916:) uses UV radiation to link proteins to RNA molecules in a tissue during splicing. A trans-acting splicing regulatory protein of interest is then precipitated using specific antibodies. When the RNA attached to that protein is isolated and cloned, it reveals the target sequences for that protein. Another method for identifying RNA-binding proteins and mapping their binding to pre-mRNA transcripts is "Microarray Evaluation of Genomic Aptamers by shift (MEGAshift)".net This method involves an adaptation of the "Systematic Evolution of Ligands by Exponential Enrichment (SELEX)" method together with a microarray-based readout. Use of the MEGAshift method has provided insights into the regulation of alternative splicing by allowing for the identification of sequences in pre-mRNA transcripts surrounding alternatively spliced exons that mediate binding to different splicing factors, such as ASF/SF2 and PTB. This approach has also been used to aid in determining the relationship between RNA secondary structure and the binding of splicing factors.
3755:ΔFosB is an essential transcription factor implicated in the molecular and behavioral pathways of addiction following repeated drug exposure. The formation of ΔFosB in multiple brain regions, and the molecular pathway leading to the formation of AP-1 complexes is well understood. The establishment of a functional purpose for ΔFosB has allowed further determination as to some of the key aspects of its molecular cascades, involving effectors such as GluR2 (87,88), Cdk5 (93) and NFkB (100). Moreover, many of these molecular changes identified are now directly linked to the structural, physiological and behavioral changes observed following chronic drug exposure (60,95,97,102). New frontiers of research investigating the molecular roles of ΔFosB have been opened by epigenetic studies, and recent advances have illustrated the role of ΔFosB acting on DNA and histones, truly as a
427:
in some cases, but decrease the probability in other cases, depending on context. The context within which regulatory elements act includes cis-acting context that is established by the presence of other RNA sequence features, and trans-acting context that is established by cellular conditions. For example, some cis-acting RNA sequence elements influence splicing only if multiple elements are present in the same region so as to establish context. As another example, a cis-acting element can have opposite effects on splicing, depending on which proteins are expressed in the cell (e.g., neuronal versus non-neuronal PTB). The adaptive significance of splicing silencers and enhancers is attested by studies showing that there is strong selection in human genes against mutations that produce new silencers or disrupt existing enhancers.
924:
protein-coding genes assembled from numerous RNA sequencing experiments across a variety of human tissues. This comprehensive analysis has led to the identification of numerous isoforms with more confidently predicted structure and potentially superior function compared to canonical isoforms in the latest human gene database. By integrating structural predictions with expression and evolutionary evidence, this approach has demonstrated the potential of protein structure prediction as a tool for refining the annotation of the human genome.
661:) as enriched in the non-constitutive exons suggesting that protein isoforms may display functional diversity due to the alteration of functional modules within these regions. Such functional diversity achieved by isoforms is reflected by their expression patterns and can be predicted by machine learning approaches. Comparative studies indicate that alternative splicing preceded multicellularity in evolution, and suggest that this mechanism might have been co-opted to assist in the development of multicellular organisms.
3759:(34). As a consequence of our improved understanding of ΔFosB in addiction, it is possible to evaluate the addictive potential of current medications (119), as well as use it as a biomarker for assessing the efficacy of therapeutic interventions (121,122,124). Some of these proposed interventions have limitations (125) or are in their infancy (75). However, it is hoped that some of these preliminary findings may lead to innovative treatments, which are much needed in addiction.
557:
31:
382:
enhancer element may serve as a repressor when bound to its splicing element in the context of an exon, and vice versa. The secondary structure of the pre-mRNA transcript also plays a role in regulating splicing, such as by bringing together splicing elements or by masking a sequence that would otherwise serve as a binding element for a splicing factor. Together, these elements form a "splicing code" that governs how splicing will occur under different cellular conditions.
758:. One study found that a relatively small percentage (383 out of over 26000) of alternative splicing variants were significantly higher in frequency in tumor cells than normal cells, suggesting that there is a limited set of genes which, when mis-spliced, contribute to tumor development. It is believed however that the deleterious effects of mis-spliced transcripts are usually safeguarded and eliminated by a cellular posttranscriptional quality control mechanism termed
147:
155:
504:
419:
366:
611:
573:, or programmed cell death. Increased expression of Fas receptor in skin cells chronically exposed to the sun, and absence of expression in skin cancer cells, suggests that this mechanism may be important in elimination of pre-cancerous cells in humans. If exon 6 is skipped, the resulting mRNA encodes a soluble Fas protein that does not promote apoptosis. The inclusion or skipping of the exon depends on two antagonistic proteins,
242:
69:, where it increases the number of proteins that can be encoded by the genome. In humans, it is widely believed that ~95% of multi-exonic genes are alternatively spliced to produce functional alternative products from the same gene but many scientists believe that most of the observed splice variants are due to splicing errors and the actual number of biologically relevant alternatively spliced genes is much lower.
447:
82:
is large and comes from 5/6 of the 32kb adenovirus genome. This is much larger than any of the individual adenovirus mRNAs present in infected cells. Researchers found that the primary RNA transcript produced by adenovirus type 2 in the late phase was spliced in many different ways, resulting in mRNAs encoding different viral proteins. In addition, the primary transcript contained multiple
279:
3789:
For these reasons, ΔFosB is considered a primary and causative transcription factor in creating new neural connections in the reward centre, prefrontal cortex, and other regions of the limbic system. This is reflected in the increased, stable and long-lasting level of sensitivity to cocaine and other
707:
Changes in the RNA processing machinery may lead to mis-splicing of multiple transcripts, while single-nucleotide alterations in splice sites or cis-acting splicing regulatory sites may lead to differences in splicing of a single gene, and thus in the mRNA produced from a mutant gene's transcripts. A
597:
This mechanism is an example of exon definition in splicing. A spliceosome assembles on an intron, and the snRNP subunits fold the RNA so that the 5' and 3' ends of the intron are joined. However, recently studied examples such as this one show that there are also interactions between the ends of the
530:
undergo alternative splicing via the alternative acceptor site mode. The gene Tra encodes a protein that is expressed only in females. The primary transcript of this gene contains an intron with two possible acceptor sites. In males, the upstream acceptor site is used. This causes a longer version of
720:
and proteomics analyses have revealed striking differential expression of splice isoforms of key proteins in important cancer pathways. It is not always clear whether such aberrant patterns of splicing contribute to the cancerous growth, or are merely consequence of cellular abnormalities associated
712:
affect splicing rather than directly affecting coding sequences. A more recent study indicates that one-third of all hereditary diseases are likely to have a splicing component. Regardless of exact percentage, a number of splicing-related diseases do exist. As described below, a prominent example of
484:
well, so that U2AF proteins bind poorly to it without assistance from splicing activators. This 3' splice acceptor site is therefore not used in males. Females, however, produce the splicing activator
Transformer (Tra) (see below). The SR protein Tra2 is produced in both sexes and binds to an ESE in
426:
In general, the determinants of splicing work in an inter-dependent manner that depends on context, so that the rules governing how splicing is regulated form a splicing code. The presence of a particular cis-acting RNA sequence element may increase the probability that a nearby site will be spliced
258:
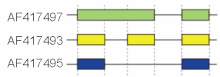
3 gene. Comparing the exonic structure shown in the first line (green) with the one in the second line (yellow) shows intron retention, whereas the comparison between the second and the third spliceform (yellow vs. blue) exhibits exon skipping. A model nomenclature to uniquely designate all possible
237:
mechanism rather than alternative splicing; by starting transcription at different points, transcripts with different 5'-most exons can be generated. At the other end, multiple polyadenylation sites provide different 3' end points for the transcript. Both of these mechanisms are found in combination
81:
produces five primary transcripts early in its infectious cycle, prior to viral DNA replication, and an additional one later, after DNA replication begins. The early primary transcripts continue to be produced after DNA replication begins. The additional primary transcript produced late in infection
61:
process during gene expression that allows a single gene to produce different splice variants. For example, some exons of a gene may be included within or excluded from the final RNA product of the gene. This means the exons are joined in different combinations, leading to different splice variants.
