070178.005123
4227:10.1146/annurev.bi.59.070190.002035
3969:Mendel RR, Hänsch R (August 2002).
3869:10.1146/annurev.bi.52.070183.003431
3697:Balch WE, Wolfe RS (January 1979).
3662:The Journal of Biological Chemistry
3656:Taylor CD, Wolfe RS (August 1974).
3351:Søballe B, Poole RK (August 1999).
3006:The Journal of Biological Chemistry
3000:Rouvière PE, Wolfe RS (June 1988).
2887:The Journal of Biological Chemistry
349:rather than as an enzyme cofactor.
24:
5272:
4998:10.4159/harvard.9780674366701.c141
4267:10.1111/j.1432-1033.1979.tb12934.x
404:aldehyde ferredoxin oxidoreductase
89:. Cofactors typically differ from
25:
6066:
5834:
5302:
5021:Friedkin M, Lehninger AL (1949).
1675:nicotinamide adenine dinucleotide
1651:nicotinamide adenine dinucleotide
624:(TPP), which is tightly bound in
211:
164:nicotinamide adenine dinucleotide
4326:Di Carlo SE, Collins HL (2001).
4254:European Journal of Biochemistry
3947:10.1111/j.1432-1033.1997.00919.x
3934:European Journal of Biochemistry
3414:10.1111/j.1742-4658.2006.05607.x
3186:Banerjee R, Ragsdale SW (2003).
2856:Farrell SO, Campbell MK (2009).
2588:Lane TW, Morel FM (April 2000).
2151:"Pyruvate Dehydrogenase Complex"
698:, such as the electron carriers
5220:
5203:
5144:
5100:Davidson VL, Wilmot CM (2013).
5093:
5014:
4981:
4938:
4911:
4878:
4835:
4811:
4778:
4728:
4703:
4669:
4620:
4585:
4534:
4483:
4426:
4385:
4342:
4282:
4241:
4204:
4155:
4104:
4065:
4003:
3962:
3918:
3883:
3846:
3819:
3739:
3690:
3649:
3590:
3541:
3510:
3467:
3388:
3301:
3266:
3231:
3179:
3143:
3034:
2993:
2966:
2874:
2849:
2820:
2791:
2748:
2697:
2640:
2581:
2530:
2503:
2468:
2419:
2384:
2343:
2300:
2265:
2241:
2214:
1249:
257:Trends in Biochemistry Sciences
4885:Lipmann F (1 September 1945).
4492:Journal of Molecular Evolution
3975:Journal of Experimental Botany
3928:, Thauer RK (September 1997).
3764:10.1080/07315724.2001.10719063
2924:Journal of Molecular Evolution
2881:Morey AV, Juni E (June 1968).
2352:"The biochemistry of chromium"
2143:
2118:
2098:
2033:
2008:
1983:
1949:
13:
1:
5287:. Oxford: Blackwell Science.
5107:Annual Review of Biochemistry
5040:10.1016/S0021-9258(18)56879-4
4904:10.1016/S0021-9258(18)43110-9
4749:10.1016/s0076-6879(85)13062-4
4678:Comprehensive Enzyme Kinetics
4606:10.1016/S0300-9084(02)01404-9
4402:10.1016/S0065-2911(08)60135-6
4292:Annual Review of Biochemistry
4214:Annual Review of Biochemistry
3856:Annual Review of Biochemistry
3715:10.1128/JB.137.1.256-263.1979
3675:10.1016/S0021-9258(19)42403-4
3287:10.1016/j.plipres.2005.04.001
3019:10.1016/S0021-9258(18)68417-0
2900:10.1016/S0021-9258(18)93372-7
2286:10.1016/S0300-595X(85)80005-0
2105:de Bolster, M. W. G. (1997).
1943:
1260:Chemical group(s) transferred
743:Chemical group(s) transferred
439:proteins, which consist of a
276:
170:(CoA), and a metal ion (Mg).
4460:10.1016/0014-5793(96)00631-X
4394:How did bacteria come to be?
4351:Chemistry & Biodiversity
4024:10.1016/j.bbamcr.2006.03.013
3188:"The many faces of vitamin B
2235:10.1016/0968-0004(79)90255-X
1977:10.1016/0307-4412(94)90088-4
1877:tryptophan tryptophylquinone
1827:extract greatly accelerated
1704:
173:Organic cofactors are often
7:
5366:
3567:10.1096/fasebj.10.4.8647346
3476:Appl. Microbiol. Biotechnol
3373:10.1099/13500872-145-8-1817
2125:Jordan F, Patel MS (2004).
1933:Bioorganometallic chemistry
1911:
1715:Organic cofactors, such as
1177:Flavin adenine dinucleotide
771:2-carbon groups, α cleavage
638:flavin adenine dinucleotide
373:. Calcium is, therefore, a
160:flavin adenine dinucleotide
10:
6071:
3746:Crane FL (December 2001).
3453:10.1016/j.cbpa.2007.01.010
2445:10.1016/j.cell.2007.11.028
2040:Nelson DL, Cox MM (2008).
1903:that bind to and activate
1812:
1808:
1708:
610:
599:
339:impaired glucose tolerance
280:
6012:
5955:
5833:
5675:
5668:
5523:
5515:Michaelis–Menten kinetics
5487:
5456:
5425:
5374:
4932:10.1002/hlca.193601901199
4248:Mitchell P (March 1979).
4180:10.1042/0264-6021:3470001
3488:10.1007/s00253-006-0421-7
3059:10.1007/s00018-007-6423-5
2944:10.1007/s00239-017-9821-9
2769:10.1007/s00775-007-0318-7
2350:Vincent JB (April 2000).
1991:"coenzymes and cofactors"
1866:Protein-derived cofactors
1742:on Earth. The nucleotide
849:Amino and carboxyl groups
429:Thalassiosira weissflogii
5407:Diffusion-limited enzyme
5326:Medical Subject Headings
4920:Biochemische Zeitschrift
3904:10.2174/0929867043455521
3252:10.2174/1389203033487199
2975:Curr Opin Investig Drugs
2356:The Journal of Nutrition
2114:. Pure & Appl. Chem.
1928:Organometallic chemistry
1573:Pyrroloquinoline quinone
727:Vitamins and derivatives
301:, the list of essential
154:(TPP), covalently bound
69:that is required for an
5233:Molecular Endocrinology
5178:10.1073/pnas.1304733110
4168:The Biochemical Journal
3703:Journal of Bacteriology
3624:10.1073/pnas.83.12.4238
3240:Curr. Protein Pept. Sci
2723:10.1186/1472-6807-11-13
2559:10.1126/science.7878465
2477:Journal of Biochemistry
2321:10.1002/biof.5520110301
1891:Non-enzymatic cofactors
1815:History of biochemistry
1733:last universal ancestor
1619:Tetrahydromethanopterin
182:adenosine monophosphate
36:succinate dehydrogenase
5213:Molecular Cell Biology
4804:10.1098/rspb.1906.0070
4560:10.1186/1471-2148-3-26
4363:10.1002/cbdv.200790055
4090:10.1006/abbi.2001.2368
4012:Biochim. Biophys. Acta
3322:10.1080/20014091084209
2710:BMC Structural Biology
2615:10.1073/pnas.090091397
2405:10.1089/thy.1997.7.177
1841:Hans von Euler-Chelpin
1829:alcoholic fermentation
1654:
1269:Adenosine triphosphate
752:Thiamine pyrophosphate
634:pyruvate dehydrogenase
630:pyruvate decarboxylase
622:thiamine pyrophosphate
592:
392:bacteria of the genus
295:bioinorganic chemistry
152:thiamine pyrophosphate
140:pyruvate dehydrogenase
51:
5500:Eadie–Hofstee diagram
5433:Allosteric regulation
5332:The CoFactor Database
4741:Methods in Enzymology
2806:. New York: Horwood.
