445:
generation step. In other words, structure reduction is efficient when structural constraints are provided, and structure assembly is faster without constraints. First, the useless connections are eliminated, and then the substructures are assembled to build structures. Thus, GEN copes with the constraints in a more efficient way by combining these methods. GEN removes the connections creating the forbidden structures, and then the connection matrices are filled based on substructure information. The method does not accept overlaps among substructures. Once the structure is built in the matrix representation, the saturated molecule is stored in the output list. The COCOA method was further improved and a new generator was built, HOUDINI. It relies on two data structures: a square matrix of compounds representing all bonds in a hyper structure is constructed, and second, substructure representation is used to list atom-centred fragments. In the structure generation, HOUDINI maps all the atom-centred fragments onto the hyper structure.
328:. Orderly generation is performed to cope with this exhaustivity. Many assembly algorithms, such as OMG, MOLGEN and Jean-Loup Faulon's structure generator, are orderly generation methods. Jean-Loup Faulon's structure generator relies on equivalence classes over atoms. Atoms with the same interaction type and element are grouped in the same equivalence class. Rather than extending all atoms in a molecule, one atom from each class is connected with other atoms. Similar to the former generator, Julio Peironcely's structure generator, OMG, takes atoms and substructures as inputs and extends the structures using a
310:
2049:
158:'s orderly generation method. Although their reports are from the 1970s, these studies are still the fundamental references for structure generators. In the orderly generation method, specific order-check functions are performed on graph representatives, such as vectors. For example, MOLGEN performs a descending order check while filling rows of adjacency matrices. This descending order check is based on an input valence distribution. The literature classifies generators into two major types: structure assembly and structure reduction. The
459:
223:
the algorithm assembles substructures with overlaps to construct structures. ASSEMBLE overcomes overlapping by including a “neighbouring atom tag”. The generator is purely mathematical and does not involve the interpretation of any spectral data. Spectral data are used for structure scoring and substructure information. Based on the molecular formula, the generator forms bonds between pairs of atoms, and all the extensions are checked against the given constraints. If the process is considered as a
367:. MASS, SMOG and Ivan Bangov's algorithm are good examples in the literature. MASS is a method of mathematical synthesis. First, it builds all incidence matrices for a given molecular formula. The atom valences are then used as the input for matrix generation. The matrices are generated by considering all the possible interactions among atoms with respect to the constraints and valences. The benefit of constructive search algorithms is their low memory usage. SMOG is a successor of MASS.
80:
394:'s research group. As a type of assembly method, building blocks, such as ring systems and atom fragments, are used in the structure generation. Every intermediate structure is extended by adding building blocks in all possible ways. To reduce the number of duplicates, Brendan McKay's canonical path augmentation method is used. To overcome the combinatorial explosion in the generation, applicability domain and ring systems are detected based on inverse
260:
filling in missing bond information. This platform could generate structures with any arbitrary size of molecules; however, molecular formulas with more than 30 heavy atoms are too time consuming for practical applications. This limitation highlighted the need for a new CASE system. SENECA was developed to eliminate the shortcomings of LUCY. To overcome the limitations of the exhaustive approach, SENECA was developed as a
4622:
436:
starts with the hypergraph, and the structures decrease in size at each step. Bonds are deleted based on the substructures. If a substructure is no longer in the hypergraph, the substructure is removed from the constraints. Overlaps in the substructures were also considered due to the hypergraphs. The earliest reduction-based structure generator is COCOA, an exhaustive and
398:
analysis. The applicability domain, or target area, is described based on given biological as well as pharmaceutical activity information from QSPR/QSAR. In that study, monotonically changed descriptors (MCD) are used to describe applicability domains. For every extension in intermediate structures,
444:
The structure generator GEN by Simona
Bohanec combines two tasks: structure assembly and structure reduction. Like COCOA, the initial state of the problem is a hyper structure. Both assembly and reduction methods have advantages and disadvantages, and the GEN tool avoids these disadvantages in the
440:
bond-removal method. Generated fragments are described as atom-centred fragments to optimize storage, comparable to circular fingerprints and atom signatures. Rather than storing structures, only the list of first neighbours of each atom is stored. The main disadvantage of reduction methods is the
222:
Substantial contributions were made by Craig
Shelley and Morton Munk, who published a large number of CASE papers in this field. The first of these papers reported a structure generator, ASSEMBLE. The algorithm is considered one of the earliest assembly methods in the field. As the name indicates,
435:
Unlike these assembly methods, reduction methods make all the bonds between atom pairs, generating a hypergraph. Then, the size of the graph is reduced with respect to the constraints. First, the existence of substructures in the hypergraph is checked. Unlike assembly methods, the generation tree
411:
based on the given reaction rules. Its core algorithm is a breadth-first method, generating structures by applying reaction rules to each source compound. Structure generation and enumeration are performed based on
Brendan McKay's canonical augmentation method. RetroPath 2.0 provides a variety of
335:
OMG generates structures based on the canonical augmentation method from
Brendan McKay's NAUTY package. The algorithm calculates canonical labelling and then extends structures by adding one bond. To keep the extension canonical, canonical bonds are added. Although NAUTY is an efficient tool for
198:
data; second, the assembly of these substructures based on a set of construction rules. Hidetsugu Abe and the other contributors published the first paper on CHEMICS, which is a CASE tool comprising several structure generation methods. The program relies on a predefined non-overlapping fragment
259:
structure elucidation method based on the HMBC data of unknown molecules, and involves an exhaustive 2-step structure generation process where first all combinations of interpretations of HMBC signals are implemented in a connectivity matrix, which is then completed by a deterministic generator
114:
and fragments. These generators are the core of CASE systems. In a generator, the molecular formula is the basic input. If fragments are obtained from the experimental data, they can also be used as inputs to accelerate structure generation. The first structure generators were versions of graph
227:, the first node of the tree is an atom set with substructures if any are provided by the spectral data. By extending the molecule with a bond, an intermediate structure is built. Each intermediate structure can be represented by a node in the generation tree. ASSEMBLE was developed with a
280:
is reached. In the generation, this transformation relies on equations based on Jean-Loup Faulon's rules. LSD (Logic for
Structure Determination) is an important contribution from French scientists. The tool uses spectral data information such as HMBC and
386:
data of an unknown molecule. Compared to many other generators, MOLGEN approaches the problem from different angles. The key feature of MOLGEN is generating structures without building all the intermediate structures and without generating duplicates.
382:. MOLGEN is an orderly generation method. Many different versions of MOLGEN have been developed, and they provide various functions. Based on the users' needs, different types of inputs can be used. For example, MOLGEN-MS allows users to input
601:, due to the computational explosion, many structure generators output only connected chemical graphs. Thus, the connectivity check is one of the mandatory intermediate steps in structure generation because the aim is to generate fully
305:
into fragments, components and segments was performed as an application of integer partitioning. These fragments were then used as building blocks in the structure generator. This structure generator was part of a CASE system, ESESOC.
293:
and, notably, one of the developers of MASS, Mikhail
Elyashberg. COCON is another NMR based structure generator, relying on theoretical data sets for structure generation. Except J-HMBC and J-COSY, all NMR types can be used as inputs.
2559:
Shin-ichi Sasaki; Hidetsugu Abe; Yuji Hirota; Yoshiaki Ishida; Yoshihiro Kudo; Shukichi Ochiai; Keiji Saito; Tohru
Yamasaki (1 November 1978). "CHEMICS-F: A Computer Program System for Structure Elucidation of Organic Compounds".
4091:
Robert C Glem; Andreas Bender; Catrin H Arnby; Lars
Carlsson; Scott Boyer; James Smith (1 March 2006). "Circular fingerprints: flexible molecular descriptors with applications from physical chemistry to ADME".
2975:
K. A. Blinov; M. E. Elyashberg; S. G. Molodtsov; A. J. Williams; E. R. Martirosian (1 April 2001). "An expert system for automated structure elucidation utilizing 1H-1H, 13C-1H and 15N-1H 2D NMR correlations".
238:
The efficiency and exhaustivity of generators are also related to the data structures. Unlike previous methods, AEGIS was a list-processing generator. Compared to adjacency matrices, list data requires less
219:. The secondary and tertiary component sets are built layer-by-layer starting with these primary components. These component sets are represented as vectors and are used as building blocks in the process.
1773:. In molecular graphs, canonical labelling and molecular symmetry detection are implementations of automorphism groups. Although there are well known canonical labelling methods in the field, such as
3461:
Mohammad Mahdi
Jaghoori; Sung-Shik T.Q. Jongmans; Frank de Boer; Julio Peironcely; Jean-Loup Faulon; Theo Reijmers; Thomas Hankemeier (December 2013). "PMG: Multi-core Metabolite Identification".
324:
labelling and duplicate checks. As for many other generators, the tree approach is the skeleton of Jean-Loup Faulon's structure generators. However, considering all possible extensions leads to a
4657:
355:
generation algorithms. In contrast to previous methods, these methods build all the connectivity matrices without building intermediate structures. In these algorithms, canonicity criteria and
441:
massive size of the hypergraphs. Indeed, for molecules with unknown structures, the size of the hyper structure becomes extremely large, resulting in a proportional increase in the run time.
2697:(April 1981). "Applications of artificial intelligence for chemical inference. 37. GENOA: a computer program for structure elucidation utilizing overlapping and alternative substructures".
4647:
146:
After DENDRAL, another mathematical method, MASS, a tool for mathematical synthesis and analysis of molecular structures, was reported. As with CONGEN, the MASS algorithm worked as an
3549:
I. P. Bangov; K. D. Kanev (February 1988). "Computer-assisted structure generation from a gross formula: II. Multiple bond unsaturated and cyclic compounds. Employment of fragments".