773:
groups to DNA, a modification that often has regulatory effects. Several abnormally spliced DNMT3B mRNAs are found in tumors and cancer cell lines. In two separate studies, expression of two of these abnormally spliced mRNAs in mammalian cells caused changes in the DNA methylation patterns in those
748:
In fact, there is actually a reduction of alternative splicing in cancerous cells compared to normal ones, and the types of splicing differ; for instance, cancerous cells show higher levels of intron retention than normal cells, but lower levels of exon skipping. Some of the differences in splicing
698:
found no large differences in frequency of alternatively spliced genes among humans and any of the other animals tested. Another study, however, proposed that these results were an artifact of the different numbers of ESTs available for the various organisms. When they compared alternative splicing
648:
Genuine alternative splicing occurs in both protein-coding genes and non-coding genes to produce multiple products (proteins or non-coding RNAs). External information is needed in order to decide which product is made, given a DNA sequence and the initial transcript. Since the methods of regulation
543:
protein from binding to the polypyrimidine tract. This prevents the use of this junction, shifting the spliceosome binding to the downstream acceptor site. Splicing at this point bypasses the stop codon, which is excised as part of the intron. The resulting mRNA encodes an active Tra protein, which
316:
The typical eukaryotic nuclear intron has consensus sequences defining important regions. Each intron has the sequence GU at its 5' end. Near the 3' end there is a branch site. The nucleotide at the branchpoint is always an A; the consensus around this sequence varies somewhat. In humans the branch
128:
In 2021, it was discovered that the genome of adenovirus type 2, the adenovirus in which alternative splicing was first identified, was able to produce a much greater variety of splice variants than previously thought. By using next generation sequencing technology, researchers were able to update
3709:
that define a state of addiction. ... A large body of literature has demonstrated that such ΔFosB induction in D1-type NAc neurons increases an animal's sensitivity to drug as well as natural rewards and promotes drug self-administration, presumably through a process of positive reinforcement
3708:
DESPITE THE IMPORTANCE OF NUMEROUS PSYCHOSOCIAL FACTORS, AT ITS CORE, DRUG ADDICTION INVOLVES A BIOLOGICAL PROCESS: the ability of repeated exposure to a drug of abuse to induce changes in a vulnerable brain that drive the compulsive seeking and taking of drugs, and loss of control over drug use,
344:
protein factors, binds to the branchpoint A within the branch site. The complex at this stage is known as the spliceosome A complex. Formation of the A complex is usually the key step in determining the ends of the intron to be spliced out, and defining the ends of the exon to be retained. (The U
919:
Use of reporter assays makes it possible to find the splicing proteins involved in a specific alternative splicing event by constructing reporter genes that will express one of two different fluorescent proteins depending on the splicing reaction that occurs. This method has been used to isolate
858:
sequences, but this requires sequencing of very large numbers of ESTs. Most EST libraries come from a very limited number of tissues, so tissue-specific splice variants are likely to be missed in any case. High-throughput approaches to investigate splicing have, however, been developed, such as:
629:
in humans, produces a single primary RNA transcript, which is alternatively spliced in multiple ways to produce over 40 different mRNAs. Equilibrium among differentially spliced transcripts provides multiple mRNAs encoding different products that are required for viral multiplication. One of the
838:
Deep sequencing technologies have been used to conduct genome-wide analyses of both unprocessed and processed mRNAs; thus providing insights into alternative splicing. For example, results from use of deep sequencing indicate that, in humans, an estimated 95% of transcripts from multiexon genes
634:
gene, in which exon 2 is a cassette exon that may be skipped or included. The inclusion of tat exon 2 in the RNA is regulated by competition between the splicing repressor hnRNP A1 and the SR protein SC35. Within exon 2 an exonic splicing silencer sequence (ESS) and an exonic splicing enhancer
381:
regulatory sites (silencers and enhancers) on the pre-mRNA. However, as part of the complexity of alternative splicing, it is noted that the effects of a splicing factor are frequently position-dependent. That is, a splicing factor that serves as a splicing activator when bound to an intronic
639:
of multiple A1 molecules, extending into the 5' donor site upstream of exon 2 and preventing the binding of the core splicing factor U2AF35 to the polypyrimidine tract. If SC35 binds to the ESE, it prevents A1 binding and maintains the 5' donor site in an accessible state for assembly of the
923:
Recent advancements in protein structure prediction have facilitated the development of new tools for genome annotation and alternative splicing anlaysis. For instance, isoform.io, a platform guided by protein structure predictions, has evaluated hundreds of thousands of isoforms of human
312:
are determined during the splicing process. The regulation and selection of splice sites are done by trans-acting splicing activator and splicing repressor proteins as well as cis-acting elements within the pre-mRNA itself such as exonic splicing enhancers and exonic splicing silencers.
834:
Transcriptome-wide analysis of alternative splicing is typically performed by high-throughput RNA-sequencing. Most commonly, by short-read sequencing, such as by
Illumina instrumentation. But even more informative, by long-read sequencing, such as by Nanopore or PacBio instrumentation.
129:
the human adenovirus type 2 transcriptome and document the presence of 904 splice variants produced by the virus through a complex pattern of alternative splicing. Very few of these splice variants have been shown to be functional, a point that the authors raise in their paper.
825:
Recent provocative studies point to a key function of chromatin structure and histone modifications in alternative splicing regulation. These insights suggest that epigenetic regulation determines not only what parts of the genome are expressed but also how they are spliced.
585:
Binding of TIA-1 protein to an intronic splicing enhancer site stabilizes binding of the U1 snRNP. The resulting 5' donor site complex assists in binding of the splicing factor U2AF to the 3' splice site upstream of the exon, through a mechanism that is not yet known (see
485:
exon 4; if Tra is present, it binds to Tra2 and, along with another SR protein, forms a complex that assists U2AF proteins in binding to the weak polypyrimidine tract. U2 is recruited to the associated branchpoint, and this leads to inclusion of exon 4 in the mRNA.
904:
from tissues of interest. The probe cDNAs bind to DNA from the exons that are included in mRNAs in their tissue of origin, or to DNA from the boundary where two exons have been joined. This can reveal the presence of particular alternatively spliced mRNAs.
680:. This finding led to speculation that the perceived greater complexity of humans, or vertebrates generally, might be due to higher rates of alternative splicing in humans than are found in invertebrates. However, a study on samples of 100,000
932:
There is a collection of alternative splicing databases. These databases are useful for finding genes having pre-mRNAs undergoing alternative splicing and alternative splicing events or to study the functional impact of alternative splicing.
393:
are sites to which splicing repressor proteins bind, reducing the probability that a nearby site will be used as a splice junction. These can be located in the intron itself (intronic splicing silencers, ISS) or in a neighboring exon
581:
The 5' donor site in the intron downstream from exon 6 in the pre-mRNA has a weak agreement with the consensus sequence, and is not bound usually by the U1 snRNP. If U1 does not bind, the exon is skipped (see "a" in accompanying
568:
protein are produced by alternative splicing. Two normally occurring isoforms in humans are produced by an exon-skipping mechanism. An mRNA including exon 6 encodes the membrane-bound form of the Fas receptor, which promotes
753:
of trans-acting splicing factors. Others may be produced by changes in the relative amounts of splicing factors produced; for instance, breast cancer cells have been shown to have increased levels of the splicing factor
356:
linkage. In the second transesterification, the 3' end of the intron is cleaved from the downstream exon, and the two exons are joined by a phosphodiester bond. The intron is then released in lariat form and degraded.
109:
mRNA contains exons 1–4, and terminates after a polyadenylation site in exon 4. Another mRNA is produced from this pre-mRNA by skipping exon 4, and includes exons 1–3, 5, and 6. It encodes a protein known as CGRP
406:
are sites to which splicing activator proteins bind, increasing the probability that a nearby site will be used as a splice junction. These also may occur in the intron (intronic splicing enhancers, ISE) or exon
652:
Alternative splicing may provide evolutionary flexibility. A single point mutation may cause a given exon to be occasionally excluded or included from a transcript during splicing, allowing production of a new
468:
contain 6 exons. In males, exons 1,2,3,5,and 6 are joined to form the mRNA, which encodes a transcriptional regulatory protein required for male development. In females, exons 1,2,3, and 4 are joined, and a
2925:
Roest
Crollius H, Jaillon O, Bernot A, Dasilva C, Bouneau L, Fischer C, et al. (June 2000). "Estimate of human gene number provided by genome-wide analysis using Tetraodon nigroviridis DNA sequence".
721:
with cancer. For certain types of cancer, like in colorectal and prostate, the number of splicing errors per cancer has been shown to vary greatly between individual cancers, a phenomenon referred to as
1554:
Maki R, Roeder W, Traunecker A, Sidman C, Wabl M, Raschke W, Tonegawa S (May 1981). "The role of DNA rearrangement and alternative RNA processing in the expression of immunoglobulin delta genes".
598:
exon. In this particular case, these exon definition interactions are necessary to allow the binding of core splicing factors prior to assembly of the spliceosomes on the two flanking introns.
3790:
drugs, and tendency to relapse even after long periods of abstinence. These newly constructed networks function very efficiently via new pathways as soon as drugs of abuse are further taken
62:
In the case of protein-coding genes, the proteins translated from these splice variants may contain differences in their amino acid sequence and in their biological functions (see Figure).
2516:"SC35 and heterogeneous nuclear ribonucleoprotein A/B proteins bind to a juxtaposed exonic splicing enhancer/exonic splicing silencer element to regulate HIV-1 tat exon 2 splicing"
254:
These modes describe basic splicing mechanisms, but may be inadequate to describe complex splicing events. For instance, the figure to the right shows 3 spliceforms from the mouse
867:. These methods can be used to screen for polymorphisms or mutations in or around splicing elements that affect protein binding. When combined with splicing assays, including
1061:
Pan Q, Shai O, Lee LJ, Frey BJ, Blencowe BJ (December 2008). "Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing".
4556:
4291:
Sommer, Markus J.; Cha, Sooyoung; Varabyou, Ales; Rincon, Natalia; Park, Sukhwan; Minkin, Ilia; Pertea, Mihaela; Steinegger, Martin; Salzberg, Steven L. (2022-12-15).