2802:Understanding enzymes
1965:Biochemical Education
1881:X-ray crystallography
1852:Otto Heinrich Warburg
1813:Further information:
1799:Glutathione Reductase
1795:Lactate Dehydrogenase
1787:Alcohol Dehydrogenase
1709:Further information:
1644:
1385:and lipid head groups
1378:Cytidine triphosphate
1294:S-Adenosyl methionine
1139:Flavin mononucleotide
600:Further information:
590:
572:Alcohol dehydrogenase
508:Glucose 6-phosphatase
367:allosteric regulation
355:nitric oxide synthase
281:Further information:
33:
5510:Lineweaver–Burk plot
5245:10.1210/me.2008-0201
3786:on 16 December 2008.
2757:J. Biol. Inorg. Chem
2369:10.1093/jn/130.4.715
2155:Chemistry LibreTexts
1938:Cofactor engineering
1845:Fritz Albert Lipmann
1773:Changes in coenzymes
1054:and formimino groups
1023:Tetrahydrofolic acid
740:Additional component
596:Iron–sulfur clusters
443:ring coordinated to
406:of the thermophilic
359:protein phosphatases
237:iron–sulfur clusters
162:(FAD), cosubstrates
5169:2013PNAS..110.8069H
4856:1929NW.....17..624.
4844:Naturwissenschaften
4504:1976JMolE...7..101W
4451:1996FEBSL.390..119O
4332:Advan. Physiol. Edu
4125:1979Natur.280..843S
3615:1986PNAS...83.4238N
3441:Curr Opin Chem Biol
3197:Annu. Rev. Biochem.
3153:Annu. Rev. Biochem.
3047:Cell. Mol. Life Sci
2936:2017JMolE..85..205H
2665:2005Natur.435...42L
2606:2000PNAS...97.4627L
2551:1995Sci...267.1463C
2428:"Calcium signaling"
2426:Clapham DE (2007).
2223:Trends Biochem. Sci
1923:Inorganic chemistry
1860:Albert L. Lehninger
1839:sugar phosphate by
1590:Tetrahydrobiopterin
832:Pyridoxal phosphate
636:. Other coenzymes,
602:Iron–sulfur protein
412:Pyrococcus furiosus
272:Inorganic cofactors
233:inorganic cofactors
142:at the junction of
114:essential nutrients
44:iron–sulfur centers
5469:Enzyme superfamily
5402:Enzyme promiscuity
5314:2016-10-05 at the
4959:10.1007/BF01874172
4864:10.1007/BF01506215
4824:. Nobel Foundation
4512:10.1007/BF01732468
3988:10.1093/jxb/erf038
3119:10.1042/BJ20061638
2016:"Enzyme Cofactors"
1655:
593:
576:Carbonic anhydrase
467:Cytochrome oxidase
421:carbonic anhydrase
337:deficiency causes
112:and other organic
52:
6037:
6036:
6008:
6007:
5625:
5624:
5318:(Powerpoint file)
5309:Cofactors lecture
5294:978-0-86542-793-8
5079:10.1021/bi700468t
5073:(18): 5283–5292.
4758:978-0-12-182013-8
4721:978-0-495-10935-8
3839:978-1-4020-0178-9
3812:978-0-943088-39-6
3534:978-0-86542-793-8
2867:978-0-495-39041-1
2842:978-1-57259-153-0
2813:978-0-85312-307-1
2798:Palmer T (1981).
2192:10.1002/prot.1093
2136:978-0-8247-4062-7
2091:978-0-12-492540-3
1905:receptor proteins
1885:mass spectroscopy
1727:. Such universal
1659:functional groups
1634:
1633:
1523:Nucleotide sugars
1247:
1246:
585:
584:
540:Nitrate reductase
474:Ferrous or Ferric
255:A 1980 letter in
249:prosthetic groups
148:citric acid cycle
102:organic molecules
79:chemical reaction
63:chemical compound
16:(Redirected from
6062:
5673:
5672:
5652:
5645:
5638:
5629:
5628:
5505:Hanes–Woolf plot
5448:Enzyme activator
5443:Enzyme inhibitor
5417:Enzyme catalysis
5361:
5354:
5347:
5338:
5337:
5322:Enzyme+cofactors
5298:
5286:
5267:
5266:
5256:
5224:
5218:
5217:
5207:
5201:
5200:
5190:
5180:
5148:
5142:
5141:
5131:
5097:
5091:
5090:
5062:
5053:
5052:
5042:
5018:
5012:
5011:
4985:
4979:
4978:
4942:
4936:
4935:
4915:
4909:
4908:
4906:
4882:
4876:
4875:
4839:
4833:
4832:
4830:
4829:
4823:
4815:
4809:
4808:
4806:
4782:
4776:
4775:
4774:
4773:
4732:
4726:
4725:
4707:
4701:
4700:
4673:
4667:
4666:
4656:
4624:
4618:
4617:
4589:
4583:
4582:
4572:
4562:
4538:
4532:
4531:
4487:
4481:
4480:
4462:
4430:
4424:
4423:
4392:Koch AL (1998).
4389:
4383:
4382:
4346:
4340:
4339:
4323:
4317:
4316:
4286:
4280:
4279:
4269:
4245:
4239:
4238:
4208:
4202:
4201:
4191:
4159:
4153:
4152:
4133:10.1038/280843a0
4108:
4102:
4101:
4069:
4063:
4062:
4042:
4036:
4035:
4007:
4001:
4000:
3990:
3981:(375): 1689–98.
3966:
3960:
3959:
3949:
3922:
3916:
3915:
3887:
3881:
3880:
3850:
3844:
3843:
3823:
3817:
3816:
3804:
3794:
3788:
3787:
3782:. Archived from
3743:
3737:
3736:
3726:
3694:
3688:
3687:
3677:
3653:
3647:
3646:
3636:
3626:
3594:
3588:
3587:
3569:
3545:
3539:
3538:
3514:
3508:
3507:
3471:
3465:
3464:
3436:
3427:
3426:
3416:
3392:
3386:
3385:
3375:
3357:
3348:
3342:
3341:
3305:
3299:
3298:
3270:
3264:
3263:
3235:
3229:
3228:
3183:
3177:
3176:
3147:
3141:
3140:
3130:
3098:
3089:
3088:
3070:
3053:(7–8): 892–905.
3038:
3032:
3031:
3021:
2997:
2991:
2990:
2970:
2964:
2963:
2930:(5–6): 205–218.
2919:
2913:
2912:
2902:
2878:
2872:
2871:
2853:
2847:
2846:
2834:
2824:
2818:
2817:
2805:
2795:
2789:
2788:
2752:
2746:
2745:
2735:
2725:
2701:
2695:
2694:
2676:
2644:
2638:
2637:
2627:
2617:
2585:
2579:
2578:
2545:(5203): 1463–9.
2534:
2528:
2527:
2507:
2501:
2500:
2472:
2466:
2465:
2447:
2423:
2417:
2416:
2388:
2382:
2381:
2371:
2347:
2341:
2340:
2304:
2298:
2297:
2269:
2263:
2262:
2260:
2259:
2245:
2239:
2238:
2218:
2212:
2211:
2175:
2166:
2165:
2163:
2162:
2147:
2141:
2140:
2122:
2116:
2115:
2113:
2102:
2096:
2095:
2083:
2073:
2056:
2055:
2037:
2031:
2030:
2028:
2027:
2018:. Archived from
2012:
2006:
2005:
2003:
2002:
1993:. Archived from
1987:
1981:
1980:
1962:
1953:
1918:Enzyme catalysis
1777:Candida boidinii
1254:
1253:
986:Pantothenic acid
731:
730:
710:, which carries
618:prosthetic group
548:Xanthine oxidase
450:
449:
423:from the marine
363:adenylate kinase
347:thyroid hormones
121:prosthetic group
21:
6070:
6069:
6065:
6064:
6063:
6061:
6060:
6059:
6040:
6039:
6038:
6033:
6004:
5951:
5946:
5938:
5829:
5820:
5812:
5796:
5782:
5772:
5764:
5756:
5746:
5736:
5726:
5704:
5690:
5664:
5656:
5626:
5621:
5533:Oxidoreductases
5519:
5495:Enzyme kinetics
5483:
5479:List of enzymes
5452:
5421:
5392:Catalytic triad
5370:
5365:
5316:Wayback Machine
5305:
5295:
5279:Bugg T (1997).