2514:
Hidetsugu. Abe; Peter C. Jurs (September 1975). "Automated chemical structure analysis of organic molecules with a molecular structure generator and pattern recognition techniques".
4642:
243:. As no spectral data was interpreted in this system, the user needed to provide substructures as inputs. Structure generators can also vary based on the type of data used, such as
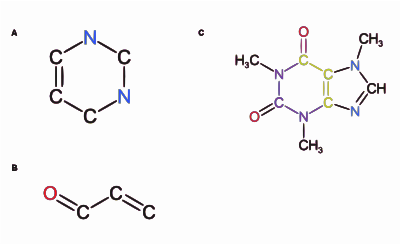
344:, and the developers released PMG (Parallel Molecule Generator). MOLGEN outperforms PMG using only 1 core; however, PMG outperforms MOLGEN by increasing the number of cores to 10.
1502:
589:, and the maximum number of bonds a chemical element can make. For example, carbon's valence is 4. In a chemical graph, an atom is saturated if it reaches its valence. A graph is
854:
3227:
Jean-Loup Faulon; Carla J Churchwell; Donald P Visco (1 May 2003). "The signature molecular descriptor. 2. Enumerating molecules from their extended valence sequences".
1113:
1673:
1582:
1387:
536:
1751:
1619:
1259:
1180:
1077:
231:
to facilitate use. The second version of ASSEMBLE was released in 2000. Another assembly method is GENOA. Compared to ASSEMBLE and many other generators, GENOA is a
3906:(13 July 2017). "druGAN: An Advanced Generative Adversarial Autoencoder Model for de Novo Generation of New Molecules with Desired Molecular Properties in Silico".
1349:
1002:
4135:
Jean-Loup Faulon; Michael J Collins; Robert D Carr (1 March 2004). "The signature molecular descriptor. 4. Canonizing molecules using extended valence sequences".
705:
199:
library. CHEMICS generates different types of component sets ranked from primary to tertiary based on component complexity. The primary set contains atoms, i.e.,
1282:
1771:
1716:
1693:
1639:
1544:
1524:
1427:
1407:
1309:
1224:
1204:
1145:
1042:
1022:
970:
950:
926:
906:
880:
783:
763:
640:
576:
556:
407:
in the generation makes the generation process more efficient. For example, a well-known tool called RetroPath is used for molecular structure enumeration and
316:
Molecular structure generation is explained step by step. Starting from a set of atoms, bonds are added between atom pairs until reaching saturated structures.
399:
the MCDs are updated. The usage of MCDs reduces the search space in the generation process. In the QSPR/QSAR based structure generation, there is the lack of
276:
is calculated to evaluate the structure and its spectral properties. By transforming this structure into another structure, the process continues until the
2887:
Jean-Loup Faulon (January 1996). "Stochastic
Generator of Chemical Structure. 2. Using Simulated Annealing To Search the Space of Constitutional Isomers".
3780:
Baudoin Delépine; Thomas Duigou; Pablo Carbonell; Jean-Loup Faulon (9 December 2017). "RetroPath2.0: A retrosynthesis workflow for metabolic engineers".
248:
2836:(November 2001). "SENECA: A Platform-Independent, Distributed, and Parallel System for Computer-Assisted Structure Elucidation in Organic Chemistry".
1319:
of the graph. An automorphism permutes the vertices of a graph; in other words, it maps a graph onto itself. This action is edge-vertex preserving. If
3463:
3184:
Jean-Loup Faulon (1 September 1994). "Stochastic Generator of Chemical Structure. 1. Application to the Structure Elucidation of Large Molecules".
395:
48:
2744:
2650:
3278:
Julio E Peironcely; Miguel Rojas-Chertó; Davide Fichera; Theo Reijmers; Leon Coulier; Jean-Loup Faulon; Thomas Hankemeier (17 September 2012).
320:
A series of stochastic generators was reported by Jean-Loup Faulon. The software, MOLSIG, was integrated into this stochastic generator for
2036:
4652:
3506:
M. S. Molchanova; V. V. Shcherbukhin; N. S. Zefirov (January 1996). "Computer Generation of Molecular Structures by the SMOG Program".
4040:
Christie BD; Munk ME (1 May 1988). "Structure generation by reduction: a new strategy for computer-assisted structure elucidation".
2060:
3594:"MOLGEN-MS: Evaluation of low resolution electron impact mass spectra with MS classification and exhaustive structure generation"
60:
2332:
Karina A. Gulyaeva; Irina L. Artemieva (2020). "The Ontological Approach in Organic Chemistry Intelligent System Development".
3141:
Junfeng Hao; Lu Xu; Changyu Hu (October 2000). "Expert system for elucidation of structures of organic compounds (ESESOC)".
489:
2742:
H.J. Luinge; J.H. Van Der Maas (June 1990). "AEGIS, an algorithm for the exhaustive generation of irredundant structures".
131:
to search for life on Mars. CONGEN dealt well with overlaps in substructures. The overlaps among substructures rather than
4518:"MAYGEN: an open-source chemical structure generator for constitutional isomers based on the orderly generation principle"
115:
generators modified for chemical purposes. One of the first structure generators was CONGEN, originally developed for the
602:
1839:
2648:
Martin Badertscher; Andrew Korytko; Klaus-Peter Schulz; et al. (May 2000). "Assemble 2.0: a structure generator".
351:
method, such as Igor Faradjev's algorithm, and an additional solution to memory problems. Branch-and-bound methods are
194:-based structure generator. The algorithm had two steps: first, the prediction of the substructure from low-resolution
4186:
Simona Bohanec (1 May 1995). "Structure Generation by the Combination of Structure Reduction and Structure Assembly".
3347:
Jean Loup Faulon (1 July 1992). "On using graph-equivalent classes for the structure elucidation of large molecules".
2371:
V.V. Serov; M.E. Elyashberg; L.A. Gribov (April 1976). "Mathematical synthesis and analysis of molecular structures".
3096:
Chang-Yu Hu; Lu Xu (November 1994). "Principles for structure generation of organic isomers from molecular formula".
2289:
1774:
163:
4626:
2699:
1777:
and ALATIS, NAUTY is a commonly used software package for automorphism group calculations and canonical labelling.
3782:
3395:
2603:
Craig A. Shelley; Morton E. Munk (November 1981). "Case, a computer model of the structure elucidation process".
235:
substructure search-based algorithm, and it assembles different substructures by also considering the overlaps.
2207:(1 January 1987). "Prediction of the folding of short polypeptide segments by uniform conformational sampling".
2516:
2373:
290:
3390:
370:
Unlike previous methods, MOLGEN is the only maintained efficient generic structure generator, developed as a
63:(CASE). CASE systems again have regained interest for the structure elucidation of unknowns in computational
4284:
G. Massiot; J. M. Nuzillard (July 1992). "Computer-assisted elucidation of structures of natural products".
3460:
437:
103:
3447:
1434:
745:
The second line of this table shows a permutation of the first line. The multiplication of permutations,
590:
289:. A well-known commercial CASE system, StrucEluc, also features a NMR based generator. This tool is from
4382:
3839:
3284:
3033:
2079:
379:
286:
285:
data to generate all possible structures. LSD is an open source structure generator released under the
2278:
794:
4455:
2287:(June 1993). "DENDRAL: A case study of the first expert system for scientific hypothesis formation".
420:
175:
585:
of a vertex is its number of connections. In a chemical graph, the maximum degree of an atom is its
4662:
1315:, called blocks of the set. The set of permutations preserving the set system is used to build the
273:
159:
2790:(20 September 1996). "LUCY—A Program for Structure Elucidation from NMR Correlation Experiments".
336:
graph canonical labelling, OMG is approximately 2000 times slower than MOLGEN. The problem is the
3908:
2444:
404:
325:
56:
3835:"Molecular structures enumeration and virtual screening in the chemical space with RetroPath2.0"
3277:
2064:
4286:
3670:(26 November 2014). "Ring-System-Based Exhaustive Structure Generation for Inverse-QSPR/QSAR".
3098:
2692:
2605:
2497:"Algorithms for group actions: Homomorphism principle and orderly generation applied to graphs"
232:
28:
2647:
1082:
3969:
3729:
3672:
2932:
2209:
1644:
1549:
1354:
582:
503:
481:
337:
216:
68:
4444:
3779:
3615:(14 June 2016). "Ring system-based chemical graph generation for de novo molecular design".
1721:
1589:
1229:
1150:
1052:
309:
4376:
Stephen R Heller; Alan McNaught; Igor Pletnev; Stephen Stein; Dmitrii Tchekhovskoi (2015).
3958:
2157:
1322:
1124:
975:
883:
786:
371:
352:
329:
256:
240:
3424:
2930:
Jean-Marc Nuzillard; Massiot Georges (January 1991). "Logic for structure determination".
2473:
681:
332:
method. This tree extension terminates when all the branches reach saturated structures.