225:
In addition to these primary modes of alternative splicing, there are two other main mechanisms by which different mRNAs may be generated from the same gene; multiple
4344:"An atlas of alternative splicing profiles and functional associations reveals new regulatory programs and genes that simultaneously express multiple major isoforms"
839:
undergo alternative splicing, with a number of pre-mRNA transcripts spliced in a tissue-specific manner. Functional genomics and computational approaches based on
795:
has been found to be associated with increased levels of the SF2/ASF in breast cancer cells. The abnormal isoform of the Ron protein encoded by this mRNA leads to
699:
frequencies in random subsets of genes from each organism, the authors concluded that vertebrates do have higher rates of alternative splicing than invertebrates.
593:, where PTB can bind. If PTB binds, it inhibits the effect of the 5' donor complex on the binding of U2AF to the acceptor site, resulting in exon skipping (see c).
3401:"Transcriptome instability in colorectal cancer identified by exon microarray analyses: Associations with splicing factor expression levels and patient survival"
3301:"A new class of protein cancer biomarker candidates: differentially expressed splice variants of ERBB2 (HER2/neu) and ERBB1 (EGFR) in breast cancer cell lines"
1401:
Nevins JR, Darnell JE (December 1978). "Steps in the processing of Ad2 mRNA: poly(A)+ nuclear sequences are conserved and poly(A) addition precedes splicing".
117:
Since then, many other examples of biologically relevant alternative splicing have been found in eukaryotes. The "record-holder" for alternative splicing is a
4503:
AStalavista (Alternative
Splicing landscape visualization tool), a method for the computationally exhaustive classification of Alternative Splicing Structures
135:"An outstanding question is what roles the menagerie of novel RNAs play or whether they are spurious molecules generated by an overloaded splicing machinery."
774:
cells. Cells with one of the abnormal mRNAs also grew twice as fast as control cells, indicating a direct contribution to tumor development by this product.
217:. If the retained intron is in the coding region, the intron must encode amino acids in frame with the neighboring exons, or a stop codon or a shift in the
150:
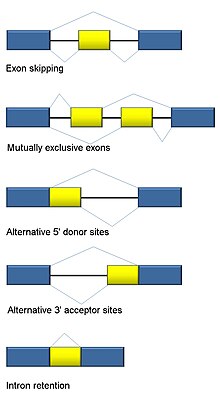
Traditional classification of basic types of alternative RNA splicing events. Exons are represented as blue and yellow blocks, introns as lines in between.
851:
that are released during splicing, the determination of branch site sequences, and the large-scale mapping of branchpoints in human pre-mRNA transcripts.
213:: A sequence may be spliced out as an intron or simply retained. This is distinguished from exon skipping because the retained sequence is not flanked by
473:
signal in exon 4 causes cleavage of the mRNA at that point. The resulting mRNA is a transcriptional regulatory protein required for female development.
843:
have also been developed to integrate RNA-seq data to predict functions for alternatively spliced isoforms. Deep sequencing has also aided in the
4600:
4549:
539:(Sxl). The Sxl protein is a splicing repressor that binds to an ISS in the RNA of the Tra transcript near the upstream acceptor site, preventing
2557:"A second exon splicing silencer within human immunodeficiency virus type 1 tat exon 2 represses splicing of Tat mRNA and binds protein hnRNP H"
791:. An important property of cancerous cells is their ability to move and invade normal tissue. Production of an abnormally spliced transcript of
3621:"Abnormally spliced beta-globin mRNAs: a single point mutation generates transcripts sensitive and insensitive to nonsense-mediated mRNA decay"
352:
reactions. In the first transesterification, 5' end of the intron is cleaved from the upstream exon and joined to the branch site A by a 2',5'-
1444:
Rosenfeld MG, Amara SG, Roos BA, Ong ES, Evans RM (March 1981). "Altered expression of the calcitonin gene associated with RNA polymorphism".
4199:
Tuerk C, Gold L (August 1990). "Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase".
3061:
López-Bigas N, Audit B, Ouzounis C, Parra G, Guigó R (March 2005). "Are splicing mutations the most frequent cause of hereditary disease?".
725:. Transcriptome instability has further been shown to correlate grealty with reduced expression level of splicing factor genes. Mutation of
2825:"Functional and evolutionary analysis of alternatively spliced genes is consistent with an early eukaryotic origin of alternative splicing"
536:
835:
Transcriptome-wide analyses can for example be used to measure the amount of deviating alternative splicing, such as in a cancer cohort.
4542:
749:
in cancerous cells may be due to the high frequency of somatic mutations in splicing factor genes, and some may result from changes in
3771:
Biliński P, Wojtyła A, Kapka-Skrzypczak L, Chwedorowicz R, Cyranka M, Studziński T (2012). "Epigenetic regulation in drug addiction".
4763:
4758:
1323:
Leff SE, Rosenfeld MG, Evans RM (1986). "Complex transcriptional units: diversity in gene expression by alternative RNA processing".
893:
4768:
4743:
2655:"Alternative splicing in concert with protein intrinsic disorder enables increased functional diversity in multicellular organisms"
640:
spliceosome. Competition between the activator and repressor ensures that both mRNA types (with and without exon 2) are produced.
399:
535:. The resulting mRNA encodes a truncated protein product that is inactive. Females produce the master sex determination protein
4841:
398:, ESS). They vary in sequence, as well as in the types of proteins that bind to them. The majority of splicing repressors are
2065:"Next-generation SELEX identifies sequence and structural determinants of splicing factor binding in human pre-mRNA sequence"
1931:
17:
385:
There are two major types of cis-acting RNA sequence elements present in pre-mRNAs and they have corresponding trans-acting
874:
assays, the functional effects of polymorphisms or mutations on the splicing of pre-mRNA transcripts can then be analyzed.
4505:
249:
gene. Directionality of transcription from 5' to 3' is shown from left to right. Exons and introns are not drawn to scale.
4244:"High-throughput binding analysis determines the binding specificity of ASF/SF2 on alternatively spliced human pre-mRNAs"
2330:"Assembly of specific SR protein complexes on distinct regulatory elements of the Drosophila doublesex splicing enhancer"
658:
4514:
4610:
114:). Examples of alternative splicing in immunoglobin gene transcripts in mammals were also observed in the early 1980s.
348:
The U4,U5,U6 complex binds, and U6 replaces the U1 position. U1 and U4 leave. The remaining complex then performs two
105:
was found to be alternatively spliced in mammalian cells. The primary transcript from this gene contains 6 exons; the
4565:
1954:"Using positional distribution to identify splicing elements and predict pre-mRNA processing defects in human genes"
1358:
Chow LT, Broker TR (October 1978). "The spliced structures of adenovirus 2 fiber message and the other late mRNAs".
4059:"Revealing global regulatory features of mammalian alternative splicing using a quantitative microarray platform"
1007:
920:
mutants affecting splicing and thus to identify novel splicing regulatory proteins inactivated in those mutants.
340:, U4, U5, and U6 (U3 is not involved in mRNA splicing). U1 binds to the 5' GU and U2, with the assistance of the
111:
2969:
Brett D, Pospisil H, Valcárcel J, Reich J, Bork P (January 2002). "Alternative splicing and genome complexity".
3452:"Abnormal RNA splicing and genomic instability after induction of DNMT3A mutations by CRISPR/Cas9 gene editing"
3499:
Kim E, Goren A, Ast G (January 2008). "Insights into the connection between cancer and alternative splicing".
2766:"Systematically differentiating functions for alternatively spliced isoforms through integrating RNA-seq data"
221:
will cause the protein to be non-functional. This is the rarest mode in mammals but the most common in plants.
4529:
2174:
Barash Y, Calarco JA, Gao W, Pan Q, Wang X, Shai O, et al. (May 2010). "Deciphering the splicing code".
967:
207:: An alternative 3' splice junction (acceptor site) is used, changing the 5' boundary of the downstream exon.
198:
1824:
Ner-Gaon, Hadas; Halachmi, Ronit; Savaldi-Goldstein, Sigal; Rubin, Eitan; Ophir, Ron; Fluhr, Robert (2004).
759:
308:
there can be more than 100 introns and exons in one transcribed pre-mRNA.) The exons to be retained in the
4520:
Stamms-lab.net: Research Group dealing with alternative
Splicing issues and mis-splicing in human diseases
4605:
1649:
Westergren
Jakobsson A, Segerman B, Wallerman O, Lind SB, Zhao H, Rubin CJ, et al. (November 2020).
4342:
Tapial J, Ha KC, Sterne-Weiler T, Gohr A, Braunschweig U, Hermoso-Pulido A, et al. (October 2017).
3580:"Identification of alternatively spliced mRNA variants related to cancers by genome-wide ESTs alignment"
1781:
Matlin AJ, Clark F, Smith CW (May 2005). "Understanding alternative splicing: towards a cellular code".
708:
study in 2005 involving probabilistic analyses indicated that greater than 60% of human disease-causing
840:
234:
4442:
Rodriguez JM, Maietta P, Ezkurdia I, Pietrelli A, Wesselink JJ, Lopez G, et al. (January 2013).
742:
722:
3537:
Ghigna C, Giordano S, Shen H, Benvenuto F, Castiglioni F, Comoglio PM, et al. (December 2005).
2472:"Regulation of Fas alternative splicing by antagonistic effects of TIA-1 and PTB on exon definition"
668:
and other genome sequencing has shown that humans have only about 30% more genes than the roundworm
657:
without loss of the original protein. Studies have identified intrinsically disordered regions (see
4864:
3252:"Aberrant RNA splicing in cancer; expression changes and driver mutations of splicing factor genes"
732:
408:
395:
178:
158:
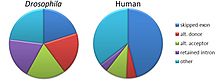
Relative frequencies of types of alternative splicing events differ between humans and fruit flies.