5275:
5273:Further reading
5270:
5225:
5221:
5216:(4th ed.).
5208:
5204:
5163:(20): 8069–74.
5149:
5145:
5098:
5094:
5063:
5056:
5019:
5015:
5008:
4986:
4982:
4943:
4939:
4916:
4912:
4883:
4879:
4840:
4836:
4827:
4825:
4821:
4817:
4816:
4812:
4797:(526): 369–75.
4783:
4779:
4771:
4769:
4759:
4733:
4729:
4722:
4708:
4704:
4697:
4687:10.1007/b100340
4675:
4674:
4670:
4645:10.1002/pro.227
4639:(10): 2125–38.
4633:Protein Science
4625:
4621:
4590:
4586:
4539:
4535:
4488:
4484:
4431:
4427:
4412:
4390:
4386:
4347:
4343:
4324:
4320:
4287:
4283:
4246:
4242:
4209:
4205:
4160:
4156:
4119:(5725): 843–4.
4109:
4105:
4070:
4066:
4043:
4039:
4008:
4004:
3967:
3963:
3923:
3919:
3892:Curr. Med. Chem
3888:
3884:
3851:
3847:
3840:
3824:
3820:
3813:
3795:
3791:
3744:
3740:
3695:
3691:
3668:(15): 4879–85.
3654:
3650:
3609:(12): 4238–42.
3595:
3591:
3546:
3542:
3535:
3517:Bugg T (1997).
3515:
3511:
3472:
3468:
3437:
3430:
3393:
3389:
3355:
3349:
3345:
3306:
3302:
3281:(2–3): 125–53.
3275:Prog. Lipid Res
3271:
3267:
3236:
3232:
3191:
3184:
3180:
3148:
3144:
3099:
3092:
3039:
3035:
2998:
2994:
2971:
2967:
2920:
2916:
2893:(11): 3009–19.
2879:
2875:
2868:
2854:
2850:
2843:
2825:
2821:
2814:
2796:
2792:
2753:
2749:
2702:
2698:
2674:10.1038/435042a
2645:
2641:
2586:
2582:
2535:
2531:
2508:
2504:
2473:
2469:
2424:
2420:
2389:
2385:
2348:
2344:
2305:
2301:
2270:
2266:
2257:
2255:
2247:
2246:
2242:
2219:
2215:
2176:
2169:
2160:
2158:
2149:
2148:
2144:
2137:
2123:
2119:
2111:
2103:
2099:
2092:
2074:
2059:
2052:
2038:
2034:
2025:
2023:
2014:
2013:
2009:
2000:
1998:
1989:
1988:
1984:
1960:
1954:
1950:
1946:
1914:
1893:
1873:phosphorylation
1868:
1817:
1811:
1740:history of life
1713:
1707:
1639:
1528:Monosaccharides
1383:Diacylglycerols
1274:Phosphate group
1252:
1225:
1187:
1149:
1033:
991:
962:
914:
876:
868:Methylcobalamin
842:
804:
762:
729:
680:
673:
666:
659:
613:
604:
598:
578:
574:
546:
542:
514:
510:
494:
490:
481:
390:nitrogen-fixing
285:
279:
274:
266:catalytic cycle
214:
87:enzyme kinetics
28:
23:
22:
15:
12:
11:
5:
6068:
6058:
6057:
6052:
6035:
6034:
6032:
6031:
6016:
6014:
6010:
6009:
6006:
6005:
6003:
6002:
5997:
5992:
5987:
5982:
5977:
5972:
5967:
5961:
5959:
5953:
5952:
5950:
5949:
5944:
5940:
5936:
5932:
5927:
5922:
5917:
5912:
5907:
5885:
5880:
5875:
5870:
5865:
5860:
5855:
5850:
5845:
5839:
5837:
5831:
5830:
5828:
5827:
5822:
5818:
5810:
5804:
5798:
5794:
5784:
5780:
5770:
5762:
5758:
5754:
5748:
5744:
5738:
5734:
5728:
5724:
5706:
5702:
5692:
5688:
5681:
5679:
5670:
5666:
5665:
5655:
5654:
5647:
5640:
5632:
5623:
5622:
5620:
5619:
5606:
5593:
5580:
5567:
5554:
5541:
5527:
5525:
5521:
5520:
5518:
5517:
5512:
5507:
5502:
5497:
5491:
5489:
5485:
5484:
5482:
5481:
5476:
5471:
5466:
5460:
5458:
5457:Classification
5454:
5453:
5451:
5450:
5445:
5440:
5435:
5429:
5427:
5423:
5422:
5420:
5419:
5414:
5409:
5404:
5399:
5394:
5389:
5384:
5378:
5376:
5372:
5371:
5364:
5363:
5356:
5349:
5341:
5335:
5334:
5329:
5319:
5304:
5303:External links
5301:
5300:
5299:
5293:
5274:
5271:
5269:
5268:
5239:(10): 2213–4.
5219:
5202:
5143:
5092:
5054:
5013:
5006:
4980:
4953:(1–2): 55–63.
4937:
4910:
4877:
4834:
4810:
4777:
4757:
4727:
4720:
4702:
4695:
4668:
4619:
4584:
4547:BMC Evol. Biol
4533:
4482:
4425:
4410:
4384:
4341:
4318:
4281:
4240:
4203:
4154:
4103:
4064:
4037:
4002:
3961:
3917:
3882:
3845:
3838:
3818:
3811:
3789:
3738:
3689:
3648:
3589:
3540:
3533:
3509:
3466:
3447:(2): 195–202.
3428:
3387:
3366:(8): 1817–30.
3343:
3316:(3): 183–223.
3300:
3265:
3230:
3189:
3178:
3142:
3090:
3033:
3012:(17): 7913–6.
2992:
2965:
2914:
2873:
2866:
2848:
2841:
2819:
2812:
2790:
2747:
2696:
2639:
2600:(9): 4627–31.
2580:
2529:
2502:
2467:
2438:(6): 1047–58.
2418:
2383:
2342:
2299:
2264:
2240:
2229:(3): N62–N63.