8:
3832:
2974:
2833:
2787:
2070:
586:
485:
277:
265:
224:
191:
107:
2161:
1264:
190:
in the molecule. One of the earliest attempts was made by Hidetsugu Abe in 1975 using a
106:
generation problems. Molecular structures are graphs with chemical constraints such as
4595:
4568:
4544:
4517:
4485:
4450:
4414:
4377:
4009:
3978:
3964:
3871:
3834:
3316:
3279:
3065:
3028:
2558:
2261:
2180:
2145:
2111:
2074:
1756:
1701:
1678:
1624:
1529:
1509:
1412:
1392:
1316:
1294:
1209:
1189:
1130:
1027:
1007:
955:
935:
911:
891:
865:
768:
748:
625:
561:
541:
400:
360:
341:
321:
94:. The overlap of these substructures is highlighted in green in the caffeine structure
24:
4375:
4090:
3921:
3901:
2945:
2663:
2618:
182:
connected to consider all possible extensions. If substructures are obtained from the
4600:
4549:
4490:
4472:
4419:
4401:
4350:
4303:
4258:
4250:
4203:
4160:
4152:
4109:
4101:
4065:
4057:
4014:
3996:
3933:
3925:
3876:
3858:
3807:
3799:
3754:
3746:
3697:
3689:
3640:
3632:
3593:
3566:
3523:
3480:
3412:
3364:
3321:
3303:
3252:
3244:
3201:
3158:
3115:
3111:
3070:
3052:
3001:
2993:
2949:
2904:
2861:
2853:
2807:
2761:
2757:
2716:
2667:
2622:
2577:
2533:
2496:
2461:
2390:
2386:
2345:
2306:
2302:
2280:
2234:
2226:
2185:
2116:
2098:
1285:
1183:
408:
383:
298:
269:
183:
120:
52:
3833:
Mathilde Koch; Thomas Duigou; Pablo Carbonell; Jean-Loup Faulon (19 December 2017).
2048:
186:, the generation starts with these substructures. These substructures provide known
4590:
4580:
4539:
4529:
4498:
4480:
4464:
4427:
4409:
4391:
4358:
4342:
4311:
4295:
4266:
4242:
4211:
4195:
4168:
4144:
4117:
4073:
4049:
4022:
4004:
3988:
3941:
3917:
3903:
3884:
3866:
3848:
3815:
3791:
3762:
3738:
3705:
3681:
3648:
3624:
3574:
3558:
3531:
3515:
3488:
3472:
3428:
3420:
3404:
3372:
3356:
3329:
3311:
3293:
3260:
3236:
3209:
3193:
3166:
3150:
3123:
3107:
3078:
3060:
3042:
3009:
2985:
2957:
2941:
2912:
2896:
2869:
2845:
2815:
2799:
2769:
2753:
2724:
2708:
2675:
2659:
2630:
2614:
2585:
2569:
2541:
2525:
2477:
2469:
2453:
2435:
2398:
2382:
2353:
2337:
2314:
2298:
2284:
2242:
2218:
2175:
2165:
2124:
2106:
2088:
1959:
1116:
886:
348:
302:
151:
147:
36:
4230:
2170:
2093:
493:
136:
124:
44:
32:
4451:"Unique identifiers for small molecules enable rigorous labeling of their atoms"
3795:
3476:
3226:
2341:
488:
represent atoms and bonds, respectively. The bond order corresponds to the edge
458:
419:
Besides these mathematical structure generation methods, the implementations of
4585:
4534:
4333:(March 1999). "Combinatorial algorithms: generation, enumeration, and search".
3724:
3667:
3612:
3505:
2439:
2204:
2055:
593:
if there is at least one path between each pair of vertices. Although chemical
497:
391:
264:
method to find optimal solutions. The systems comprise two stochastic methods:
228:
155:
140:
4396:
4134:
3853:
3628:
3408:
1869:
4636:
4567:
McKay, Brendan D.; Yirik, Mehmet Aziz; Steinbeck, Christoph (December 2022).
4476:
4405:
4354:
4307:
4254:
4207:
4156:
4105:
4061:
4000:
3960:
3929:
3862:
3803:
3750:
3693:
3636:
3570:
3527:
3484:
3416:
3368:
3307:
3248:
3205:
3162:
3119:
3056:
2997:
2953:
2908:
2857:
2811:
2765:
2720:
2694:
2671:
2626:
2581:
2537:
2465:
2394:
2349:
2310:
2262:"DENDRAL-a computer program for generating and filtering chemical structures"
2230:
2102:
375:
187:
179:
4362:
3722:
3578:
3492:
3170:
2357:
2128:
4604:
4553:
4516:
Yirik, Mehmet Aziz; Sorokina, Maria; Steinbeck, Christoph (December 2021).
4494:
4446:
4423:
4330:
4299:
4262:
4164:
4113:
4018:
3992:
3937:
3880:
3811:
3758:
3742:
3727:(1 January 2010). "Exhaustive Structure Generation for Inverse-QSPR/QSAR".
3701:
3685:
3665:
3644:
3610:
3325:
3298:
3256:
3074:
3047:
3005:
2865:
2803:
2457:
2189:
2120:
859:
413:
364:
195:
64:
4502:
4468:
4431:
4346:
4315:
4270:
4215:
4172:
4121:
4077:
4069:
4026:
3945:
3888:
3819:
3766:
3709:
3652:
3535:
3432:
3376:
3333:
3264:
3213:
3127:
3082:
3013:
2989:
2961:
2916:
2873:
2819:
2773:
2728:
2679:
2634:
2589:
2545:
2481:
2442:(1979). "Orderly algorithms for generating restricted classes of graphs".
2416:
Faradzev, IA (1978). "Constructive enumeration of combinatorial objects".
2402:
2318:
2246:
2238:
2222:
1995:
614:
424:
356:
261:
79:
40:
4228:
4199:
4053:
3360:
3197:
2712:
2573:
2529:
2017:
3562:
3154:
2693:
Raymond E. Carhart; Dennis H. Smith; Neil A. B. Gray; James G. Nourse;
1289:
111:
4328:
4246:
4148:
3519:
3240:
2900:
2849:
2501:
DIMACS Series in Discrete Mathematics and Theoretical Computer Science
1977:
464:
150:
generator. Many mathematical generators are descendants of efficient
3393:; Adolfo Piperno (January 2014). "Practical graph isomorphism, II".
2370:
2331:
1803:
4229:
A. Korytko; K-P Schulz; M. S. Madison; Munk ME (1 September 2003).
3983:
3965:"Application of Generative Autoencoder in de Novo Molecular Design"
3959:
Thomas Blaschke; Marcus Olivecrona; Ola Engkvist; Jürgen Bajorath;
1941:
1785:
The available software packages and their links are listed below.
605:
molecules. A molecule is saturated if all its atoms are saturated.
204:
31:. The development of such software packages is a research topic of
2929:
1893:
2434:
1127:
of a group is the number of elements in the group. Let us assume
617:
is a rearrangement of these elements. An example is given below:
598:
594:
116:
4231:"HOUDINI: A New Approach to Computer-Based Structure Generation"
1925:
1147:
is a set of integers. Under the function composition operation,
4621:
1312:
412:
workflows such as isomer transformation, enumeration, QSAR and
212:
208:
200:
4658:
Knowledge articles published in peer-reviewed literature (J2W)
3902:
Artur Kadurin; Sergey Nikolenko; Kuzma Khrabrov; Alex Aliper;
3389:
2602:
35:. Chemical graph generators are used in areas such as virtual
23:
is a software package to generate computer representations of
858:
The combination of two permutations is also a permutation. A
2741:
2513:
2144:
Yirik, Mehmet Aziz; Steinbeck, Christoph (5 January 2021).
2053:
This article was adapted from the following source under a
282:
244:
174:
The generation process starts with a set of atoms from the
132:
128:
4648:
Knowledge articles published in PLOS Computational Biology
3499:
3271:
3140:
2494:
4283:
3591:
3029:"Theoretical NMR correlations based Structure Discussion"
1780:
252:
4643:
Knowledge articles published in peer-reviewed literature
1909:
3548:
1823:
608:
340:
of all the intermediate structures. OMG has since been
4515:
3952:
2735:
2641:
2202:
119:
project, the first artificial intelligence project in
4235:
Journal of Chemical Information and Computer Sciences
4188:
Journal of Chemical Information and Computer Sciences
4137:
Journal of Chemical Information and Computer Sciences
4042:
Journal of Chemical Information and Computer Sciences
3592:
Kerber, A; Laue, R; Meringer, M; Varmuza, K. (2001).
3508:
Journal of Chemical Information and Computer Sciences
3349:
Journal of Chemical Information and Computer Sciences
3229:
Journal of Chemical Information and Computer Sciences
3186:
Journal of Chemical Information and Computer Sciences
2889:
Journal of Chemical Information and Computer Sciences
2838:
Journal of Chemical Information and Computer Sciences
2562:
Journal of Chemical Information and Computer Sciences
2495:
Grüner, T; Laue, R; Meringer, M; Bayreuth, U (1997).
1759:
1724:
1704:
1681:
1647:
1627:
1592:
1552:
1532:
1512:
1437:
1415:
1395:
1357:
1325:
1297:
1267:
1232:
1212:
1192:
1186:, the set of all permutations over X. If the size of
1153:
1133:
1085:
1055:
1030:
1010:
978:
958:
938:
914:
894:
868:
797:
771:
751:
684:
628:
564:
544:
506:
301:-based structure generator. The decomposition of the
143:
calculations were performed for duplicate detection.
4569:"Surge: a fast open-source chemical graph generator"
4378:"InChI, the IUPAC International Chemical Identifier"
4277:
3895:
3454:
3220:
2968:
2923:
2196:
272:. First, a random structure is generated; then, its
86:
Two substructures of a caffeine molecule are given,
4566:
2832:
2786:
4369:
4039:
3773:
2364:
1765:
1745:
1710:
1687:
1667:
1633:
1613:
1576:
1538:
1518:
1496:
1421:
1401:
1381:
1343:
1303:
1276:
1253:
1218:
1198:
1174:
1139:
1107:
1071:
1036:
1016:
996:
964:
944:
920:
900:
874:
848:
777:
757:
699:
634:
578:is the set of edges, which represents the bonds.