90:
3902:"Deviating alternative splicing as a molecular subtype of microsatellite stable colorectal cancer"
2882:
Ewing B, Green P (June 2000). "Analysis of expressed sequence tags indicates 35,000 human genes".
1216:(September 1977). "An amazing sequence arrangement at the 5' ends of adenovirus 2 messenger RNA".
4811:
4632:
3721:
Ruffle JK (November 2014). "Molecular neurobiology of addiction: what's all the (Δ)FosB about?".
3190:"A global view of cancer-specific transcript variants by subtractive transcriptome-wide analysis"
3153:
Skotheim RI, Nees M (2007). "Alternative splicing in cancer: noise, functional, or systematic?".
1651:"The Human Adenovirus Type 2 Transcriptome: An Amazing Complexity of Alternatively Spliced mRNAs"
1497:"Calcitonin mRNA polymorphism: peptide switching associated with alternative RNA splicing events"
676:
526:
304:
4525:
Alternative
Splicing of ion channels in the brain, connected to mental and neurological diseases
4103:"The search for alternative splicing regulators: new approaches offer a path to a splicing code"
415:
family. Such proteins contain RNA recognition motifs and arginine and serine-rich (RS) domains.
855:
681:
670:
3539:"Cell motility is controlled by SF2/ASF through alternative splicing of the Ron protooncogene"
2121:"Splicing regulation: from a parts list of regulatory elements to an integrated splicing code"
4874:
1026:
2470:
Izquierdo JM, Majós N, Bonnal S, Martínez C, Castelo R, Guigó R, et al. (August 2005).
1599:"Drosophila Dscam is an axon guidance receptor exhibiting extraordinary molecular diversity"
814:
has been identified as the causal mechanism involved in the induction and maintenance of an
4869:
4208:
4057:
Pan Q, Shai O, Misquitta C, Zhang W, Saltzman AL, Mohammad N, et al. (December 2004).
4013:
3201:
2836:
2777:
2666:
2653:
Romero PR, Zaidi S, Fang YY, Uversky VN, Radivojac P, Oldfield CJ, et al. (May 2006).
2183:
1965:
1720:
1508:
1453:
1272:
694:
665:
477:
322:
765:
One example of a specific splicing variant associated with cancers is in one of the human
716:
Abnormally spliced mRNAs are also found in a high proportion of cancerous cells. Combined
197:: An alternative 5' splice junction (donor site) is used, changing the 3' boundary of the
8:
4688:
1106:"Systematic evaluation of isoform function in literature reports of alternative splicing"
913:
636:
386:
349:
4391:
Louadi Z, Yuan K, Gress A, Tsoy O, Kalinina OV, Baumbach J, et al. (January 2021).
4212:
4152:"A rapid high-throughput method for mapping ribonucleoproteins (RNPs) on human pre-mRNA"
4017:
3205:
2840:
2781:
2670:
2555:
Jacquenet S, Méreau A, Bilodeau PS, Damier L, Stoltzfus CM, Branlant C (November 2001).
2187:
2063:
Reid DC, Chang BL, Gunderson SI, Alpert L, Thompson WA, Fairbrother WG (December 2009).
1969:
1724:
1512:
1457:
1336:
1276:
4468:
4443:
4419:
4393:"DIGGER: exploring the functional role of alternative splicing in protein interactions"
4392:
4368:
4343:
4319:
4292:
4268:
4243:
4176:
4151:
4127:
4102:
4039:
3977:
3952:
3877:
3852:
3828:
3803:
3746:
3694:
3685:
3669:
3650:
3476:
3451:
3427:
3400:
3376:
3349:
3325:
3300:
3281:
3224:
3189:
3130:
3105:
3086:
3038:
3013:
2994:
2951:
2907:
2859:
2824:
2800:
2765:
2738:
2713:
2689:
2654:
2630:
2602:
2449:
2395:
2370:
2251:
2226:
2207:
2148:
2120:
2089:
2064:
2037:
2012:
1988:
1953:
1899:
1874:
1806:
1743:
1708:
1675:
1650:
1628:
1579:
1477:
1426:
1383:
1241:
1213:
1189:
1164:
1145:
1132:
1105:
1086:
1038:
520:
481:
238:
with alternative splicing and provide additional variety in mRNAs derived from a gene.
226:
2302:
2275:
1615:
1598:
1531:
1496:
1295:
1260:
4683:
4473:
4424:
4373:
4324:
4273:
4224:
4181:
4132:
4080:
4031:
3982:
3933:
3882:
3833:
3819:
3780:
3738:
3699:
3642:
3601:
3560:
3516:
3481:
3432:
3381:
3330:
3273:
3229:
3170:
3135:
3078:
3043:
2986:
2943:
2899:
2864:
2805:
2743:
2694:
2635:
2578:
2537:
2493:
2441:
2400:
2351:
2307:
2256:
2199:
2153:
2094:
2042:
1993:
1927:
1904:
1855:
1847:
1842:
1825:
1798:
1748:
1680:
1620:
1571:
1567:
1536:
1495:
Rosenfeld MG, Lin CR, Amara SG, Stolarsky L, Roos BA, Ong ES, Evans RM (March 1982).
1469:
1418:
1414:
1375:
1371:
1340:
1300:
1233:
1229:
1194:
1137:
1078:
1030:
811:
411:, ESE). Most of the activator proteins that bind to ISEs and ESEs are members of the
4534:
3750:
3654:
3619:
Danckwardt S, Neu-Yilik G, Thermann R, Frede U, Hentze MW, Kulozik AE (March 2002).
3285:
3090:
2955:
2911:
2714:"The emerging era of genomic data integration for analyzing splice isoform function"
2453:
1810:
1632:
1583:
1430:
1387:
1149:
1042:
4463:
4455:
4414:
4404:
4363:
4355:
4314:
4304:
4263:
4255:
4216:
4171:
4163:
4122:
4114:
4070:
4043:
4021:
3972:
3964:
3923:
3913:
3872:
3864:
3823:
3815:
3730:
3689:
3681:
3632:
3591:
3550:
3508:
3471:
3463:
3450:
Banaszak LG, Giudice V, Zhao X, Wu Z, Gao S, Hosokawa K, et al. (March 2018).
3422:
3412:
3371:
3361:
3320:
3312:
3263:
3219:
3209:
3162:
3125:
3117:
3070:
3033:
3025:
2998:
2978:
2935:
2891:
2854:
2844:
2795:
2785:
2733:
2725:
2684:
2674:
2625:
2617:
2568:
2527:
2483:
2431:
2390:
2382:
2341:
2297:
2287:
2246:
2238:
2211:
2191:
2143:
2135:
2084:
2076:
2032:
2024:
1983:
1973:
1894:
1886:
1837:
1790:
1738:
1728:
1670:
1662:
1610:
1597:
Schmucker D, Clemens JC, Shu H, Worby CA, Xiao J, Muda M, et al. (June 2000).
1563:
1526:
1516:
1481:
1461:
1410:
1367:
1332:
1290:
1280:
1245:
1225:
1184:
1176:
1127:
1117:
1104:
Bhuiyan SA, Ly S, Phan M, Huntington B, Hogan E, Liu CC, Liu J, Pavlidis P (2018).
1090:
1070:
1022:
3350:"Transcriptome instability as a molecular pan-cancer characteristic of carcinomas"
3074:
854:
More historically, alternatively spliced transcripts have been found by comparing
402:(hnRNPs) such as hnRNPA1 and polypyrimidine tract binding protein (PTB). Splicing
4642:
4625:
4620:
4595:
4509:
4075:
4058:
3734:
3555:
3538:
3214:
3166:
2790:
2488:
2471:
2292:
1733:
864:
770:
750:
654:
649:
are inherited, this provides novel ways for mutations to affect gene expression.
470:
230:
83:
3316:
4806:
4705:
4590:
4585:
4502:
4259:
3868:
3467:
2659:
Proceedings of the
National Academy of Sciences of the United States of America
2028:
1958:
Proceedings of the
National Academy of Sciences of the United States of America
1826:"Intron retention is a major phenomenon in alternative splicing in Arabidopsis"
1501:
Proceedings of the
National Academy of Sciences of the United States of America
1265:
Proceedings of the National Academy of Sciences of the United States of America
1180:
972:
860:
819:
788:
635:
sequence (ESE) overlap. If A1 repressor protein binds to the ESS, it initiates
353:
3512:
3399:
Sveen A, Agesen TH, Nesbakken A, Rognum TO, Lothe RA, Skotheim RI (May 2011).