2213:
2167:
2142:
2135:
2117:
2097:
2090:
2057:
2051:978-1429224161
2050:
2032:
2007:
1982:
1947:
1945:
1942:
1941:
1940:
1935:
1930:
1925:
1920:
1913:
1910:
1892:
1889:
1867:
1864:
1856:Herman Kalckar
1810:
1807:
1706:
1703:
1671:dehydrogenases
1638:
1635:
1632:
1631:
1626:
1621:
1615:
1614:
1601:
1592:
1586:
1585:
1580:
1575:
1569:
1568:
1555:
1550:
1544:
1543:
1530:
1525:
1519:
1518:
1505:
1499:
1493:
1492:
1487:
1482:
1476:
1475:
1462:
1453:
1447:
1446:
1433:
1428:
1422:
1421:
1411:
1406:
1400:
1399:
1386:
1380:
1374:
1373:
1360:
1355:
1349:
1348:
1343:
1338:
1332:
1331:
1326:
1321:
1315:
1314:
1301:
1296:
1290:
1289:
1276:
1271:
1265:
1264:
1261:
1258:
1251:
1248:
1245:
1244:
1235:
1230:
1227:
1223:
1217:
1211:
1210:
1197:
1192:
1189:
1185:
1179:
1173:
1172:
1159:
1154:
1151:
1147:
1141:
1135:
1134:
1121:
1116:
1113:
1110:
1104:
1103:
1090:
1084:Carbonyl group
1081:
1078:
1075:
1069:
1068:
1055:
1041:
1035:
1031:
1025:
1019:
1018:
1005:
996:
993:
989:
983:
977:
976:
963:
960:
957:
954:
948:
942:
941:
928:
919:
916:
912:
906:
900:
899:
886:
881:
878:
874:
870:
864:
863:
850:
847:
844:
840:
834:
828:
827:
814:
809:
806:
802:
796:
786:
785:
772:
769:
764:
760:
754:
748:
747:
744:
741:
738:
735:
728:
725:
678:
671:
664:
657:
612:
609:
597:
594:
583:
582:
580:DNA polymerase
569:
563:
562:
557:
551:
550:
537:
531:
530:
525:
519:
518:
516:DNA polymerase
505:
499:
498:
476:
470:
469:
464:
458:
457:
454:
375:cell signaling
303:trace elements
283:Metalloprotein
278:
275:
273:
270:
213:
212:Classification
210:
204:in an ancient
133:nonpolypeptide
98:inorganic ions
26:
9:
6:
4:
3:
2:
6067:
6056:
6053:
6051:
6048:
6047:
6045:
6030:
6029:
6023:
6022:
6018:
6017:
6015:
6011:
6001:
5998:
5996:
5993:
5991:
5988:
5986:
5983:
5981:
5978:
5976:
5973:
5971:
5968:
5966:
5963:
5962:
5960:
5958:
5954:
5948:
5941:
5939:
5933:
5931:
5928:
5926:
5923:
5921:
5920:Molybdopterin
5918:
5916:
5913:
5911:
5908:
5905:
5901:
5897:
5893:
5889:
5886:
5884:
5881:
5879:
5876:
5874:
5873:Cofactor F430
5871:
5869:
5866:
5864:
5861:
5859:
5856:
5854:
5851:
5849:
5846:
5844:
5841:
5840:
5838:
5836:
5832:
5826:
5825:Coenzyme F420
5823:
5816:
5808:
5807:Phylloquinone
5805:
5802:
5801:Ascorbic acid
5799:
5792:
5788:
5785:
5778:
5774:
5766:
5759:
5752:
5749:
5742:
5739:
5732:
5729:
5722:
5718:
5714:
5710:
5707:
5700:
5696:
5693:
5686:
5683:
5682:
5680:
5678:
5674:
5671:
5667:
5663:
5660:
5653:
5648:
5646:
5641:
5639:
5634:
5633:
5630:
5617:
5613:
5612:
5607:
5604:
5600:
5599:
5594:
5591:
5587:
5586:
5581:
5578:
5574:
5573:
5568:
5565:
5561:
5560:
5555:
5552:
5548:
5547:
5542:
5539:
5535:
5534:
5529:
5528:
5526:
5522:
5516:
5513:
5511:
5508:
5506:
5503:
5501:
5498:
5496:
5493:
5492:
5490:
5486:
5480:
5477:
5475:
5474:Enzyme family
5472:
5470:
5467:
5465:
5462:
5461:
5459:
5455:
5449:
5446:
5444:
5441:
5439:
5438:Cooperativity
5436:
5434:
5431:
5430:
5428:
5424:
5418:
5415:
5413:
5410:
5408:
5405:
5403:
5400:
5398:
5397:Oxyanion hole
5395:
5393:
5390:
5388:
5385:
5383:
5380:
5379:
5377:
5373:
5369:
5362:
5357:
5355:
5350:
5348:
5343:
5342:
5339:
5333:
5330:
5327:
5323:
5320:
5317:
5313:
5310:
5307:
5306:
5296:
5290:
5285:
5284:
5277:
5276:
5264:
5260:
5255:
5250:
5246:
5242:
5238:
5234:
5230:
5223:
5215:
5214:
5206:
5198:
5194:
5189:
5184:
5179:
5174:
5170:
5166:
5162:
5158:
5154:
5147:
5139:
5135:
5130:
5125:
5121:
5117:
5113:
5109:
5108:
5103:
5096:
5088:
5084:
5080:
5076:
5072:
5068:
5061:
5059:
5050:
5046:
5041:
5036:
5033:(2): 611–23.
5032:
5028:
5027:J. Biol. Chem
5024:
5017:
5009:
5007:9780674366701
5003:
4999:
4995:
4991:
4984:
4976:
4972:
4968:
4964:
4960:
4956:
4952:
4948:
4941:
4933:
4929:
4925:
4921:
4914:
4905:
4900:
4897:(1): 173–90.
4896:
4892:
4891:J. Biol. Chem
4888:
4881:
4873:
4869:
4865:
4861:
4857:
4853:
4850:(31): 624–5.
4849:
4845:
4838:
4820:
4814:
4805:
4800:
4796:
4792:
4788:
4781:
4768:
4764:
4760:
4754:
4750:
4746:
4742:
4738:
4731:
4723:
4717:
4713:
4706:
4698:
4696:0-306-46712-7
4692:
4688:
4684:
4680:
4679:
4672:
4664:
4660:
4655:
4650:
4646:
4642:
4638:
4634:
4630:
4623:
4615:
4611:
4607:
4603:
4600:(9): 877–88.
4599:
4595:
4588:
4580:
4576:
4571:
4566:
4561:
4556:
4552:
4548:
4544:
4537:
4529:
4525:
4521:
4517:
4513:
4509:
4505:
4501:
4497:
4493:
4486:
4478:
4474:
4470:
4466:
4461:
4456:
4452:
4448:
4445:(2): 119–23.
4444:
4440:
4436:
4429:
4421:
4417:
4413:
4411:9780120277407
4407:
4403:
4399:
4395:
4388:
4380:
4376:
4372:
4368:
4364:
4360:
4357:(4): 633–55.
4356:
4352:
4345:
4337:
4333:
4329:
4322:
4314:
4310:
4306:
4302:
4298:
4294:
4293:
4285:
4277:
4273:
4268:
4263:
4259:
4255:
4251:
4244:
4236:
4232:
4228:
4224:
4220:
4216:
4215:
4207:
4199:
4195:
4190:
4185:
4181:
4177:
4173:
4169:
4165:
4158:
4150:
4146:
4142:
4138:
4134:
4130:
4126:
4122:
4118:
4114:
4107:
4099:
4095:
4091:
4087:
4084:(2): 149–57.
4083:
4079:
4075:
4068:
4060:
4056:
4052:
4048:
4041:
4033:
4029:
4025:
4021:
4018:(7): 621–35.
4017:
4013:
4006:
3998:
3994:
3989:
3984:
3980:
3976:
3972:
3965:
3957:
3953:
3948:
3943:
3940:(3): 919–24.
3939:
3935:
3931:
3927:
3921:
3913:
3909:
3905:
3901:
3897:
3893:
3886:
3878:
3874:
3870:
3866:
3862:
3858:
3857:
3849:
3841:
3835:
3831:
3830:
3822:
3814:
3808:
3803:
3802:
3793:
3785:
3781:
3777:
3773:
3769:
3765:
3761:
3757:
3753:
3749:
3742:
3734:
3730:
3725:
3720:
3716:
3712:
3709:(1): 256–63.
3708:
3704:
3700:
3693:
3685:
3681:
3676:
3671:
3667:
3663:
3659:
3652:
3644:
3640:
3635:
3630:
3625:
3620:
3616:
3612:
3608:
3604:
3600:
3593:
3585:
3581:
3577:
3573:
3568:
3563:
3560:(4): 471–80.
3559:
3555:
3554:FASEB Journal
3551:
3544:
3536:
3530:
3526:
3522:
3521:
3513:
3505:
3501:
3497:
3493:
3489:
3485:
3482:(3): 265–75.
3481:
3477:
3470:
3462:
3458:
3454:
3450:
3446:
3442:
3435:
3433:
3424:
3420:
3415:
3410:
3406:
3402:
3398:
3391:
3383:
3379:
3374:
3369:
3365:
3361:
3354:
3347:
3339:
3335:
3331:
3327:
3323:
3319:
3315:
3311:
3304:
3296:
3292:
3288:
3284:
3280:
3276:
3269:
3261:
3257:
3253:
3249:
3246:(3): 217–29.
3245:
3241:
3234:
3226:
3222:
3218:
3214:
3210:
3206:
3202:
3199:
3198:
3193:
3182:
3174:
3170:
3166:
3162:
3158:
3155:
3154:
3146:
3138:
3134:
3129:
3124:
3120:
3116:
3113:(2): 205–18.