570:
550:
530:
480:In a graph representing a chemical structure, the
390:In the field, the studies recent to 2021 are from
135:were used as the building blocks. For the case of
4438:
4322:
4222:
4128:
4634:
4033:
3716:
3464:Electronic Notes in Theoretical Computer Science
3383:
3346:
3183:
3095:
2886:
2686:
3659:
3604:
3542:
2745:Chemometrics and Intelligent Laboratory Systems
2651:Chemometrics and Intelligent Laboratory Systems
2596:
2428:
427:models, are the novel directions of the field.
4185:
4084:
3826:
2978:Fresenius' Journal of Analytical Chemistry
2143:
102:Molecular structure generation is a branch of
3026:
2826:
2780:
2552:
2334:Advances in Intelligent Systems and Computing
3617:Journal of Computer - Aided Molecular Design
2037:Simplified molecular-input line-entry system
16:Software for visualizing chemical structures
4179:
2507:
2325:
1049:For each element of G, there is an element
3340:
3177:
3020:
2880:
2272:
2259:
47:with specified properties, called inverse
4594:
4584:
4543:
4533:
4484:
4413:
4395:
4094:IDrugs: the Investigational Drugs Journal
4008:
3982:
3870:
3852:
3315:
3297:
3134:
3064:
3046:
2179:
2169:
2137:
2110:
2092:
558:is the set of vertices, i.e., atoms, and
123:. DENDRAL was developed as a part of the
3448:"The Benchmark for Structure Generators"
3089:
2415:
882:, is a set of elements together with an
457:
308:
78:
2792:Angewandte Chemie International Edition
538:is described as a chemical graph where
166:are the criteria used for comparison.
61:computer-assisted structure elucidation
4635:
2279:Robert K. Lindsay; Bruce G. Buchanan;
1781:List of available structure generators
597:are one of the main interests of many
430:
403:of the generated structures. Usage of
84:Overlapping substructures of caffeine.
3445:
3143:Science in China. Series B: Chemistry
1753:, is the set of all automorphisms on
476:Graph representation of the molecule.
448:
347:A constructive search algorithm is a
169:
1497:{\displaystyle a({u,v})=(a(u),a(v))}
609:Symmetry groups for molecular graphs
4449:; Hamid R Eghbalnia (23 May 2017).
500:. A vertex and edge-labelled graph
178:. In structure assembly, atoms are
13:
3723:Tomoyuki Miyao; Masamoto Arakawa;
1698:The automorphism group of a graph
928:such that the following are true:
472:Molecular structure of serotonin.
453:
14:
4674:
4653:Externally peer reviewed articles
4614:
4445:Hesam Dashti; William M Westler;
3922:10.1021/ACS.MOLPHARMACEUT.7B00346
3666:Tomoyuki Miyao; Hiromasa Kaneko;
3611:Tomoyuki Miyao; Hiromasa Kaneko;
3551:Journal of Mathematical Chemistry
4620:
2700:The Journal of Organic Chemistry
2047:
1546:is an automorphism of the graph
849:{\displaystyle (a*b)(x)=a(b(x))}
314:Breadth-first search generation.
4560:
4509:
3585:
3439:
3396:Journal of Symbolic Computation
498:vertex and edge-labelled graphs
297:In 1994, Hu and Xu reported an
154:methods from Igor Faradjev and
3280:"OMG: Open Molecule Generator"
3027:Jochen Junker (28 July 2011).
2488:
2409:
2374:Journal of Molecular Structure
2253:
1740:
1734:
1661:
1649:
1608:
1596:
1571:
1559:
1491:
1488:
1482:
1473:
1467:
1461:
1455:
1441:
1376:
1364:
1338:
1326:
1248:
1242:
1169:
1163:
843:
840:
834:
828:
819:
813:
810:
798:
694:
688:
525:
513:
1:
3598:Advances in Mass Spectrometry
2946:10.1016/S0040-4020(01)80878-4
2664:10.1016/S0169-7439(00)00056-3
2619:10.1016/S0003-2670(01)95416-9
2260:Sutherland, G. (1967-02-15).
2042:
3112:10.1016/0003-2670(94)90044-2
2758:10.1016/0169-7439(90)80131-O
2387:10.1016/0022-2860(76)80018-X
2303:10.1016/0004-3702(93)90068-M
2171:10.1371/journal.pcbi.1008504
2094:10.1371/JOURNAL.PCBI.1008504
463:Graph representation of the
287:General Public License (GPL)
7:
3796:10.1016/J.YMBEN.2017.12.002
3477:10.1016/J.ENTCS.2013.11.005
2342:10.1007/978-981-32-9343-4_7
2146:"Chemical graph generators"
2075:"Chemical graph generators"
2030:
10:
4679:
4586:10.1186/s13321-022-00604-9
4573:Journal of Cheminformatics
4535:10.1186/s13321-021-00529-9
4522:Journal of Cheminformatics
4383:Journal of Cheminformatics
3840:Journal of Cheminformatics
3285:Journal of Cheminformatics
3034:Journal of Cheminformatics
2840:(in English and English).
2150:PLOS Computational Biology
2080:PLOS Computational Biology
380:computational group theory
74:
4397:10.1186/S13321-015-0068-4
3854:10.1186/S13321-017-0252-9
3629:10.1007/S10822-016-9916-1
3409:10.1016/J.JSC.2013.09.003
1351:is an edge of the graph,
613:For a set of elements, a
2266:DEPT OF COMPUTER SCIENCE
1108:{\displaystyle g*g^{-1}}
53:organic synthesis design
21:chemical graph generator
3909:Molecular Pharmaceutics
2445:Journal of Graph Theory
2290:Artificial Intelligence
1668:{\displaystyle {(u,v)}}
1577:{\displaystyle G=(E,V)}
1382:{\displaystyle G=(V,E)}
531:{\displaystyle G=(V,E)}
374:platform by a group of
326:combinatorial explosion
229:user-friendly interface
4329:Donald Lawson Kreher;
4300:10.1002/PCA.2800030403
4287:Phytochemical Analysis
3993:10.1002/MINF.201700123
3743:10.1002/MINF.200900038
3686:10.1002/MINF.201400072
3299:10.1186/1758-2946-4-21
3099:Analytica Chimica Acta
3048:10.1186/1758-2946-3-27
2804:10.1002/ANIE.199619841
2606:Analytica Chimica Acta
2458:10.1002/JGT.3190030210
2418:Colloq. Internat. CNRS
2268:. Stanford University.
1767:
1747:
1746:{\displaystyle Aut(G)}
1712:
1689:
1669:
1635:
1615:
1614:{\displaystyle a(u,v)}
1578:
1540:
1520:
1498:
1423:
1403:
1383:
1345:
1305:
1278:
1255:
1254:{\displaystyle Sym(X)}
1220:
1200:
1176:
1175:{\displaystyle Sym(X)}
1141:
1109:
1073:
1072:{\displaystyle g^{-1}}
1038:
1018:
998:
966:
946:
922:
902:
876:
850:
779:
759:
701:
636:
572:
552:
532:
477:
317:
160:algorithmic complexity
99:
4469:10.1038/SDATA.2017.73
4347:10.1145/309739.309744
3970:Molecular Informatics
3783:Metabolic Engineering
3730:Molecular Informatics
3673:Molecular Informatics
2990:10.1007/S002160100757
2223:10.1002/BIP.360260114
1842:.softwarepreservation
1835:DENDRAL CONGEN+GENOA
1768:
1748:
1713:
1690:
1670:
1636:
1616:
1579:
1541:
1521:
1499:
1424:
1404:
1384:
1346:
1344:{\displaystyle (u,v)}
1306:
1288:systems consist of a
1279:
1256:
1221:
1201:
1177:
1142:
1110:
1074:
1039:
1019:
999:
997:{\displaystyle g*I=g}
967:
947:
923:
903:
877:
851:
780:
760:
702:
637:
581:In graph theory, the
573:
553:
533:
461:
423:, such as generative
378:as an application of
312:
82:
69:computational biology
4629:at Wikimedia Commons
3963:(13 December 2017).
3450:– via Blogger.
3446:Yirik, M.A. (2020).
2517:Analytical Chemistry
2281:Edward A. Feigenbaum
1757:
1722:
1702:
1679:
1645:
1625:
1590:
1550:
1530:
1510:
1435:
1413:
1409:is a permutation of
1393:
1355:
1323:
1295:
1265:
1230:
1226:, then the order of
1210:
1190:
1151:
1131:
1083:
1053:
1028:
1008:
976:
956:
936:
932:There is an element
912:
892:
866:
795:
787:function composition
769:
749:
700:{\displaystyle f(x)}
682:
626:
562:
542:
504:
405:retrosynthesis paths
359:checks are based on
330:breadth-first search
67:, a current area of
27:adhering to certain
4200:10.1021/CI00025A017
4054:10.1021/CI00058A009
3600:. pp. 939–940.
3361:10.1021/CI00008A013
3198:10.1021/CI00021A031
2834:Christoph Steinbeck
2788:Christoph Steinbeck
2713:10.1021/JO00321A037
2574:10.1021/CI60016A007
2530:10.1021/AC60361A007
2436:Charles J. Colbourn
2162:2021PLSCB..17E8504Y
2071:Christoph Steinbeck
2069:Mehmet Aziz Yirik;
1004:, for all elements
789:, as shown below.
492:, and as a result,
431:Structure reduction
361:automorphism groups
266:simulated annealing
192:pattern recognition
29:boundary conditions
25:chemical structures
4331:Douglas R. Stinson
3680:(11–12): 764–778.
3563:10.1007/BF01166467
3155:10.1007/BF02969496
2503:. pp. 113–22.
2073:(5 January 2021).