2729:
2386:
2276:"Single nucleotide polymorphism-based validation of exonic splicing enhancers"
2227:"Positive selection acting on splicing motifs reflects compensatory evolution"
1122:
544:
itself is a regulator of alternative splicing of other sex-related genes (see
4858:
3637:
3620:
3366:
2420:"Expression of CD95 (Fas) in sun-exposed human skin and cutaneous carcinomas"
1851:
871:
796:
317:
site consensus sequence is yUnAy. The branch site is followed by a series of
282:
Spliceosome A complex defines the 5' and 3' ends of the intron before removal
255:
246:
218:
39:
4220:
4026:
4001:
2849:
2679:
1978:
65:
Biologically relevant alternative splicing occurs as a normal phenomenon in
38:. Protein A includes all of the exons, whereas Proteins B and C result from
4652:
4573:
4477:
4428:
4377:
4328:
4277:
4185:
4136:
4084:
4035:
3986:
3953:"Large-scale mapping of branchpoints in human pre-mRNA transcripts in vivo"
3937:
3886:
3837:
3784:
3770:
3742:
3703:
3646:
3605:
3596:
3579:
3564:
3520:
3485:
3436:
3385:
3334:
3277:
3233:
3174:
3139:
3082:
3047:
2990:
2947:
2903:
2868:
2809:
2747:
2698:
2639:
2582:
2573:
2556:
2541:
2532:
2515:
2497:
2445:
2404:
2346:
2329:
2311:
2260:
2203:
2157:
2098:
2046:
1997:
1908:
1859:
1802:
1752:
1684:
1624:
1521:
1285:
1198:
1141:
1082:
1034:
943:
565:
556:
476:
This is an example of exon skipping. The intron upstream from exon 4 has a
374:
273:
58:
30:
4459:
4409:
4359:
4228:
3951:
Taggart AJ, DeSimone AM, Shih JS, Filloux ME, Fairbrother WG (June 2012).
3348:
Sveen A, Johannessen B, Teixeira MR, Lothe RA, Skotheim RI (August 2014).
2355:
2242:
1575:
1540:
1473:
1344:
4831:
4678:
4637:
3918:
3901:
3268:
3251:
3029:
1890:
1666:
1648:
1422:
1379:
1304:
1237:
329:
328:
Splicing of mRNA is performed by an RNA and protein complex known as the
154:
146:
4309:
4118:
3928:
2764:
Eksi R, Li HD, Menon R, Wen Y, Omenn GS, Kretzler M, Guan Y (Nov 2013).
2195:
2080:
1952:
Lim KH, Ferraris L, Filloux ME, Raphael BJ, Fairbrother WG (July 2011).
877:
In microarray analysis, arrays of DNA fragments representing individual
4002:"Predictive identification of exonic splicing enhancers in human genes"
2621:
2603:"Aberrant RNA splicing and its functional consequences in cancer cells"
2436:
2419:
2139:
1709:"A general definition and nomenclature for alternative splicing events"
909:
885:
622:
532:
503:
412:
378:
318:
106:
102:
94:
78:
3968:
3121:
1823:
531:
exon 2 to be included in the processed transcript, including an early
418:
365:
4494:
A General Definition and Nomenclature for Alternative Splicing Events
4293:"Structure-guided isoform identification for the human transcriptome"
3250:
Sveen A, Kilpinen S, Ruusulehto A, Lothe RA, Skotheim RI (May 2016).
1465:
815:
610:
570:
488:
464:
191:: One of two exons is retained in mRNAs after splicing, but not both.
66:
3851:
Luco RF, Allo M, Schor IE, Kornblihtt AR, Misteli T (January 2011).
2418:
Filipowicz E, Adegboyega P, Sanchez RL, Gatalica Z (February 2002).
1794:
601:
2417:
1074:
709:
299:
182:
162:
Five basic modes of alternative splicing are generally recognized.
3417:
2982:
2939:
2924:
2895:
807:
4798:
4778:
4773:
4444:"APPRIS: annotation of principal and alternative splice isoforms"
4441:
4167:
962:
937:
848:
755:
717:
241:
214:
98:
35:
3618:
1165:"Alternative splicing may not be the key to proteome complexity"
4753:
4748:
4728:
4664:
4615:
4497:
2274:
Fairbrother WG, Holste D, Burge CB, Sharp PA (September 2004).
1261:"Spliced segments at the 5' terminus of adenovirus 2 late mRNA"
949:
737:
727:
446:
291:
3578:
Hui L, Zhang X, Wu X, Lin Z, Wang Q, Li Y, Hu G (April 2004).
2371:"The role of U2AF35 and U2AF65 in enhancer-dependent splicing"
4821:
4816:
4738:
4733:
4723:
4718:
4713:
4673:
4493:
4241:
3764:
3347:
3188:
He C, Zhou F, Zuo Z, Cheng H, Zhou R (2009). Bauer JA (ed.).
3060:
2554:
1707:
Sammeth M, Foissac S, Guigó R (August 2008). Brent MR (ed.).
897:
829:
783:
540:
341:
333:
122:
4519:
4242:
Chang B, Levin J, Thompson WA, Fairbrother WG (March 2010).
3249:
3155:
The International Journal of Biochemistry & Cell Biology
2469:
589:
Exon 6 contains a pyrimidine-rich exonic splicing silencer,
278:
233:
sites. Use of multiple promoters is properly described as a
4836:
4826:
4668:
4524:
3804:"Natural rewards, neuroplasticity, and non-drug addictions"
3795:
3536:
3014:"Different levels of alternative splicing among eukaryotes"
2968:
2273:
901:
878:
803:
766:
626:
574:
309:
295:
174:
4000:
Fairbrother WG, Yeh RF, Sharp PA, Burge CB (August 2002).
3950:
3899:
3398:
1553:
1494:
245:
Schematic cutoff from 3 splicing structures in the murine
4341:
4149:
2513:
1951:
618:
287:
86:
sites, giving different 3' ends for the processed mRNAs.
3999:
2514:
Zahler AM, Damgaard CK, Kjems J, Caputi M (March 2004).
2062:
1596:
1211:
900:) have been used. The array is then probed with labeled
89:
In 1981, the first example of alternative splicing in a
4290:
4248:
Combinatorial Chemistry & High Throughput Screening
3850:
2013:"Role of RNA structure in regulating pre-mRNA splicing"
1103:
377:
proteins (repressors and activators) and corresponding
345:
nomenclature derives from their high uridine content).
337:
181:
or retained. This is the most common mode in mammalian
125:, which could potentially have 38,016 splice variants.
4150:
Watkins KH, Stewart A, Fairbrother W (December 2009).
3146:
2652:
2368:
1008:"Mechanisms of alternative pre-messenger RNA splicing"
302:, the mean is 4–5 exons and introns; in the fruit fly
27:
Process by which a gene can code for multiple proteins
4564:
4056:
3449:
1872:
1443:
745:
as compared to their isogenic wildtype counterparts.
77:
Alternative splicing was first observed in 1977. The
4515:
IsoPred: computationally predicted isoform functions
2823:
Irimia M, Rukov JL, Penny D, Roy SW (October 2007).
1162:
802:
Overexpression of a truncated splice variant of the
4390:
2822:
2173:
2058:
2056:
1706:
360:
1873:Gao K, Masuda A, Matsuura T, Ohno K (April 2008).
1322:
3773:Annals of Agricultural and Environmental Medicine
560:Alternative splicing of the Fas receptor pre-mRNA
4856:
2712:Li HD, Menon R, Omenn GS, Guan Y (August 2014).
2711:
2600:
2465:
2463:
2369:Graveley BR, Hertel KJ, Maniatis T (June 2001).
2053:
1875:"Human branch point consensus sequence is yUnAy"
1780:
1258:
769:genes. Three DNMT genes encode enzymes that add
630:differentially spliced transcripts contains the
577:and polypyrimidine tract-binding protein (PTB).
551:
435:
286:When the pre-mRNA has been transcribed from the
3900:Strømme JM, Johannessen B, Skotheim RI (2023).
3714:
3853:"Epigenetics in alternative pre-mRNA splicing"
3723:The American Journal of Drug and Alcohol Abuse
3661:
3612:
3298:
3187:
2763:
1060:
259:splicing patterns has recently been proposed.
97:gene was characterized. The gene encoding the
4550:
3577:
3532:
3530:
3443:
3245:
3243:
2596:
2594:
2592:
2509:
2507:
2460:
2327:
2224:
1644:
1642:
1400:
968:Polyadenylation § Alternative polyadenylation
810:– in a specific population of neurons in the
684:(EST) each from human, mouse, rat, cow, fly (
267:
4530:BIPASS: Web Services in Alternative Splicing
3571:
3152:
2548:
2010:
1259:Berget SM, Moore C, Sharp PA (August 1977).
34:Alternative splicing produces three protein
4100:
3498:
3054:
3011:
1547:
1488:
1437:
4557:
4543:
3527:
3392:
3341:
3240:
3103:
2881:
2589:
2504:
2411:
1702:
1700:
1698:
1696:
1694:
1639:
1357:
888:exon microarray) or exon/exon boundaries (
830:Genome-scale (transcriptome-wide) analysis
674:, and only about twice as many as the fly
4467:
4418:
4408:
4367:
4318:
4308:
4267:
4198:
4175:
4126:
4074:
4025:
3976:
3957:Nature Structural & Molecular Biology
3927:
3917:
3876:
3827:
3693:
3636:
3595:
3554:
3475:
3426:
3416:
3375:
3365:
3324:
3267:
3223:
3213:
3129:
3037:
2858:
2848:
2799:
2789:
2737:
2688:
2678:
2629:
2572:
2531:
2487:
2435:
2394:
2345:
2301:
2291:
2250:
2169:
2167:
2147:
2118:
2088:
2036:
1987:
1977:
1898:
1841:
1776:
1774:
1772:
1770:
1768:
1766:
1764:
1762:
1742:
1732:
1674:
1614:
1590:
1530:
1520:
1394:
1351:
1294:
1284:
1188:
1131:
1121:
1056:
1054:
1052:
863:-based analyses, RNA-binding assays, and
4096:
4094:
3844:
3670:"Cellular basis of memory for addiction"
2918:
2362:
2323:
2321:
2225:Ke S, Zhang XH, Chasin LA (April 2008).