3112:
3108:
3104:
3097:
3095:
3086:
3082:
3078:
3074:
3069:
3064:
3060:
3056:
3052:
3048:
3044:
3037:
3029:
3025:
3020:
3015:
3011:
3007:
3003:
2996:
2988:
2984:
2981:(10): 912–5.
2980:
2976:
2969:
2961:
2957:
2953:
2949:
2945:
2941:
2937:
2933:
2929:
2925:
2918:
2910:
2906:
2901:
2896:
2892:
2888:
2884:
2877:
2869:
2863:
2859:
2852:
2844:
2838:
2833:
2832:
2823:
2815:
2809:
2804:
2803:
2794:
2786:
2782:
2778:
2774:
2770:
2766:
2763:(2): 157–70.
2762:
2758:
2751:
2743:
2739:
2734:
2729:
2724:
2719:
2715:
2711:
2707:
2700:
2692:
2688:
2684:
2680:
2675:
2670:
2666:
2662:
2658:
2654:
2650:
2643:
2635:
2631:
2626:
2621:
2616:
2611:
2607:
2603:
2599:
2595:
2591:
2584:
2576:
2572:
2568:
2564:
2560:
2556:
2552:
2548:
2544:
2540:
2533:
2525:
2521:
2517:
2513:
2506:
2498:
2494:
2490:
2486:
2483:(4): 685–98.
2482:
2478:
2471:
2463:
2459:
2455:
2451:
2446:
2441:
2437:
2433:
2429:
2422:
2414:
2410:
2406:
2402:
2399:(2): 177–81.
2398:
2394:
2387:
2379:
2375:
2370:
2365:
2361:
2357:
2353:
2346:
2338:
2334:
2330:
2326:
2322:
2318:
2315:(3): 149–62.
2314:
2310:
2303:
2295:
2291:
2287:
2283:
2280:(3): 513–43.
2279:
2275:
2268:
2254:
2250:
2244:
2236:
2232:
2228:
2224:
2217:
2209:
2205:
2201:
2197:
2193:
2189:
2186:(3): 282–91.
2185:
2181:
2174:
2172:
2156:
2152:
2146:
2138:
2132:
2128:
2121:
2110:
2109:
2101:
2093:
2087:
2082:
2081:
2072:
2070:
2068:
2066:
2064:
2062:
2053:
2047:
2043:
2036:
2022:on 2003-05-05
2021:
2017:
2011:
1997:on 1999-08-26
1996:
1992:
1986:
1978:
1974:
1970:
1966:
1959:
1952:
1948:
1939:
1936:
1934:
1931:
1929:
1926:
1924:
1921:
1919:
1916:
1915:
1909:
1906:
1902:
1898:
1888:
1886:
1882:
1878:
1874:
1863:
1861:
1857:
1853:
1848:
1846:
1842:
1838:
1834:
1830:
1826:
1822:
1821:Arthur Harden
1816:
1806:
1804:
1800:
1796:
1792:
1788:
1784:
1780:
1778:
1774:
1770:
1768:
1767:
1762:
1757:
1754:, with early
1753:
1749:
1745:
1741:
1736:
1734:
1730:
1726:
1722:
1718:
1712:
1702:
1699:
1695:
1691:
1686:
1684:
1680:
1676:
1672:
1666:
1664:
1660:
1652:
1649:reactions of
1648:
1643:
1630:
1627:
1625:
1622:
1620:
1617:
1616:
1613:
1609:
1605:
1602:
1600:
1596:
1593:
1591:
1588:
1587:
1584:
1581:
1579:
1576:
1574:
1571:
1570:
1567:
1563:
1559:
1556:
1554:
1553:Sulfate group
1551:
1549:
1546:
1545:
1542:
1538:
1534:
1531:
1529:
1526:
1524:
1521:
1520:
1517:
1513:
1509:
1506:
1503:
1500:
1498:
1497:Molybdopterin
1495:
1494:
1491:
1488:
1486:
1483:
1481:
1478:
1477:
1474:
1470:
1466:
1463:
1461:
1457:
1454:
1452:
1449:
1448:
1445:
1441:
1437:
1434:
1432:
1429:
1427:
1424:
1423:
1420:
1416:
1412:
1410:
1407:
1405:
1402:
1401:
1398:
1394:
1390:
1387:
1384:
1381:
1379:
1376:
1375:
1372:
1368:
1364:
1361:
1359:
1356:
1354:
1351:
1350:
1347:
1344:
1342:
1339:
1337:
1334:
1333:
1330:
1327:
1325:
1322:
1320:
1317:
1316:
1313:
1309:
1305:
1302:
1300:
1297:
1295:
1292:
1291:
1288:
1284:
1280:
1277:
1275:
1272:
1270:
1267:
1266:
1263:Distribution
1262:
1259:
1256:
1255:
1243:
1239:
1236:
1234:
1231:
1228:
1221:
1218:
1216:
1215:Coenzyme F420
1213:
1212:
1209:
1205:
1201:
1198:
1196:
1193:
1190:
1183:
1180:
1178:
1175:
1174:
1171:
1167:
1163:
1160:
1158:
1155:
1152:
1145:
1142:
1140:
1137:
1136:
1133:
1129:
1125:
1122:
1120:
1117:
1114:
1111:
1109:
1108:Ascorbic acid
1106:
1105:
1102:
1098:
1094:
1091:
1089:
1085:
1082:
1079:
1076:
1074:
1071:
1070:
1067:
1063:
1059:
1056:
1053:
1049:
1045:
1042:
1039:
1036:
1029:
1026:
1024:
1021:
1020:
1017:
1013:
1009:
1006:
1004:
1000:
997:
994:
987:
984:
982:
979:
978:
975:
971:
967:
964:
958:
955:
952:
949:
947:
944:
943:
940:
936:
932:
929:
927:
923:
920:
917:
910:
907:
905:
902:
901:
898:
894:
890:
887:
885:
882:
879:
877:
871:
869:
866:
865:
862:
858:
854:
851:
848:
845:
838:
835:
833:
830:
829:
826:
822:
818:
815:
813:
810:
807:
800:
797:
795:
791:
788:
787:
784:
780:
776:
773:
770:
768:
767:pyrophosphate
765:
758:
755:
753:
750:
749:
746:Distribution
745:
742:
739:
736:
733:
732:
724:
722:
718:
713:
709:
705:
701:
697:
693:
689:
685:
681:
674:
667:
660:
653:
649:
647:
643:
639:
635:
631:
627:
626:transketolase
623:
619:
608:
603:
589:
581:
577:
573:
570:
568:
565:
564:
561:
558:
556:
553:
552:
549:
545:
541:
538:
536:
533:
532:
529:
526:
524:
521:
520:
517:
513:
509:
506:
504:
501:
500:
497:
493:
488:
484:
480:
477:
475:
472:
471:
468:
465:
463:
460:
459:
455:
452:
451:
448:
446:
442:
438:
433:
431:
430:
426:
422:
418:
414:
413:
409:
405:
401:
397:
396:
391:
387:
383:
378:
376:
372:
368:
364:
360:
356:
352:
348:
344:
340:
336:
332:
328:
324:
320:
316:
312:
308:
304:
300:
296:
292:
289:
284:
269:
267:
263:
258:
253:
251:
250:
245:
240:
238:
234:
230:
226:
222:
219:
209:
207:
203:
199:
195:
191:
187:
183:
180:
176:
171:
169:
165:
161:
157:
153:
149:
145:
141:
136:
134:
130:
126:
122:
117:
115:
111:
107:
103:
99:
94:
92:
88:
84:
80:
76:
73:'s role as a
72:
68:
64:
61:
57:
49:
45:
41:
37:
32:
19:
6025:
6019:
5925:Mycofactocin
5915:Methanofuran
5835:non-vitamins
5669:Active forms
5661:
5611:Translocases
5608:
5595:
5582:
5569:
5556:
5546:Transferases
5543:
5530:
5411:
5387:Binding site
5282:
5236:
5232:
5222:
5212:
5205:
5160:
5156:
5146:
5111:
5105:
5095:
5070:
5067:Biochemistry
5066:
5030:
5026:
5016:
4989:
4983:
4950:
4946:
4940:
4923:
4919:
4913:
4894:
4890:
4880:
4847:
4843:
4837:
4826:. Retrieved
4813:
4794:
4790:
4780:
4770:, retrieved
4740:
4730:
4712:Biochemistry
4711:
4705:
4677:
4671:
4636:
4632:
4622:
4597:
4593:
4587:
4550:
4546:
4536:
4498:(2): 101–4.