2018:structuregenerator
1763:
1743:
1708:
1685:
1665:
1631:
1611:
1574:
1536:
1516:
1494:
1419:
1399:
1379:
1341:
1301:
1277:{\displaystyle n!}
1274:
1251:
1216:
1196:
1172:
1137:
1105:
1069:
1034:
1014:
994:
962:
942:
918:
898:
872:
846:
785:, is defined as a
775:
755:
697:
632:
568:
548:
528:
478:
449:Mathematical basis
363:from mathematical
318:
270:genetic algorithms
170:Structure assembly
100:
59:or in systems for
4625:Media related to
4247:10.1021/CI034057R
4149:10.1021/CI0341823
3520:10.1021/CI950393Z
3241:10.1021/CI020346O
2940:(22): 3655–3664.
2901:10.1021/CI950179A
2850:10.1021/CI000407N
2798:(17): 1984–1986.
2524:(11): 1829–1835.
2028:
2027:
1766:{\displaystyle V}
1711:{\displaystyle G}
1688:{\displaystyle E}
1675:is an element of
1634:{\displaystyle E}
1621:is an element of
1539:{\displaystyle V}
1519:{\displaystyle a}
1422:{\displaystyle V}
1402:{\displaystyle a}
1304:{\displaystyle X}
1219:{\displaystyle n}
1199:{\displaystyle X}
1140:{\displaystyle X}
1037:{\displaystyle G}
1017:{\displaystyle g}
965:{\displaystyle G}
945:{\displaystyle I}
921:{\displaystyle G}
901:{\displaystyle *}
875:{\displaystyle G}
778:{\displaystyle b}
758:{\displaystyle a}
743:
742:
635:{\displaystyle x}
571:{\displaystyle E}
551:{\displaystyle V}
409:virtual screening
384:mass spectrometry
303:molecular formula
299:integer partition
255:data. LUCY is an
184:experimental data
176:molecular formula
121:organic chemistry
112:bond multiplicity
4670:
4624:
4609:
4608:
4598:
4588:
4564:
4558:
4557:
4547:
4537:
4513:
4507:
4506:
4488:
4442:
4436:
4435:
4417:
4399:
4373:
4367:
4366:
4326:
4320:
4319:
4281:
4275:
4274:
4241:(5): 1434–1446.
4226:
4220:
4219:
4183:
4177:
4176:
4132:
4126:
4125:
4088:
4082:
4081:
4037:
4031:
4030:
4012:
3986:
3977:(1–2): 1700123.
3956:
3950:
3949:
3916:(9): 3098–3104.
3904:Alex Zhavoronkov
3899:
3893:
3892:
3874:
3856:
3830:
3824:
3823:
3777:
3771:
3770:
3737:(1–2): 111–125.
3720:
3714:
3713:
3663:
3657:
3656:
3608:
3602:
3601:
3589:
3583:
3582:
3546:
3540:
3539:
3503:
3497:
3496:
3458:
3452:
3451:
3443:
3437:
3436:
3391:Brendan D. McKay
3387:
3381:
3380:
3344:
3338:
3337:
3319:
3301:
3275:
3269:
3268:
3224:
3218:
3217:
3192:(5): 1204–1218.
3181:
3175:
3174:
3138:
3132:
3131:
3093:
3087:
3086:
3068:
3050:
3024:
3018:
3017:
2984:(7–8): 709–714.
2972:
2966:
2965:
2927:
2921:
2920:
2884:
2878:
2877:
2844:(6): 1500–1507.
2830:
2824:
2823:
2784:
2778:
2777:
2739:
2733:
2732:
2707:(8): 1708–1718.
2690:
2684:
2683:
2645:
2639:
2638:
2600:
2594:
2593:
2556:
2550:
2549:
2511:
2505:
2504:
2492:
2486:
2485:
2432:
2426:
2425:
2413:
2407:
2406:
2368:
2362:
2361:
2329:
2323:
2322:
2285:Joshua Lederberg
2276:
2270:
2269:
2257:
2251:
2250:
2200:
2194:
2193:
2183:
2173:
2141:
2133:
2132:
2114:
2096:
2065:reviewer reports
2058:
2051:
2024:
2021:
2019:
2008:
2005:
2003:
2001:
1999:
1997:
1986:
1983:
1981:
1979:
1968:
1966:/pmgcoordination
1965:
1963:
1961:
1950:
1947:
1945:
1943:
1932:
1929:
1927:
1916:
1913:
1911:
1900:
1897:
1895:
1884:
1881:
1879:
1877:
1875:
1873:
1871:
1860:
1857:
1855:
1853:
1851:
1849:
1847:
1845:
1843:
1841:
1830:
1827:
1825:
1814:
1811:
1809:
1807:
1805:
1788:
1787:
1772:
1770:
1769:
1764:
1752:
1750:
1749:
1744:
1717:
1715:
1714:
1709:
1694:
1692:
1691:
1686:
1674:
1672:
1671:
1666:
1664:
1640:
1638:
1637:
1632:
1620:
1618:
1617:
1612:
1583:
1581:
1580:
1575:
1545:
1543:
1542:
1537:
1525:
1523:
1522:
1517:
1503:
1501:
1500:
1495:
1454:
1428:
1426:
1425:
1420:
1408:
1406:
1405:
1400:
1388:
1386:
1385:
1380:
1350:
1348:
1347:
1342:
1310:
1308:
1307:
1302:
1283:
1281:
1280:
1275:
1260:
1258:
1257:
1252:
1225:
1223:
1222:
1217:
1205:
1203:
1202:
1197:
1181:
1179:
1178:
1173:
1146:
1144:
1143:
1138:
1117:identity element
1115:is equal to the
1114:
1112:
1111:
1106:
1104:
1103:
1078:
1076:
1075:
1070:
1068:
1067:
1043:
1041:
1040:
1035:
1023:
1021:
1020:
1015:
1003:
1001:
1000:
995:
971:
969:
968:
963:
951:
949:
948:
943:
927:
925:
924:
919:
907:
905:
904:
899:
887:binary operation
881:
879:
878:
873:
855:
853:
852:
847:
784:
782:
781:
776:
764:
762:
761:
756:
706:
704:
703:
698:
641:
639:
638:
633:
620:
619:
577:
575:
574:
569:
557:
555:
554:
549:
537:
535:
534:
529:
401:synthesizability
349:branch-and-bound
180:combinatorically
152:branch-and-bound
148:adjacency matrix
127:launched by the
51:, as well as in
45:molecular design
4678:
4677:
4673:
4672:
4671:
4669:
4668:
4667:
4663:Cheminformatics
4633:
4632:
4627:Chemical graphs
4617:
4612:
4565:
4561:
4514:
4510:
4456:Scientific Data
4443:
4439:
4374:
4370:
4335:ACM SIGACT News
4327:
4323:
4282:
4278:
4227:
4223:
4184:
4180:
4133:
4129:
4089:
4085:
4038:
4034:
3957:
3953:
3900:
3896:
3831:
3827:
3778:
3774:
3721:
3717:
3664:
3660:
3609:
3605:
3590:
3586:
3547:
3543:
3504:
3500:
3459:
3455:
3444:
3440:
3388:
3384:
3345:
3341:
3276:
3272:
3225:
3221:
3182:
3178:
3139:
3135:
3094:
3090:
3025:
3021:
2973:
2969:
2928:
2924:
2885:
2881:
2831:
2827:
2785:
2781:
2740:
2736:
2691:
2687:
2646:
2642:
2601:
2597:
2557:
2553:
2512:
2508:
2493:
2489:
2433:
2429:
2414:
2410:
2369:
2365:
2330:
2326:
2277:
2273:
2258:
2254:
2203:Bruccoleri RE;
2201:
2197:
2156:(1): e1008504.
2142:
2138:
2087:(1): e1008504.
2068:
2054:
2052:
2045:
2033:
2016:
1994:
1976:
1958:
1940:
1924:
1908:
1892:
1868:
1852:/DENDRAL-CONGEN
1838:
1822:
1802:
1783:
1758:
1755:
1754:
1723:
1720:
1719:
1703:
1700:
1699:
1696:
1680:
1677:
1676:
1648:
1646:
1643:
1642:
1626:
1623:
1622:
1591:
1588:
1587:
1551:
1548:
1547:
1531:
1528:
1527:
1511:
1508:
1507:
1504:
1444:
1436:
1433:
1432:
1414:
1411:
1410:
1394:
1391:
1390:
1356:
1353:
1352:
1324:
1321:
1320:
1296:
1293:
1292:
1266:
1263:
1262:
1231:
1228:
1227:
1211:
1208:
1207:
1191:
1188:
1187:
1152:
1149:
1148:
1132:
1129:
1128:
1096:
1092:
1084:
1081:
1080:
1060:
1056:
1054:
1051:
1050:
1029:
1026:
1025:
1009:
1006:
1005:
977:
974:
973:
957:
954:
953:
937:
934:
933:
913:
910:
909:
893:
890:
889:
867:
864:
863:
856:
796:
793:
792:
770:
767:
766:
750:
747:
746:
683:
680:
679:
627:
624:
623:
611:
563:
560:
559:
543:
540:
539:
505:
502:
501:
494:chemical graphs
456:
454:Chemical graphs
451:
433:
421:neural networks
172:
125:Mariner program
77:
33:cheminformatics
17:
12:
11:
5:
4676:
4666:
4665:
4660:
4655:
4650:
4645:
4631:
4630:
4616:
4615:External links
4613:
4611:
4610:
4559:
4508:
4447:John L Markley
4437:
4368:
4321:
4294:(4): 153–159.
4276:
4221:
4194:(3): 494–503.
4178:
4143:(2): 427–436.
4127:
4100:(3): 199–204.
4083:
4032:
3951:
3894:
3825:
3772:
3725:Kimito Funatsu
3715:
3668:Kimito Funatsu
3658:
3623:(5): 425–446.
3613:Kimito Funatsu
3603:
3584:
3541:
3514:(4): 888–899.