1318:
1316:
1314:
1163:Tress ML, Abascal F, Valencia A (2017).
1027:10.1146/annurev.biochem.72.121801.161720
643:
614:Alternative splicing of HIV-1 tat exon 2
609:
555:
502:
445:
417:
400:heterogeneous nuclear ribonucleoproteins
364:
277:
240:
153:
145:
29:
4050:
3893:
3667:
3492:
3299:Omenn GS, Guan Y, Menon R (July 2014).
2759:
2757:
2114:
2112:
2110:
2108:
1691:
1001:
999:
997:
995:
993:
991:
989:
987:
731:has been demonstrated to contribute to
602:Repressor-activator competition: HIV-1
14:
4857:
3720:
3181:
2816:
2646:
2164:
1947:
1945:
1943:
1926:. Amsterdam: Elsevier Academic Press.
1783:Nature Reviews. Molecular Cell Biology
1759:
1049:
4538:
4101:David CJ, Manley JL (February 2008).
4091:
3801:
3456:Blood Cells, Molecules & Diseases
2875:
2318:
1921:
1915:
1311:
1005:
713:splicing-related diseases is cancer.
2754:
2328:Lynch KW, Maniatis T (August 1996).
2105:
984:
3104:Ward AJ, Cooper TA (January 2010).
3005:
2561:The Journal of Biological Chemistry
2520:The Journal of Biological Chemistry
2011:Warf MB, Berglund JA (March 2010).
1940:
1866:
1337:10.1146/annurev.bi.55.070186.005303
659:Intrinsically unstructured proteins
24:
3674:Dialogues in Clinical Neuroscience
2218:
177:may be spliced out of the primary
25:
4886:
4566:Post-transcriptional modification
4487:
4156:Journal of Visualized Experiments
2601:Fackenthal JD, Godley LA (2008).
3820:10.1016/j.neuropharm.2011.03.010
3686:10.31887/DCNS.2013.15.4/enestler
1843:10.1111/j.1365-313X.2004.02172.x
1212:Chow LT, Gelinas RE, Broker TR,
361:Regulatory elements and proteins
4435:
4384:
4335:
4284:
4235:
4192:
4143:
3993:
3944:
3906:JCO Clinical Cancer Informatics
3292:
3097:
2962:
2705:
2610:Disease Models & Mechanisms
2267:
2004:
1817:
112:calcitonin gene related peptide
3106:"The pathobiology of splicing"
3012:Kim E, Magen A, Ast G (2006).
2017:Trends in Biochemical Sciences
1252:
1205:
1169:Trends in Biochemical Sciences
1156:
1097:
13:
1:
3075:10.1016/j.febslet.2005.02.047
2119:Wang Z, Burge CB (May 2008).
1616:10.1016/S0092-8674(00)80878-8
1325:Annual Review of Biochemistry
1015:Annual Review of Biochemistry
978:
552:Exon definition: Fas receptor
262:
4076:10.1016/j.molcel.2004.12.004
3735:10.3109/00952990.2014.933840
3668:Nestler EJ (December 2013).
3556:10.1016/j.molcel.2005.10.026
3215:10.1371/journal.pone.0004732
3167:10.1016/j.biocel.2007.02.016
2791:10.1371/journal.pcbi.1003314
2489:10.1016/j.molcel.2005.06.015
2293:10.1371/journal.pbio.0020268
1734:10.1371/journal.pcbi.1000147
1568:10.1016/0092-8674(81)90325-1
1415:10.1016/0092-8674(78)90071-5
1372:10.1016/0092-8674(78)90019-3
1230:10.1016/0092-8674(77)90180-5
927:
760:nonsense-mediated mRNA decay
741:-mutated cell lines exhibit
507:Alternative splicing of the
489:Alternative acceptor sites:
325:– then by AG at the 3' end.
72:
7:
3317:10.1016/j.jprot.2014.04.012
956:
847:detection of the transient
430:
10:
4891:
4454:(Database issue): D110-7.
4260:10.2174/138620710790980522
3869:10.1016/j.cell.2010.11.056
3802:Olsen CM (December 2011).
3468:10.1016/j.bcmd.2017.12.002
2770:PLOS Computational Biology
2029:10.1016/j.tibs.2009.10.004
1713:PLOS Computational Biology
1181:10.1016/j.tibs.2016.08.008
841:multiple instance learning
702:
271:
268:General splicing mechanism
235:transcriptional regulation
4797:
4704:
4660:
4651:
4581:
4572:
3513:10.1016/j.tig.2007.10.001
2730:10.1016/j.tig.2014.05.005
2387:10.1017/S1355838201010317
1123:10.1186/s12864-018-5013-2
743:transcriptome instability
723:transcriptome instability
564:Multiple isoforms of the
409:exonic splicing enhancers
396:exonic splicing silencers
373:Splicing is regulated by
205:Alternative acceptor site
3638:10.1182/blood.V99.5.1811
3367:10.1186/1471-2164-15-672
3110:The Journal of Pathology
2829:BMC Evolutionary Biology
733:hematologic malignancies
450:Alternative splicing of
189:Mutually exclusive exons
141:
51:alternative RNA splicing
4633:Poly(A)-binding protein
4221:10.1126/science.2200121
4107:Genes & Development
4027:10.1126/science.1073774
2850:10.1186/1471-2148-7-188
2680:10.1073/pnas.0507916103
2334:Genes & Development
1979:10.1073/pnas.1101135108
777:Another example is the
682:expressed sequence tags
677:Drosophila melanogaster
527:Drosophila melanogaster
480:that doesn't match the
4448:Nucleic Acids Research
4397:Nucleic Acids Research
3597:10.1038/sj.onc.1207362
3018:Nucleic Acids Research
2574:10.1074/jbc.M104070200
2533:10.1074/jbc.M312743200
2347:10.1101/gad.10.16.2089
1879:Nucleic Acids Research
1522:10.1073/pnas.79.6.1717
1286:10.1073/pnas.74.8.3171
671:Caenorhabditis elegans
664:Research based on the
615:
561:
515:
455:
423:
370:
290:, it includes several
283:
250:
195:Alternative donor site
159:
151:
43:
4360:10.1101/gr.220962.117
3305:Journal of Proteomics
2243:10.1101/gr.070268.107
644:Adaptive significance
613:
559:
506:
449:
421:
368:
281:
244:
157:
149:
55:differential splicing
33:
18:Alternatively spliced
4696:Alternative splicing
3919:10.1200/CCI.22.00159
3269:10.1038/onc.2015.318
1667:10.1128/JVI.01869-20
695:Arabidopsis thaliana
666:Human Genome Project
478:polypyrimidine tract
387:RNA-binding proteins
323:polypyrimidine tract
57:, is an alternative
47:Alternative splicing
4460:10.1093/nar/gks1058
4410:10.1093/nar/gkaa768
4310:10.7554/eLife.82556
4213:1990Sci...249..505T
4119:10.1101/gad.1643108
4018:2002Sci...297.1007F
3206:2009PLoSO...4.4732H
2841:2007BMCEE...7..188I
2782:2013PLSCB...9E3314E
2671:2006PNAS..103.8390R
2196:10.1038/nature09000
2188:2010Natur.465...53B
2081:10.1261/rna.1821809
1970:2011PNAS..10811093H
1725:2008PLSCB...4E0147S
1655:Journal of Virology
1513:1982PNAS...79.1717R
1458:1981Natur.290...63R
1277:1977PNAS...74.3171B
914:immunoprecipitation
637:cooperative binding
458:Pre-mRNAs from the
422:Splicing activation
369:Splicing repression
350:transesterification
173:: in this case, an
4807:5′ cap methylation
4508:2009-12-11 at the
3501:Trends in Genetics
3030:10.1093/nar/gkl924
2718:Trends in Genetics
2622:10.1242/dmm.000331
2437:10.1002/cncr.10277
2140:10.1261/rna.876308
1891:10.1093/nar/gkn073
616:
562:
516:
482:consensus sequence
456:
424:
371:
284:
251:
160:
152:
44:
4852:
4851:
4793:
4792:
4789:
4788:
4706:pre-mRNA factors
4403:(D1): D309–D318.
4354:(10): 1759–1768.
4012:(5583): 1007–13.
3969:10.1038/nsmb.2327
3808:Neuropharmacology
3122:10.1002/path.2649
1933:978-0-12-175551-5
1924:Molecular biology
1830:The Plant Journal
1006:Black DL (2003).
812:nucleus accumbens
692:), and the plant
518:Pre-mRNAs of the
16:(Redirected from
4882:
4658:
4657:
4591:5′ cap formation
4579:
4578:
4559:
4552:
4545:
4536:
4535:
4482:
4481:
4471:
4439:
4433:
4432:
4422:
4412:
4388:
4382:
4381:
4371:
4339:
4333:
4332:
4322:
4312:
4288:
4282:
4281:
4271:
4239:
4233:
4232:
4207:(4968): 505–10.