4495:
4491:
4485:
4442:
4439:FEBS Letters
4438:
4428:
4393:
4387:
4354:
4350:
4344:
4335:
4331:
4321:
4296:
4290:
4284:
4257:
4253:
4243:
4218:
4212:
4206:
4171:
4167:
4157:
4116:
4112:
4106:
4081:
4077:
4067:
4050:
4046:
4040:
4015:
4011:
4005:
3978:
3974:
3964:
3937:
3933:
3920:
3898:(8): 981–6.
3895:
3891:
3885:
3860:
3854:
3848:
3832:. Springer.
3828:
3821:
3800:
3792:
3784:the original
3758:(6): 591–8.
3755:
3751:
3741:
3706:
3702:
3692:
3665:
3661:
3651:
3606:
3602:
3592:
3557:
3553:
3543:
3519:
3512:
3479:
3475:
3469:
3444:
3440:
3404:
3400:
3390:
3363:
3360:Microbiology
3359:
3346:
3313:
3309:
3303:
3278:
3274:
3268:
3243:
3239:
3233:
3200:
3195:
3181:
3156:
3151:
3145:
3110:
3106:
3050:
3046:
3036:
3009:
3005:
2995:
2978:
2974:
2968:
2927:
2923:
2917:
2890:
2886:
2876:
2858:Biochemistry
2857:
2851:
2830:
2822:
2801:
2793:
2760:
2756:
2750:
2713:
2709:
2699:
2659:(7038): 42.
2656:
2652:
2642:
2597:
2593:
2583:
2542:
2538:
2532:
2518:(2): 111–6.
2515:
2511:
2505:
2480:
2476:
2470:
2435:
2431:
2421:
2396:
2392:
2386:
2362:(4): 715–8.
2359:
2355:
2345:
2312:
2308:
2302:
2277:
2273:
2267:
2256:. Retrieved
2253:vle.du.ac.in
2252:
2243:
2226:
2222:
2216:
2183:
2179:
2159:. Retrieved
2157:. 2013-10-02
2154:
2145:
2126:
2120:
2107:
2100:
2079:
2041:
2035:
2024:. Retrieved
2020:the original
2010:
1999:. Retrieved
1995:the original
1985:
1971:(2): 93–94.
1968:
1964:
1951:
1894:
1869:
1849:
1832:
1818:
1782:
1781:
1776:
1772:
1771:
1764:
1737:
1729:conservation
1714:
1687:
1667:
1656:
1624:Methyl group
1485:Formyl group
1480:Methanofuran
1341:Methyl group
1299:Methyl group
1250:Non-vitamins
999:Acetyl group
926:alkyl groups
880:Methyl group
650:
614:
605:
434:
427:
410:
393:
379:
286:
256:
254:
247:
243:
241:
232:
220:
217:
215:
172:
137:
118:
100:and complex
95:
67:metallic ion
55:
53:
5910:Lipoic Acid
5888:Heme / Haem
5815:Menaquinone
5382:Active site
4926:: E79–E88.
4299:: 1031–78.
4260:(1): 1–20.
4174:(1): 1–16.
4053:: 595–600.
3407:(1): 1–22.
3159:: 383–415.
1789:(coenzyme:
1711:Abiogenesis
1629:Methanogens
1490:Methanogens
1460:acyl groups
1404:Glutathione
1346:Methanogens
1329:Methanogens
1238:Methanogens
1229:Amino acids
1073:Menaquinone
1003:acyl groups
884:acyl groups
717:methanogens
544:Nitrogenase
496:Hydrogenase
492:Nitrogenase
415:, and even
395:Azotobacter
386:nitrogenase
333:. Although
83:biochemical
6044:Categories
6013:Base forms
5957:metal ions
5883:Coenzyme Q
5878:Coenzyme M
5868:Coenzyme B
5731:Coenzyme A
5685:TPP / ThDP
5585:Isomerases
5559:Hydrolases
5426:Regulation
5114:: 531–50.
4828:2007-09-30
4772:2024-09-21
4338:(2): 70–1.
4221:: 355–94.
3926:Vorholt JA
3863:: 711–60.
3203:: 209–47.
3107:Biochem. J
2512:BioFactors
2309:BioFactors
2258:2018-02-07
2161:2017-05-10
2026:2007-11-17
2001:2007-11-17
1944:References
1837:nucleotide
1766:exaptation
1725:metabolism
1698:hydrolysis
1690:metabolism
1683:reductases
1612:eukaryotes
1566:eukaryotes
1541:eukaryotes
1516:eukaryotes
1473:eukaryotes
1444:eukaryotes
1419:eukaryotes
1397:eukaryotes
1371:eukaryotes
1353:Coenzyme Q
1336:Coenzyme M
1319:Coenzyme B
1312:eukaryotes
1287:eukaryotes
1220:Riboflavin
1208:eukaryotes
1182:Riboflavin
1170:eukaryotes
1144:Riboflavin
1132:eukaryotes
1101:eukaryotes
1066:eukaryotes
1028:Folic acid
1016:eukaryotes
1001:and other
981:Coenzyme A
974:eukaryotes
939:eukaryotes
909:Cobalamine
904:Cobalamine
897:eukaryotes
861:eukaryotes
837:Pyridoxine
825:eukaryotes
783:eukaryotes
708:coenzyme A
696:nucleotide
688:folic acid
535:Molybdenum
512:Hexokinase
483:Cytochrome
371:calmodulin
331:molybdenum
277:Metal ions
223:, such as
190:coenzyme A
179:nucleotide
168:coenzyme A
166:(NAD) and
144:glycolysis
129:holoenzyme
6055:Cofactors
5943:THMPT / H
5741:PLP / P5P
5662:cofactors
5464:EC number
4594:Biochimie
3338:218866247
1833:coferment
1756:ribozymes
1752:RNA world
1744:adenosine
1705:Evolution
1673:that use
1663:coenzymes
1599:electrons
1597:atom and
1578:Electrons
1456:Electrons
1451:Lipoamide
1431:Electrons
1417:and most
1409:Electrons
1358:Electrons
1324:Electrons
1240:and some
1233:Electrons
1195:Electrons
1157:Electrons
1119:Electrons
1112:Vitamin C
1088:electrons
1077:Vitamin K
1052:methylene
1038:Glutamate
873:Vitamin B
812:Electrons
692:vitamin C
646:lipoamide
523:Manganese
503:Magnesium
441:porphyrin
315:manganese
311:magnesium
299:nutrition
262:substrate
244:coenzymes
221:cofactors
206:RNA world
202:ribozymes
156:lipoamide
125:apoenzyme
106:coenzymes
58:is a non-
6028:vitamins
6021:vitamins
5935:THB / BH
5769:DHFA / H
5761:THFA / H
5677:vitamins
5488:Kinetics
5412:Cofactor
5375:Activity
5312:Archived
5263:18701638
5197:23633564
5138:23746262
5087:17439161
5049:18116985
4975:26999163
4872:20328411
4681:. 2004.