3498:
3453:
3438:
3382:
3355:(4): 338–348.
3339:
3270:
3235:(3): 721–734.
3219:
3176:
3149:(5): 503–515.
3133:
3088:
3019:
2967:
2922:
2895:(4): 731–740.
2879:
2825:
2779:
2752:(2): 157–165.
2734:
2685:
2640:
2613:(4): 507–516.
2595:
2568:(4): 211–222.
2551:
2506:
2487:
2452:(2): 187–195.
2440:Ronald C. Read
2427:
2408:
2381:(2): 381–397.
2363:
2324:
2297:(2): 209–261.
2271:
2252:
2217:(1): 137–168.
2195:
2135:
2044:
2041:
2040:
2039:
2032:
2029:
2026:
2025:
2014:
2010:
2009:
1992:
1988:
1987:
1974:
1970:
1969:
1956:
1952:
1951:
1938:
1934:
1933:
1922:
1918:
1917:
1906:
1902:
1901:
1890:
1886:
1885:
1866:
1862:
1861:
1836:
1832:
1831:
1820:
1816:
1815:
1800:
1796:
1795:
1792:
1782:
1779:
1762:
1742:
1739:
1736:
1733:
1730:
1727:
1707:
1684:
1663:
1660:
1657:
1654:
1651:
1630:
1610:
1607:
1604:
1601:
1598:
1595:
1586:
1573:
1570:
1567:
1564:
1561:
1558:
1555:
1535:
1515:
1506:A permutation
1493:
1490:
1487:
1484:
1481:
1478:
1475:
1472:
1469:
1466:
1463:
1460:
1457:
1453:
1450:
1447:
1443:
1440:
1431:
1418:
1398:
1378:
1375:
1372:
1369:
1366:
1363:
1360:
1340:
1337:
1334:
1331:
1328:
1300:
1273:
1270:
1250:
1247:
1244:
1241:
1238:
1235:
1215:
1195:
1184:symmetry group
1171:
1168:
1165:
1162:
1159:
1156:
1136:
1121:
1120:
1102:
1099:
1095:
1091:
1088:
1066:
1063:
1059:
1046:
1045:
1033:
1013:
993:
990:
987:
984:
981:
961:
941:
917:
897:
871:
845:
842:
839:
836:
833:
830:
827:
824:
821:
818:
815:
812:
809:
806:
803:
800:
791:
774:
754:
741:
740:
737:
734:
731:
728:
725:
722:
719:
716:
713:
710:
707:
696:
693:
690:
687:
676:
675:
672:
669:
666:
663:
660:
657:
654:
651:
648:
645:
642:
631:
610:
607:
567:
547:
527:
524:
521:
518:
515:
512:
509:
455:
452:
450:
447:
432:
429:
392:Kimito Funatsu
376:mathematicians
278:optimum energy
171:
168:
156:Ronald C. Read
141:symmetry group
76:
73:
57:retrosynthesis
39:generation in
15:
9:
6:
4:
3:
2:
4675:
4664:
4661:
4659:
4656:
4654:
4651:
4649:
4646:
4644:
4641:
4640:
4638:
4628:
4623:
4619:
4618:
4606:
4602:
4597:
4592:
4587:
4582:
4578:
4574:
4570:
4563:
4555:
4551:
4546:
4541:
4536:
4531:
4527:
4523:
4519:
4512:
4504:
4500:
4496:
4492:
4487:
4482:
4478:
4474:
4470:
4466:
4462:
4458:
4457:
4452:
4448:
4441:
4433:
4429:
4425:
4421:
4416:
4411:
4407:
4403:
4398:
4393:
4389:
4385:
4384:
4379:
4372:
4364:
4360:
4356:
4352:
4348:
4344:
4340:
4336:
4332:
4325:
4317:
4313:
4309:
4305:
4301:
4297:
4293:
4289:
4288:
4280:
4272:
4268:
4264:
4260:
4256:
4252:
4248:
4244:
4240:
4236:
4232:
4225:
4217:
4213:
4209:
4205:
4201:
4197:
4193:
4189:
4182:
4174:
4170:
4166:
4162:
4158:
4154:
4150:
4146:
4142:
4138:
4131:
4123:
4119:
4115:
4111:
4107:
4103:
4099:
4095:
4087:
4079:
4075:
4071:
4067:
4063:
4059:
4055:
4051:
4047:
4043:
4036:
4028:
4024:
4020:
4016:
4011:
4006:
4002:
3998:
3994:
3990:
3985:
3980:
3976:
3972:
3971:
3966:
3962:
3961:Hongming Chen
3955:
3947:
3943:
3939:
3935:
3931:
3927:
3923:
3919:
3915:
3911:
3910:
3905:
3898:
3890:
3886:
3882:
3878:
3873:
3868:
3864:
3860:
3855:
3850:
3846:
3842:
3841:
3836:
3829:
3821:
3817:
3813:
3809:
3805:
3801:
3797:
3793:
3789:
3785:
3784:
3776:
3768:
3764:
3760:
3756:
3752:
3748:
3744:
3740:
3736:
3732:
3731:
3726:
3719:
3711:
3707:
3703:
3699:
3695:
3691:
3687:
3683:
3679:
3675:
3674:
3669:
3662:
3654:
3650:
3646:
3642:
3638:
3634:
3630:
3626:
3622:
3618:
3614:
3607:
3599:
3595:
3588:
3580:
3576:
3572:
3568:
3564:
3560:
3556:
3552:
3545:
3537:
3533:
3529:
3525:
3521:
3517:
3513:
3509:
3502:
3494:
3490:
3486:
3482:
3478:
3474:
3470:
3466:
3465:
3457:
3449:
3442:
3434:
3430:
3426:
3422:
3418:
3414:
3410:
3406:
3402:
3398:
3397:
3392:
3386:
3378:
3374:
3370:
3366:
3362:
3358:
3354:
3350:
3343:
3335:
3331:
3327:
3323:
3318:
3313:
3309:
3305:
3300:
3295:
3291:
3287:
3286:
3281:
3274:
3266:
3262:
3258:
3254:
3250:
3246:
3242:
3238:
3234:
3230:
3223:
3215:
3211:
3207:
3203:
3199:
3195:
3191:
3187:
3180:
3172:
3168:
3164:
3160:
3156:
3152:
3148:
3144:
3137:
3129:
3125:
3121:
3117:
3113:
3109:
3105:
3101:
3100:
3092:
3084:
3080:
3076:
3072:
3067:
3062:
3058:
3054:
3049:
3044:
3040:
3036:
3035:
3030:
3023:
3015:
3011:
3007:
3003:
2999:
2995:
2991:
2987:
2983:
2979:
2971:
2963:
2959:
2955:
2951:
2947:
2943:
2939:
2935:
2934:
2926:
2918:
2914:
2910:
2906:
2902:
2898:
2894:
2890:
2883:
2875:
2871:
2867:
2863:
2859:
2855:
2851:
2847:
2843:
2839:
2835:
2829:
2821:
2817:
2813:
2809:
2805:
2801:
2797:
2793:
2789:
2783:
2775:
2771:
2767:
2763:
2759:
2755:
2751:
2747:
2746:
2738:
2730:
2726:
2722:
2718:
2714:
2710:
2706:
2702:
2701:
2696:
2695:Carl Djerassi
2689:
2681:
2677:
2673:
2669:
2665:
2661:
2657:
2653:
2652:
2644:
2636:
2632:
2628:
2624:
2620:
2616:
2612:
2608:
2607:
2599:
2591:
2587:
2583:
2579:
2575:
2571:
2567:
2563:
2555:
2547:
2543:
2539:
2535:
2531:
2527:
2523:
2519:
2518:
2510:
2502:
2498:
2491:
2483:
2479:
2475:
2471:
2467:
2463:
2459:
2455:
2451:
2447:
2446:
2441:
2437:
2431:
2423:
2419:
2412:
2404:
2400:
2396:
2392:
2388:
2384:
2380:
2376:
2375:
2367:
2359:
2355:
2351:
2347:
2343:
2339:
2335:
2328:
2320:
2316:
2312:
2308:
2304:
2300:
2296:
2292:
2291:
2286:
2282:
2275:
2267:
2263:
2256:
2248:
2244:
2240:
2236:
2232:
2228:
2224:
2220:
2216:
2212:
2211:
2206:
2199:
2191:
2187:
2182:
2177:
2172:
2167:
2163:
2159:
2155:
2151:
2147:
2140:
2136:
2134:
2130:
2126:
2122:
2118:
2113:
2108:
2104:
2100:
2095:
2090:
2086:
2082:
2081:
2076:
2072:
2066:
2062:
2057:
2050:
2038:
2035:
2034:
2023:
2015:
2012:
2011:
2007:
1993:
1990:
1989:
1985:
1975:
1972:
1971:
1967:
1957:
1954:
1953:
1949:
1939:
1936:
1935:
1931:
1923:
1920:
1919:
1915:
1907:
1904:
1903:
1899:
1891:
1888:
1887:
1883:
1867:
1864:
1863:
1859:
1837:
1834:
1833:
1829:
1821:
1818:
1817:
1813:
1801:
1798:
1797:
1793:
1790:
1789:
1786:
1778:
1776:
1760:
1737:
1731:
1728:
1725:
1705:
1682:
1658:
1655:
1652:
1628:
1605:
1602:
1599:
1593:
1585:
1568:
1565:
1562:
1556:
1553:
1533:
1513:
1485:
1479:
1476:
1470:
1464:
1458:
1451:
1448:
1445:
1438:
1430:
1416:
1396:
1373:
1370:
1367:
1361:
1358:
1335:
1332:
1329:
1318:
1317:automorphisms
1314:
1298:
1291:
1287:
1271:
1268:
1245:
1239:
1236:
1233:
1213:
1193:
1185:
1166:
1160:
1157:
1154:
1134:
1126:
1118:
1100:
1097:
1093:
1089:
1086:
1064:
1061:
1057:
1048:
1047:
1031:
1011:
991:
988:
985:
982:
979:
959:
939:
931:
930:
929:
915:
895:
888:
885:
869:
861:
837:
831:
825:
822:
816:
807:
804:
801:
790:
788:
772:
752:
738:
735:
732:
729:
726:
723:
720:
717:
714:
711:
708:
691:
685:
678:
677:
673:
670:
667:
664:
661:
658:
655:
652:
649:
646:
643:
629:
622:
621:
618:
616:
606:
604:
600:
596:
592:
588:
584:
579:
565:
545:
522:
519:
516:
510:
507:
499:
495:
491:
487:
483:
475:
471:
468:
466:
460:
446:
442:
439:
428:
426:
422:
417:
415:
410:
406:
402:
397:
393:
388:
385:
381:
377:
373:
372:closed-source
368:
366:
362:
358:
354:
350:
345:
343:
339:
333:
331:
327:
323:
315:
311:
307:
304:
300:
295:
292:
288:
284:
279:
275:
271:
267:
263:
258:
254:
250:
246:
242:
236:
234:
230:
226:
220:
218:
217:hybridization
215:, with their
214:
210:
206:
202:
197:
193:
189:
185:
181:
177:
167:
165:
161:
157:
153:
149:
144:
142:
138:
137:stereoisomers
134:
130:
126:
122:
118:
113:
109:
105:
97:
93:
89:
85:
81:
72:
70:
66:
62:
58:
54:
50:
46:
42:
38:
34:
30:
26:
22:
4576:
4572:
4562:
4525:
4521:
4511:
4460:
4454:
4440:
4387:
4381:
4371:
4341:(1): 33–35.