4196:
4190:
4189:
4179:
4147:
4141:
4140:
4130:
4098:
4089:
4088:
4078:
4054:
4048:
4047:
4029:
3997:
3991:
3990:
3980:
3948:
3942:
3941:
3931:
3921:
3897:
3891:
3890:
3880:
3848:
3842:
3841:
3831:
3799:
3793:
3792:
3768:
3762:
3761:
3757:molecular switch
3718:
3712:
3711:
3697:
3665:
3659:
3658:
3640:
3616:
3610:
3609:
3599:
3575:
3569:
3568:
3558:
3534:
3525:
3524:
3496:
3490:
3489:
3479:
3447:
3441:
3440:
3430:
3420:
3396:
3390:
3389:
3379:
3369:
3345:
3339:
3338:
3328:
3296:
3290:
3289:
3271:
3247:
3238:
3237:
3227:
3217:
3185:
3179:
3178:
3161:(7–8): 1432–49.
3150:
3144:
3143:
3133:
3101:
3095:
3094:
3058:
3052:
3051:
3041:
3009:
3003:
3002:
2966:
2960:
2959:
2922:
2916:
2915:
2879:
2873:
2872:
2862:
2852:
2820:
2814:
2813:
2803:
2793:
2776:(11): e1003314.
2761:
2752:
2751:
2741:
2709:
2703:
2702:
2692:
2682:
2650:
2644:
2643:
2633:
2607:
2606:(Free full text)
2598:
2587:
2586:
2576:
2567:(44): 40464–75.
2552:
2546:
2545:
2535:
2526:(11): 10077–84.
2511:
2502:
2501:
2491:
2467:
2458:
2457:
2439:
2415:
2409:
2408:
2398:
2366:
2360:
2359:
2349:
2340:(16): 2089–101.
2325:
2316:
2315:
2305:
2295:
2271:
2265:
2264:
2254:
2222:
2216:
2215:
2171:
2162:
2161:
2151:
2125:
2124:(Free full text)
2116:
2103:
2102:
2092:
2060:
2051:
2050:
2040:
2008:
2002:
2001:
1991:
1981:
1949:
1938:
1937:
1922:Clark D (2005).
1919:
1913:
1912:
1902:
1870:
1864:
1863:
1845:
1821:
1815:
1814:
1778:
1757:
1756:
1746:
1736:
1704:
1689:
1688:
1678:
1646:
1637:
1636:
1618:
1594:
1588:
1587:
1551:
1545:
1544:
1534:
1524:
1492:
1486:
1485:
1466:10.1038/290063a0
1441:
1435:
1434:
1398:
1392:
1391:
1355:
1349:
1348:
1320:
1309:
1308:
1298:
1288:
1256:
1250:
1249:
1209:
1203:
1202:
1192:
1160:
1154:
1153:
1135:
1125:
1101:
1095:
1094:
1058:
1047:
1046:
1012:
1003:
498:
497:
211:Intron retention
21:
4890:
4889:
4885:
4884:
4883:
4881:
4880:
4879:
4865:Gene expression
4855:
4854:
4853:
4848:
4785:
4700:
4647:
4643:Polyuridylation
4596:Polyadenylation
4568:
4563:
4510:Wayback Machine
4490:
4485:
4440:
4436:
4389:
4385:
4348:Genome Research
4340:
4336:
4289:
4285:
4240:
4236:
4197:
4193:
4148:
4144:
4099:
4092:
4055:
4051:
3998:
3994:
3949:
3945:
3912:(7): e2200159.
3898:
3894:
3849:
3845:
3800:
3796:
3769:
3765:
3719:
3715:
3666:
3662:
3617:
3613:
3590:(17): 3013–23.
3576:
3572:
3535:
3528:
3497:
3493:
3448:
3444:
3405:Genome Medicine
3397:
3393:
3346:
3342:
3297:
3293:
3262:(19): 2413–27.
3248:
3241:
3186:
3182:
3151:
3147:
3102:
3098:
3059:
3055:
3010:
3006:
2971:Nature Genetics
2967:
2963:
2928:Nature Genetics
2923:
2919:
2884:Nature Genetics
2880:
2876:
2821:
2817:
2762:
2755:
2710:
2706:
2651:
2647:
2605:
2599:
2590:
2553:
2549:
2512:
2505:
2468:
2461:
2416:
2412:
2367:
2363:
2326:
2319:
2272:
2268:
2231:Genome Research
2223:
2219:
2172:
2165:
2123:
2117:
2106:
2075:(12): 2385–97.
2061:
2054:
2009:
2005:
1964:(27): 11093–8.
1950:
1941:
1934:
1920:
1916:
1871:
1867:
1822:
1818:
1795:10.1038/nrm1645
1779:
1760:
1719:(8): e1000147.
1705:
1692:
1647:
1640:
1595:
1591:
1552:
1548:
1493:
1489:
1442:
1438:
1399:
1395:
1356:
1352:
1331:(1): 1091–117.
1321:
1312:
1257:
1253:
1210:
1206:
1161:
1157:
1102:
1098:
1063:Nature Genetics
1059:
1050:
1010:
1004:
985:
981:
959:
930:
865:deep sequencing
832:
820:natural rewards
751:phosphorylation
705:
686:D. melanogaster
655:protein isoform
646:
608:
554:
501:
495:
494:
471:polyadenylation
460:D. melanogaster
444:
436:Exon skipping:
433:
363:
336:designated U1,
276:
270:
265:
253:
231:polyadenylation
144:
119:D. melanogaster
93:from a normal,
84:polyadenylation
75:
28:
23:
22:
15:
12:
11:
5:
4888:
4878:
4877:
4872:
4867:
4850:
4849:
4847:
4846:
4845:
4844:
4839:
4834:
4829:
4824:
4819:
4812:mRNA decapping
4809:
4803:
4801:
4795:
4794:
4791:
4790:
4787:
4786:
4784:
4783:
4782:
4781:
4776:
4771:
4766:
4761:
4756:
4751:
4746:
4741:
4736:
4731:
4726:
4721:
4710:
4708:
4702:
4701:
4699:
4698:
4693:
4692:
4691:
4686:
4676:
4671:
4661:
4655:
4649:
4648:
4646:
4645:
4640:
4635:
4630:
4629:
4628:
4623:
4618:
4613:
4608:
4603:
4593:
4588:
4586:Precursor mRNA
4582:
4576:
4570:
4569:
4562:
4561:
4554:
4547:
4539:
4533:
4532:
4527:
4522:
4517:
4512:
4500:
4489:
4488:External links
4486:
4484:
4483:
4434:
4383:
4334:
4283:
4234:
4191:
4142:
4090:
4063:Molecular Cell
4049:
3992:
3943:
3892:
3843:
3814:(7): 1109–22.
3794:
3763:
3713:
3660:
3611:
3570:
3543:Molecular Cell
3526:
3491:
3442:
3391:
3340:
3291:
3239:
3180:
3145:
3096:
3053:
3004:
2961:
2917:
2874:
2815:
2753:
2704:
2665:(22): 8390–5.
2645:
2588:
2547:
2503:
2476:Molecular Cell
2459:
2410:
2361:
2317:
2266:
2217:
2182:(7294): 53–9.
2163:
2104:
2052:
2003:
1939:
1932:
1914:
1885:(7): 2257–67.
1865:
1836:(6): 877–885.
1816:
1758:
1690:
1638:
1589:
1546:
1507:(6): 1717–21.
1487:
1452:(5801): 63–5.
1436:
1409:(4): 1477–93.
1393:
1366:(2): 497–510.
1350:
1310:
1251:
1204:
1155:
1096:
1075:10.1038/ng.259
1069:(12): 1413–5.
1048:
1021:(1): 291–336.
982:
980:
977:
976:
975:
973:Trans-splicing
970:
965:
958:
955:
954:
953:
947:
941:
929:
926:
861:DNA microarray
831:
828:
789:proto-oncogene
704:
701:
645:
642:
607:
600:
595:
594:
587:
583:
553:
550:
524:(Tra) gene of
500:
487:
443:
434:
432:
429:
362:
359:
354:phosphodiester
272:Main article:
269:
266:
264:
261:
223:
222:
208:
202:
192:
186:
143:
140:
139:
138:
137:
136:
74:
71:
26:
9:
6:
4:
3:
2:
4887:
4876:
4873:
4871:
4868:
4866:
4863:
4862:
4860:
4843:
4840:
4838:
4835:
4833:
4830:
4828:
4825:
4823:
4820:
4818:
4815:
4814:
4813:
4810:
4808:
4805:
4804:
4802:
4800:
4796:
4780:
4777:
4775:
4772:
4770:
4767:
4765:
4762:
4760:
4757:
4755:
4752:
4750:
4747:
4745:
4742:
4740:
4737:
4735:
4732:
4730:
4727:
4725:
4722:
4720:
4717:
4716:
4715:
4712:
4711:
4709:
4707:
4703:
4697:
4694:
4690:
4687:
4685:
4682:
4681:
4680:
4677:
4675:
4672:
4670:
4666:
4663:
4662:
4659:
4656:
4654:
4650:
4644:
4641:
4639:
4636:
4634:
4631:
4627:
4624:
4622:
4619:
4617:
4614:
4612:
4609:
4607:
4604:
4602:
4599:
4598:
4597:
4594:
4592:
4589:
4587:
4584:
4583:
4580:
4577:
4575:
4571:
4567:
4560:
4555:
4553:
4548:
4546:
4541:
4540:
4537:
4531:
4528:
4526:
4523:
4521:
4518:
4516:
4513:
4511:
4507:
4504:
4501:
4499:
4495:
4492:
4491:
4479:
4475:
4470:
4465:
4461:
4457:
4453:
4449:
4445:
4438:
4430:
4426:
4421:
4416:
4411:
4406:
4402:
4398:
4394:
4387:
4379:
4375:
4370:
4365:
4361:
4357:
4353:
4349:
4345:
4338:
4330:
4326:
4321:
4316:
4311:
4306:
4302:
4298:
4294:
4287:
4279:
4275:
4270:
4265:
4261:
4257:
4254:(3): 242–52.