4663:19693930
4614:12458080
4579:14687414
4528:22282629
4477:39128865
4379:44873410
4371:17443876
4198:10727395
4098:11396917
4032:16784786
3997:12147719
3912:15078160
3780:28013583
3772:11771674
3584:11214528
3504:12634062
3496:16607521
3461:17275397
3423:17222174
3382:10463148
3330:11451208
3295:15893380
3260:12769720
3225:37393683
3217:14527323
3173:15189147
3137:17295611
3085:20415735
3077:17429582
3068:11136255
2987:17086936
2952:29177972
2785:21961142
2777:17992543
2742:21371326
2691:52819760
2683:15875011
2634:10781068
2575:20868012
2462:15087548
2454:18083096
2378:10736319
2337:19417496
2329:10875302
2208:10848692
2200:11455601
2180:Proteins
1912:See also
1901:hormones
1899:such as
1797:(NAD⁺),
1604:Bacteria
1583:Bacteria
1558:Bacteria
1533:Bacteria
1508:Bacteria
1465:Bacteria
1436:Bacteria
1415:bacteria
1389:Bacteria
1363:Bacteria
1304:Bacteria
1279:Bacteria
1257:Cofactor
1242:bacteria
1200:Bacteria
1162:Bacteria
1124:Bacteria
1093:Bacteria
1058:Bacteria
1040:residues
1008:Bacteria
966:Bacteria
931:Bacteria
922:hydrogen
889:Bacteria
853:Bacteria
817:Bacteria
775:Bacteria
757:Thiamine
734:Cofactor
652:Vitamins
528:Arginase
479:Catalase
408:archaean
400:tungsten
382:vanadium
335:chromium
175:vitamins
146:and the
110:vitamins
75:catalyst
56:cofactor
18:Coenzyme
6050:Enzymes
5598:Ligases
5368:Enzymes
5254:2582534
5188:3657804
5165:Bibcode
5129:4082410
4967:4279328
4852:Bibcode
4767:3003504
4654:2786976
4520:1263263
4500:Bibcode
4469:8706840
4447:Bibcode
4420:9889982
4235:2115763
4189:1220924
4149:3094647
4121:Bibcode
3956:9342247
3877:6137189
3684:4367810
3643:3086878
3611:Bibcode
3576:8647346
3128:1798440
3028:3131330
2960:7120148
2932:Bibcode
2909:4968184
2733:3059290
2661:Bibcode
2602:Bibcode
2567:7878465
2547:Bibcode
2539:Science
2524:3076437
2497:8947828
2413:9133680
2393:Thyroid
2294:3905079
1897:ligands
1809:History
1761:domains
1608:archaea
1562:archaea
1537:archaea
1512:archaea
1469:archaea
1440:archaea
1393:archaea
1367:archaea
1308:archaea
1283:archaea
1204:archaea
1166:archaea
1128:archaea
1097:archaea
1062:archaea
1012:archaea
970:archaea
935:archaea
893:archaea
857:archaea
821:archaea
779:archaea
737:Vitamin
721:archaea
640:(FAD),
611:Organic
419:in the
417:cadmium
402:in the
388:of the
384:in the
351:Calcium
218:organic
104:called
91:ligands
60:protein
5975:Fe, Fe
5787:AdoCbl
5751:Biotin
5659:Enzyme
5572:Lyases
5328:(MeSH)
5291:
5261:
5251:
5195:
5185:
5136:
5126:
5085:
5047:
5004:
4973:
4965:
4870:
4765:
4755:
4718:
4693:
4661:
4651:
4612:
4577:
4570:317284
4567:
4553:: 26.
4526:
4518:
4475:
4467:
4418:
4408:
4377:
4369:
4313:354490
4311:
4276:378655
4274:
4233:
4196:
4186:
4147:
4141:471057
4139:
4113:Nature
4096:
4059:351635
4057:
4030:
3995:
3954:
3910:
3875:
3836:
3809:
3778:
3770:
3733:104960
3731:
3724:218444
3721:
3682:
3641:
3634:323707
3631:
3582:
3574:
3531:
3502:
3494:
3459:
3421:
3401:FEBS J
3380:
3336:
3328:
3293:
3258:
3223:
3215:
3171:
3135:
3125:
3083:
3075:
3065:
3026:
2985:
2958:
2950:
2907:
2864:
2839:
2810:
2783:
2775:
2740:
2730:
2716:: 13.
2689:
2681:
2653:Nature
2632:
2622:
2573:
2565:
2522:
2495:
2460:
2452:
2411:
2376:
2335:
2327:
2292:
2206:
2198:
2133:
2088:
2048:
1679:reduce
1595:Oxygen
1502:Oxygen
1048:formyl
1044:Methyl
951:Biotin
946:Biotin
799:Niacin
706:, and
684:niacin
644:, and
642:biotin
560:Urease
555:Nickel
462:Cupric
425:diatom
361:, and
343:Iodine
329:, and
323:copper
319:cobalt
231:; and
225:flavin
196:, and
71:enzyme
46:, and
40:flavin
5791:MeCbl
5721:NADPH
5524:Types
4971:S2CID
4868:S2CID
4822:(PDF)
4524:S2CID
4473:S2CID
4375:S2CID
4145:S2CID
3776:S2CID
3580:S2CID
3500:S2CID
3356:(PDF)
3334:S2CID
3221:S2CID
3081:S2CID
2956:S2CID
2781:S2CID
2687:S2CID
2625:18283
2571:S2CID
2458:S2CID
2333:S2CID
2204:S2CID
2112:(PDF)
1961:(PDF)
1825:yeast
1803:NADPH
1748:redox
1647:redox
1504:atoms
1413:Some
485:(via
297:. In
288:Metal
6026:see
5858:PAPS
5853:SAMe
5777:MTHF
5717:NADP
5713:NADH
5616:list
5609:EC7
5603:list
5596:EC6
5590:list
5583:EC5
5577:list
5570:EC4
5564:list
5557:EC3
5551:list
5544:EC2
5538:list
5531:EC1
5289:ISBN
5259:PMID
5193:PMID
5134:PMID
5083:PMID
5045:PMID
5002:ISBN
4963:PMID
4763:PMID
4753:ISBN
4716:ISBN
4691:ISBN
4659:PMID
4610:PMID
4575:PMID
4516:PMID
4465:PMID
4416:PMID
4406:ISBN
4367:PMID
4309:PMID
4272:PMID
4231:PMID
4194:PMID
4137:PMID
4094:PMID
4055:PMID
4028:PMID
4016:1763
3993:PMID
3952:PMID
3908:PMID
3873:PMID
3834:ISBN
3807:ISBN
3768:PMID
3729:PMID
3680:PMID
3639:PMID
3572:PMID
3529:ISBN
3492:PMID
3457:PMID
3419:PMID
3378:PMID
3326:PMID
3291:PMID
3256:PMID
3213:PMID
3169:PMID
3133:PMID
3073:PMID
3024:PMID
2983:PMID
2948:PMID
2905:PMID
2862:ISBN
2837:ISBN
2808:ISBN
2773:PMID
2738:PMID
2679:PMID
2630:PMID
2563:PMID
2520:PMID
2493:PMID
2450:PMID
2432:Cell
2409:PMID
2374:PMID
2325:PMID
2290:PMID
2196:PMID
2131:ISBN
2086:ISBN
2046:ISBN
1883:and
1791:NAD⁺
1721:NADH
1719:and
1694:mole
1645:The
1610:and
1564:and
1539:and
1514:and
1471:and
1442:and
1426:Heme
1395:and
1369:and
1310:and
1285:and
1206:and
1168:and
1153:None
1130:and
1115:None
1099:and
1086:and
1080:None
1064:and
1014:and
972:and
956:None
937:and
918:None
895:and
859:and
846:None
823:and
794:NADP
792:and
781:and
712:acyl
702:and
567:Zinc
487:Heme
445:iron
437:heme
327:zinc
307:iron
291:ions
246:and
229:heme
158:and
48:heme
34:The
5947:MPT
5930:PQQ
5863:GSH
5848:CTP
5843:ATP
5813:),
5803:(C)
5709:NAD
5699:FAD
5695:FMN
5249:PMC
5241:doi
5183:PMC
5173:doi
5161:110
5124:PMC
5116:doi
5075:doi
5035:doi
5031:178
4994:doi
4955:doi
4928:doi
4924:287
4899:doi
4895:160
4860:doi
4799:doi
4745:doi
4683:doi
4649:PMC
4641:doi
4602:doi
4565:PMC
4555:doi
4508:doi
4455:doi
4443:390
4398:doi
4359:doi
4301:doi
4262:doi
4223:doi
4184:PMC
4176:doi
4172:347
4129:doi
4117:280
4086:doi
4082:390
4020:doi
3983:doi
3942:doi
3938:248
3900:doi
3865:doi
3760:doi
3719:PMC
3711:doi
3707:137
3670:doi
3666:249
3629:PMC
3619:doi
3562:doi
3484:doi
3449:doi
3409:doi
3405:274
3368:doi
3364:145
3318:doi
3283:doi
3248:doi
3205:doi
3161:doi
3123:PMC
3115:doi
3111:402
3063:PMC
3055:doi
3014:doi
3010:263
2940:doi
2895:doi
2891:243
2765:doi
2728:PMC
2718:doi
2669:doi
2657:435
2620:PMC
2610:doi
2555:doi
2543:267
2485:doi
2481:120
2440:doi
2436:131
2401:doi
2364:doi
2360:130
2317:doi
2282:doi
2231:doi
2188:doi
1973:doi
1805:).