4338:
4334:
4324:
4291:
4285:
4279:
4238:
4234:
4224:
4191:
4187:
4181:
4140:
4136:
4130:
4097:
4093:
4086:
4048:(2): 87–93.
4045:
4041:
4035:
3974:
3968:
3954:
3913:
3907:
3897:
3844:
3838:
3828:
3787:
3781:
3775:
3734:
3728:
3718:
3677:
3671:
3661:
3620:
3616:
3606:
3597:
3587:
3557:(1): 31–48.
3554:
3550:
3544:
3511:
3507:
3501:
3468:
3462:
3456:
3441:
3400:
3394:
3385:
3352:
3348:
3342:
3289:
3283:
3273:
3232:
3228:
3222:
3189:
3185:
3179:
3146:
3142:
3136:
3106:(1): 75–85.
3103:
3097:
3091:
3038:
3032:
3022:
2981:
2977:
2970:
2937:
2931:
2925:
2892:
2888:
2882:
2841:
2837:
2828:
2795:
2791:
2782:
2749:
2743:
2737:
2704:
2698:
2688:
2658:(1): 73–79.
2655:
2649:
2643:
2610:
2604:
2598:
2565:
2561:
2554:
2521:
2515:
2509:
2500:
2490:
2449:
2443:
2430:
2421:
2417:
2411:
2378:
2372:
2366:
2333:
2327:
2294:
2288:
2274:
2265:
2255:
2214:
2208:
2198:
2153:
2149:
2139:
2084:
2078:
2046:
1928:.sourceforge
1894:maygenerator
1784:
1697:
1505:
1122:
857:
744:
612:
580:
490:multiplicity
479:
473:
469:
462:
443:
434:
418:
414:metabolomics
389:
369:
365:group theory
346:
342:parallelized
334:
319:
313:
296:
237:
233:constructive
221:
173:
145:
101:
95:
91:
87:
83:
65:metabolomics
20:
18:
3790:: 158–170.
2933:Tetrahedron
2210:Biopolymers
1960:sourceforge
1942:sourceforge
1872:.univ-reims
972:satisfying
908:defined on
884:associative
615:permutation
425:autoencoder
357:isomorphism
257:open-source
41:drug design
4637:Categories
4463:: 170073.
4363:Q105033277
3984:1711.07839
3579:Q105033085
3493:Q105032974
3425:1394.05079
3403:: 94–112.
3171:Q105032775
2474:0404.05051
2424:: 131–135.
2358:Q105092432
2129:Q104747658
2043:References
1982:/steinbeck
1718:, denoted
1290:finite set
1079:such that
262:stochastic
251:and other
4579:(1): 24.
4528:(1): 48.
4503:Q33718167
4477:2052-4463
4432:Q21146620
4406:1758-2946
4390:(1): 23.
4355:0163-5700
4316:Q57818162
4308:0958-0344
4271:Q52009004
4255:1520-5142
4216:Q99233866
4208:1520-5142
4173:Q45023689
4157:1520-5142
4122:Q51947334
4106:1369-7056
4078:Q38594392
4062:1520-5142
4027:Q48127458
4001:1868-1743
3946:Q38681438
3930:1543-8384
3889:Q47199780
3863:1758-2946
3847:(1): 64.
3820:Q47256449
3804:1096-7176
3767:Q51758769
3751:1868-1743
3710:Q39092888
3694:1868-1743
3653:Q50627884
3637:0920-654X
3571:0259-9791
3536:Q99233768
3528:1520-5142
3485:1571-0661
3471:: 53–60.
3433:Q99301767
3417:0747-7171
3377:Q99233853
3369:1520-5142
3334:Q27499209
3308:1758-2946
3292:(1): 21.
3265:Q52016182
3249:1520-5142
3214:Q99233862
3206:1520-5142
3163:1006-9291
3128:Q99233968
3120:0003-2670
3083:Q38264559
3057:1758-2946
3041:(1): 27.
3014:Q43616194
2998:0937-0633
2962:Q57818172
2954:0040-4020
2917:Q28837961
2909:1520-5142
2874:Q28837910
2858:1520-5142
2820:Q50368945
2812:1433-7851
2774:Q99233812
2766:0169-7439
2729:Q99233344
2721:0022-3263
2680:Q99233839
2672:0169-7439
2635:Q99233261
2627:0003-2670
2590:Q99233202
2582:1520-5142
2546:Q99232471
2538:0003-2700
2482:Q99232279
2466:0364-9024
2403:Q99232065
2395:0022-2860
2350:2194-5357
2336:: 69–78.
2319:Q29387651
2311:0004-3702
2247:Q69715633
2231:0006-3525
2205:Karplus M
2103:1553-734X
2059:license (
2056:CC BY 4.0
2002:/software
1964:/projects
1846:/projects
1806:.upstream
1799:ASSEMBLE
1098:−
1090:∗
1062:−
983:∗
896:∗
805:∗
603:saturated
591:connected
467:molecule.
465:serotonin
438:recursive
396:QSPR/QSAR
322:canonical
49:QSAR/QSPR
4605:35461261
4554:34217353
4499:Wikidata
4495:28534867
4428:Wikidata
4424:26136848
4359:Wikidata
4312:Wikidata
4267:Wikidata
4263:14502476
4212:Wikidata
4169:Wikidata
4165:15032522
4118:Wikidata
4114:16523386
4074:Wikidata
4023:Wikidata
4019:29235269
3942:Wikidata
3938:28703000
3885:Wikidata
3881:29260340
3816:Wikidata
3812:29233745
3763:Wikidata
3759:27463853
3706:Wikidata
3702:27485423
3649:Wikidata
3645:27299746
3575:Wikidata
3532:Wikidata
3489:Wikidata
3429:Wikidata
3373:Wikidata
3330:Wikidata
3326:22985496
3261:Wikidata
3257:12767130
3210:Wikidata
3167:Wikidata
3124:Wikidata
3079:Wikidata
3075:21797997
3010:Wikidata
3006:11371077
2958:Wikidata
2913:Wikidata
2870:Wikidata
2866:11749575
2816:Wikidata
2770:Wikidata
2725:Wikidata
2676:Wikidata
2631:Wikidata
2586:Wikidata
2542:Wikidata
2478:Wikidata
2399:Wikidata
2354:Wikidata
2315:Wikidata
2243:Wikidata
2190:33400699
2125:Wikidata
2121:33400699
2031:See also
1850:/DENDRAL
1311:and its
599:chemists
595:mixtures
482:vertices
291:ACD Labs
196:spectral
164:run time
162:and the
108:valences
4596:9034616
4545:8254276
4486:5441290
4415:4486400
4070:3392122
4010:5836887
3872:5736515
3317:3558358
3066:3162559
2239:3801593
2181:7785115
2158:Bibcode
2112:7785115
2020:.github
2004:/MS-DOS
1984:/seneca
1973:SENECA
1948:/openmg
1921:MOLSIG
1912:.molgen
1905:MOLGEN
1896:.github
1889:MAYGEN
1429:, then
1313:subsets
587:valence
338:storage
117:DENDRAL
75:History
37:library
4603:
4593:
4552:
4542:
4501:
4493:
4483:
4475:
4430:
4422:
4412:
4404:
4361:
4353:
4314:
4306:
4269:
4261:
4253:
4214:
4206:
4171:
4163:
4155:
4120:
4112:
4104:
4076:
4068:
4060:
4025:
4017:
4007:
3999:
3944:
3936:
3928:
3887:
3879:
3869:
3861:
3818:
3810:
3802:
3765:
3757:
3749:
3708:
3700:
3692:
3651:
3643:
3635:
3577:
3569:
3534:
3526:
3491:
3483:
3431:
3423:
3415:
3375:
3367:
3332:
3324:
3314:
3306:
3263:
3255:
3247:
3212:
3204:
3169:
3161:
3126:
3118:
3081:
3073:
3063:
3055:
3012:
3004:
2996:
2960:
2952:
2915:
2907:
2872:
2864:
2856:
2818:
2810:
2772:
2764:
2727:
2719:
2678:
2670:
2633:
2625:
2588:
2580:
2544:
2536:
2480:
2472:
2464:
2401:
2393:
2356:
2348:
2317:
2309:
2245:
2237:
2229:
2188:
2178:
2127:
2119:
2109:
2101:
2013:Surge
1978:github
1926:molsig
1878:/index
1854:_GENOA
1819:COCON
1389:, and
583:degree
353:matrix
274:energy
241:memory
3979:arXiv
2006:/SMOG
1991:SMOG
1882:.html
1858:/view
1824:cocon
1812:.html
1810:/main
1794:Link
1791:Name
1775:InChI
1641:, if
1182:is a
1125:order
860:group
486:edges
188:bonds
133:atoms
104:graph
43:, in
4601:PMID
4550:PMID
4491:PMID
4473:ISSN
4420:PMID
4402:ISSN
4351:ISSN
4304:ISSN
4259:PMID
4251:ISSN
4204:ISSN
4161:PMID
4153:ISSN
4110:PMID
4102:ISSN
4066:PMID
4058:ISSN
4015:PMID
3997:ISSN
3934:PMID
3926:ISSN
3877:PMID
3859:ISSN
3808:PMID
3800:ISSN
3755:PMID
3747:ISSN
3698:PMID
3690:ISSN
3641:PMID
3633:ISSN
3567:ISSN
3524:ISSN
3481:ISSN
3413:ISSN
3365:ISSN
3322:PMID
3304:ISSN
3253:PMID
3245:ISSN
3202:ISSN
3159:ISSN
3116:ISSN
3071:PMID
3053:ISSN
3002:PMID
2994:ISSN
2950:ISSN
2905:ISSN
2862:PMID
2854:ISSN
2808:ISSN
2762:ISSN
2717:ISSN
2668:ISSN
2623:ISSN
2578:ISSN
2534:ISSN
2462:ISSN
2391:ISSN
2346:ISSN
2307:ISSN
2235:PMID
2227:ISSN
2186:PMID
2117:PMID
2099:ISSN
2067:):
2061:2021
2000:/cca
1998:.net
1980:.com
1962:.net
1955:PMG
1944:.net
1937:OMG
1930:.net
1880:_ENG
1876:/LSD
1865:LSD
1856:.zip
1844:.org
1826:.nmr
1123:The
765:and
496:are
484:and
283:COSY
268:and
249:HSQC
245:HMBC
225:tree
211:and
129:NASA
90:and
4591:PMC
4581:doi
4540:PMC
4530:doi
4481:PMC
4465:doi
4410:PMC
4392:doi
4343:doi
4296:doi
4243:doi
4196:doi
4145:doi
4050:doi
4005:PMC
3989:doi
3918:doi
3867:PMC
3849:doi
3792:doi
3739:doi
3682:doi
3625:doi
3559:doi
3516:doi
3473:doi
3469:299
3421:Zbl
3405:doi
3357:doi
3312:PMC
3294:doi
3237:doi
3194:doi
3151:doi
3108:doi
3104:298
3061:PMC
3043:doi
2986:doi
2982:369
2942:doi
2897:doi
2846:doi
2800:doi
2754:doi
2709:doi
2660:doi
2615:doi
2611:133
2570:doi
2526:doi
2470:Zbl
2454:doi
2422:260
2383:doi
2338:doi
2299:doi
2219:doi
2176:PMC
2166:doi
2107:PMC
2089:doi
2063:) (
2022:.io
1996:ccl
1914:.de
1910:www
1898:.io
1874:.fr
1870:eos
1848:/AI
1840:www
1828:.de
1808:.ch
1804:www
1584:if
1526:of
1286:Set
1261:is
1206:is
1024:of
952:in
736:10
715:11
674:11
671:10
474:(B)
470:(A)
253:NMR
96:(C)
92:(B)
88:(A)
4639::
4599:.
4589:.
4577:14
4575:.
4571:.
4548:.
4538:.
4526:13
4524:.
4520:.
4497:.
4489:.
4479:.
4471:.
4459:.
4453:.
4426:.
4418:.
4408:.
4400:.
4386:.
4380:.
4357:.
4349:.
4339:30
4337:.
4310:.
4302:.
4290:.
4265:.
4257:.
4249:.
4239:43
4237:.
4233:.
4210:.
4202:.
4192:35
4190:.
4167:.
4159:.
4151:.
4141:44
4139:.
4116:.
4108:.
4096:.
4072:.
4064:.
4056:.
4046:28
4044:.
4021:.
4013:.
4003:.
3995:.
3987:.
3975:37
3973:.
3967:.
3940:.
3932:.
3924:.
3914:14
3912:.
3883:.
3875:.
3865:.
3857:.
3843:.
3837:.
3814:.
3806:.
3798:.
3788:45
3786:.
3761:.
3753:.
3745:.
3735:29
3733:.
3704:.
3696:.
3688:.
3678:33
3676:.
3647:.
3639:.
3631:.
3621:30
3619:.
3596:.
3573:.
3565:.
3553:.
3530:.
3522:.
3512:36
3510:.
3487:.
3479:.
3467:.
3427:.
3419:.
3411:.
3401:60
3399:.
3371:.
3363:.
3353:32
3351:.
3328:.
3320:.
3310:.
3302:.
3288:.
3282:.
3259:.
3251:.
3243:.
3233:43
3231:.
3208:.
3200:.
3190:34
3188:.
3165:.
3157:.
3147:43
3145:.
3122:.
3114:.
3102:.
3077:.
3069:.
3059:.
3051:.
3037:.
3031:.
3008:.
3000:.
2992:.
2980:.
2956:.
2948:.
2938:47
2936:.
2911:.
2903:.
2893:36
2891:.
2868:.
2860:.
2852:.
2842:41
2814:.
2806:.
2796:35
2794:.
2768:.
2760:.
2748:.
2723:.
2715:.
2705:46
2703:.
2674:.
2666:.
2656:51
2654:.
2629:.
2621:.
2609:.
2584:.
2576:.
2566:18
2564:.
2540:.
2532:.
2522:47
2520:.
2499:.
2476:.
2468:.
2460:.
2448:.
2438:;
2420:.
2397:.
2389:.
2379:31
2377:.
2352:.
2344:.
2313:.
2305:.
2295:61
2293:.
2283:;
2264:.
2241:.
2233:.
2225:.
2215:26
2213:.
2184:.
2174:.
2164:.
2154:17
2152:.
2148:.
2123:.
2115:.
2105:.
2097:.
2085:17
2083:.
2077:.
1946:/p
1284:.
862:,
739:3
733:7
730:9
727:8
724:5
721:1
718:6
712:2
709:4
668:9
665:8
662:7
659:6
656:5
653:4
650:3
647:2
644:1
416:.
247:,
207:,
203:,
139:,
110:,
71:.
55:,
19:A
4607:.
4583::
4556:.
4532::
4505:.
4467::
4461:4
4434:.
4394::
4388:7
4365:.
4345::
4318:.
4298::
4292:3
4273:.
4245::
4218:.
4198::
4175:.
4147::
4124:.
4098:9
4080:.
4052::
4029:.
3991::
3981::
3948:.
3920::
3891:.
3851::
3845:9
3822:.
3794::
3769:.
3741::
3712:.
3684::
3655:.
3627::
3581:.
3561::
3555:2
3538:.
3518::
3495:.
3475::
3435:.
3407::
3379:.
3359::
3336:.
3296::
3290:4
3267:.
3239::
3216:.
3196::
3173:.
3153::
3130:.
3110::
3085:.
3045::
3039:3
3016:.
2988::
2964:.
2944::
2919:.
2899::
2876:.
2848::
2822:.
2802::
2776:.
2756::
2750:8
2731:.
2711::
2682:.
2662::
2637:.
2617::
2592:.
2572::
2548:.
2528::
2484:.
2456::
2450:3
2405:.
2385::
2360:.
2340::
2321:.
2301::
2249:.
2221::
2192:.
2168::
2160::
2131:.
2091::
1761:V
1741:)
1738:G
1735:(
1732:t
1729:u
1726:A
1706:G
1695:.
1683:E
1662:)
1659:v
1656:,
1653:u
1650:(
1629:E
1609:)
1606:v
1603:,
1600:u
1597:(
1594:a
1572:)
1569:V
1566:,
1563:E
1560:(
1557:=
1554:G
1534:V
1514:a
1492:)
1489:)
1486:v
1483:(
1480:a
1477:,
1474:)
1471:u
1468:(
1465:a
1462:(
1459:=
1456:)
1452:v
1449:,
1446:u
1442:(
1439:a
1417:V
1397:a
1377:)
1374:E
1371:,
1368:V
1365:(
1362:=
1359:G
1339:)
1336:v
1333:,
1330:u
1327:(
1299:X
1272:!
1269:n
1249:)
1246:X
1243:(
1240:m
1237:y
1234:S
1214:n
1194:X
1170:)
1167:X
1164:(
1161:m
1158:y
1155:S
1135:X
1119:.
1101:1
1094:g
1087:g
1065:1
1058:g
1044:.
1032:G
1012:g
992:g
989:=
986:I
980:g
960:G
940:I
916:G
870:G
844:)
841:)
838:x
835:(
832:b
829:(
826:a
823:=
820:)
817:x
814:(
811:)
808:b
802:a
799:(
773:b
753:a
695:)
692:x
689:(
686:f
630:x
566:E
546:V
526:)
523:E
520:,
517:V
514:(
511:=
508:G
213:S
209:O
205:N
201:C
98:.
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.