1793:),
1717:ATP
1191:ADP
995:ADP
953:(H)
808:ADP
790:NAD
704:FAD
700:NAD
628:or
453:Ion
227:or
198:NAD
194:FAD
186:ATP
65:or
6046::
6024::
6000:Zn
5995:Ni
5990:Mo
5985:Mn
5980:Mg
5970:Cu
5965:Ca
5902:,
5898:,
5894:,
5817:(K
5809:(K
5795:12
5793:(B
5789:,
5779:(B
5775:,
5773:FA
5767:,
5765:FA
5753:(B
5743:(B
5733:(B
5723:(B
5719:,
5715:,
5711:,
5701:(B
5697:,
5687:(B
5257:.
5247:.
5237:22
5235:.
5231:.
5191:.
5181:.
5171:.
5159:.
5155:.
5132:.
5122:.
5112:82
5110:.
5104:.
5081:.
5071:46
5069:.
5057:^
5043:.
5029:.
5025:.
5000:.
4969:.
4961:.
4949:.
4922:.
4893:.
4889:.
4866:.
4858:.
4848:17
4846:.
4795:78
4793:.
4789:.
4761:,
4751:,
4739:,
4689:.
4657:.
4647:.
4637:18
4635:.
4631:.
4608:.
4598:84
4596:.
4573:.
4563:.
4549:.
4545:.
4522:.
4514:.
4506:.
4494:.
4471:.
4463:.
4453:.
4441:.
4437:.
4414:.
4404:.
4373:.
4365:.
4353:.
4336:25
4334:.
4330:.
4307:.
4297:47
4295:.
4270:.
4258:95
4256:.
4252:.
4229:.
4219:59
4217:.
4192:.
4182:.
4170:.
4166:.
4143:.
4135:.
4127:.
4115:.
4092:.
4080:.
4076:.
4051:23
4049:.
4026:.
4014:.
3991:.
3979:53
3977:.
3973:.
3950:.
3936:.
3932:.
3906:.
3896:11
3894:.
3871:.
3861:52
3859:.
3774:.
3766:.
3756:20
3754:.
3750:.
3727:.
3717:.
3705:.
3701:.
3678:.
3664:.
3660:.
3637:.
3627:.
3617:.
3607:83
3605:.
3601:.
3578:.
3570:.
3558:10
3556:.
3552:.
3527:.
3525:95
3498:.
3490:.
3480:71
3478:.
3455:.
3445:11
3443:.
3431:^
3417:.
3403:.
3399:.
3376:.
3362:.
3358:.
3332:.
3324:.
3314:38
3312:.
3289:.
3279:44
3277:.
3254:.
3242:.
3219:.
3211:.
3201:72
3194:.
3190:12
3167:.
3157:73
3131:.
3121:.
3109:.
3105:.
3093:^
3079:.
3071:.
3061:.
3051:64
3049:.
3045:.
3022:.
3008:.
3004:.
2977:.
2954:.
2946:.
2938:.
2928:85
2926:.
2903:.
2889:.
2885:.
2779:.
2771:.
2761:13
2759:.
2736:.
2726:.
2714:11
2712:.
2708:.
2685:.
2677:.
2667:.
2655:.
2651:.
2628:.
2618:.
2608:.
2598:97
2596:.
2592:.
2569:.
2561:.
2553:.
2541:.
2514:.
2491:.
2479:.
2456:.
2448:.
2434:.
2430:.
2407:.
2395:.
2372:.
2358:.
2354:.
2331:.
2323:.
2313:11
2311:.
2288:.
2278:14
2276:.
2251:.
2225:.
2202:.
2194:.
2184:44
2182:.
2170:^
2153:.
2060:^
1969:22
1967:.
1963:.
1847:.
1769:.
1665:.
1606:,
1560:,
1535:,
1510:,
1467:,
1458:,
1438:,
1391:,
1365:,
1306:,
1281:,
1222:(B
1202:,
1184:(B
1164:,
1146:(B
1126:,
1095:,
1060:,
1050:,
1046:,
1030:(B
1010:,
988:(B
968:,
959:CO
933:,
924:,
913:12
911:(B
891:,
875:12
855:,
839:(B
819:,
801:(B
777:,
759:(B
723:.
686:,
682:,
679:12
675:,
668:,
661:,
447:.
432:.
398:,
357:,
325:,
321:,
317:,
313:,
309:,
239:.
192:,
188:,
54:A
42:,
5945:4
5937:4
5906:)
5904:O
5900:C
5896:B
5892:A
5890:(
5821:)
5819:2
5811:1
5797:)
5783:)
5781:9
5771:2
5763:4
5757:)
5755:7
5747:)
5745:6
5737:)
5735:5
5727:)
5725:3
5705:)
5703:2
5691:)
5689:1
5651:e
5644:t
5637:v
5618:)
5614:(
5605:)
5601:(
5592:)
5588:(
5579:)
5575:(
5566:)
5562:(
5553:)
5549:(
5540:)
5536:(
5360:e
5353:t
5346:v
5297:.
5265:.
5243::
5199:.
5175::
5167::
5140:.
5118::
5089:.
5077::
5051:.
5037::
5010:.
4996::
4977:.
4957::
4951:5
4934:.
4930::
4907:.
4901::
4874:.
4862::
4854::
4831:.
4807:.
4801::
4747::
4724:.
4699:.
4685::
4665:.
4643::
4616:.
4604::
4581:.
4557::
4551:3
4530:.
4510::
4502::
4496:7
4479:.
4457::
4449::
4422:.
4400::
4381:.
4361::
4355:4
4315:.
4303::
4278:.
4264::
4237:.
4225::
4200:.
4178::
4151:.
4131::
4123::
4100:.
4088::
4061:.
4034:.
4022::
3999:.
3985::
3958:.
3944::
3914:.
3902::
3879:.
3867::
3842:.
3815:.
3762::
3735:.
3713::
3686:.
3672::
3645:.
3621::
3613::
3586:.
3564::
3537:.
3506:.
3486::
3463:.
3451::
3425:.
3411::
3384:.
3370::
3340:.
3320::
3297:.
3285::
3262:.
3250::
3244:4
3227:.
3207::
3175:.
3163::
3139:.
3117::
3087:.
3057::
3030:.
3016::
2989:.
2979:7
2962:.
2942::
2934::
2911:.
2897::
2870:.
2845:.
2816:.
2787:.
2767::
2744:.
2720::
2693:.
2671::
2663::
2636:.
2612::
2604::
2577:.
2557::
2549::
2526:.
2516:1
2499:.
2487::
2464:.
2442::
2415:.
2403::
2397:7
2380:.
2366::
2339:.
2319::
2296:.
2284::
2261:.
2237:.
2233::
2227:4
2210:.
2190::
2164:.
2139:.
2094:.
2054:.
2029:.
2004:.
1979:.
1975::
1801:(
1653:.
1226:)
1224:2
1188:)
1186:2
1150:)
1148:2
1034:)
1032:9
992:)
990:5
961:2
915:)
843:)
841:6
805:)
803:3
763:)
761:1
677:B
672:6
670:B
665:2
663:B
658:1
656:B
489:)
50:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.