4253:
4249:
4245:
4238:
4230:
4226:
4222:
4218:
4214:
4210:
4206:
4202:
4195:
4187:
4183:
4178:
4173:
4169:
4165:
4161:
4157:
4153:
4146:
4138:
4134:
4129:
4124:
4120:
4116:
4113:(3): 279–85.
4112:
4108:
4104:
4097:
4095:
4086:
4082:
4077:
4072:
4069:(6): 929–41.
4068:
4064:
4060:
4053:
4045:
4041:
4037:
4033:
4028:
4023:
4019:
4015:
4011:
4007:
4003:
3996:
3988:
3984:
3979:
3974:
3970:
3966:
3963:(7): 719–21.
3962:
3958:
3954:
3947:
3939:
3935:
3930:
3925:
3920:
3915:
3911:
3907:
3903:
3896:
3888:
3884:
3879:
3874:
3870:
3866:
3862:
3858:
3854:
3847:
3839:
3835:
3830:
3825:
3821:
3817:
3813:
3809:
3805:
3798:
3791:
3786:
3782:
3778:
3774:
3767:
3760:
3758:
3752:
3748:
3744:
3740:
3736:
3732:
3729:(6): 428–37.
3728:
3724:
3717:
3710:
3705:
3701:
3696:
3691:
3687:
3683:
3680:(4): 431–43.
3679:
3675:
3671:
3664:
3656:
3652:
3648:
3644:
3639:
3634:
3631:(5): 1811–6.
3630:
3626:
3622:
3615:
3607:
3603:
3598:
3593:
3589:
3585:
3581:
3574:
3566:
3562:
3557:
3552:
3549:(6): 881–90.
3548:
3544:
3540:
3533:
3531:
3522:
3518:
3514:
3510:
3506:
3502:
3495:
3487:
3483:
3478:
3473:
3469:
3465:
3461:
3457:
3453:
3446:
3438:
3434:
3429:
3424:
3419:
3418:10.1186/gm248
3414:
3410:
3406:
3402:
3395:
3387:
3383:
3378:
3373:
3368:
3363:
3359:
3355:
3351:
3344:
3336:
3332:
3327:
3322:
3318:
3314:
3310:
3306:
3302:
3295:
3287:
3283:
3279:
3275:
3270:
3265:
3261:
3257:
3253:
3246:
3244:
3235:
3231:
3226:
3221:
3216:
3211:
3207:
3203:
3199:
3195:
3191:
3184:
3176:
3172:
3168:
3164:
3160:
3156:
3149:
3141:
3137:
3132:
3127:
3123:
3119:
3116:(2): 152–63.
3115:
3111:
3107:
3100:
3092:
3088:
3084:
3080:
3076:
3072:
3069:(9): 1900–3.
3068:
3064:
3057:
3049:
3045:
3040:
3035:
3031:
3027:
3024:(1): 125–31.
3023:
3019:
3015:
3008:
3000:
2996:
2992:
2988:
2984:
2983:10.1038/ng803
2980:
2976:
2972:
2965:
2957:
2953:
2949:
2945:
2941:
2940:10.1038/76118
2937:
2933:
2929:
2921:
2913:
2909:
2905:
2901:
2897:
2896:10.1038/76115
2893:
2889:
2885:
2878:
2870:
2866:
2861:
2856:
2851:
2846:
2842:
2838:
2834:
2830:
2826:
2819:
2811:
2807:
2802:
2797:
2792:
2787:
2783:
2779:
2775:
2771:
2767:
2760:
2758:
2749:
2745:
2740:
2735:
2731:
2727:
2723:
2719:
2715:
2708:
2700:
2696:
2691:
2686:
2681:
2676:
2672:
2668:
2664:
2660:
2656:
2649:
2641:
2637:
2632:
2627:
2623:
2619:
2615:
2611:
2604:
2597:
2595:
2593:
2584:
2580:
2575:
2570:
2566:
2562:
2558:
2551:
2543:
2539:
2534:
2529:
2525:
2521:
2517:
2510:
2508:
2499:
2495:
2490:
2485:
2482:(4): 475–84.
2481:
2477:
2473:
2466:
2464:
2455:
2451:
2447:
2443:
2438:
2433:
2429:
2425:
2421:
2414:
2406:
2402:
2397:
2392:
2388:
2384:
2381:(6): 806–18.
2380:
2376:
2372:
2365:
2357:
2353:
2348:
2343:
2339:
2335:
2331:
2324:
2322:
2313:
2309:
2304:
2299:
2294:
2289:
2285:
2281:
2277:
2270:
2262:
2258:
2253:
2248:
2244:
2240:
2237:(4): 533–43.
2236:
2232:
2228:
2221:
2213:
2209:
2205:
2201:
2197:
2193:
2189:
2185:
2181:
2177:
2170:
2168:
2159:
2155:
2150:
2145:
2141:
2137:
2134:(5): 802–13.
2133:
2129:
2122:
2115:
2113:
2111:
2109:
2100:
2096:
2091:
2086:
2082:
2078:
2074:
2070:
2066:
2059:
2057:
2048:
2044:
2039:
2034:
2030:
2026:
2023:(3): 169–78.
2022:
2018:
2014:
2007:
1999:
1995:
1990:
1985:
1980:
1975:
1971:
1967:
1963:
1959:
1955:
1948:
1946:
1944:
1935:
1929:
1925:
1918:
1910:
1906:
1901:
1896:
1892:
1888:
1884:
1880:
1876:
1869:
1861:
1857:
1853:
1849:
1844:
1839:
1835:
1831:
1827:
1820:
1812:
1808:
1804:
1800:
1796:
1792:
1789:(5): 386–98.
1788:
1784:
1777:
1775:
1773:
1771:
1769:
1767:
1765:
1763:
1754:
1750:
1745:
1740:
1735:
1730:
1726:
1722:
1718:
1714:
1710:
1703:
1701:
1699:
1697:
1695:
1686:
1682:
1677:
1672:
1668:
1664:
1660:
1656:
1652:
1645:
1643:
1634:
1630:
1626:
1622:
1617:
1612:
1609:(6): 671–84.
1608:
1604:
1600:
1593:
1585:
1581:
1577:
1573:
1569:
1565:
1562:(2): 353–65.
1561:
1557:
1550:
1542:
1538:
1533:
1528:
1523:
1518:
1514:
1510:
1506:
1502:
1498:
1491:
1483:
1479:
1475:
1471:
1467:
1463:
1459:
1455:
1451:
1447:
1440:
1432:
1428:
1424:
1420:
1416:
1412:
1408:
1404:
1397:
1389:
1385:
1381:
1377:
1373:
1369:
1365:
1361:
1354:
1346:
1342:
1338:
1334:
1330:
1326:
1319:
1317:
1315:
1306:
1302:
1297:
1292:
1287:
1282:
1278:
1274:
1271:(8): 3171–5.
1270:
1266:
1262:
1255:
1247:
1243:
1239:
1235:
1231:
1227:
1223:
1219:
1215:
1208:
1200:
1196:
1191:
1186:
1182:
1178:
1175:(2): 98–110.
1174:
1170:
1166:
1159:
1151:
1147:
1143:
1139:
1134:
1129:
1124:
1119:
1115:
1111:
1107:
1100:
1092:
1088:
1084:
1080:
1076:
1072:
1068:
1064:
1057:
1055:
1053:
1044:
1040:
1036:
1032:
1028:
1024:
1020:
1016:
1009:
1002:
1000:
998:
996:
994:
992:
990:
988:
983:
974:
971:
969:
966:
964:
961:
960:
951:
948:
945:
942:
939:
936:
935:
934:
925:
921:
917:
915:
911:
910:Cross-linking
906:
903:
899:
895:
891:
887:
884:
880:
875:
873:
872:reporter gene
870:
866:
862:
857:
852:
850:
846:
842:
836:
827:
823:
821:
818:to drugs and
817:
813:
809:
805:
800:
798:
797:cell motility
794:
790:
786:
785:
780:
775:
772:
768:
763:
761:
757:
752:
746:
744:
740:
739:
734:
730:
729:
724:
719:
714:
711:
700:
697:
696:
691:
687:
683:
679:
678:
673:
672:
667:
662:
660:
656:
650:
641:
638:
633:
628:
624:
620:
612:
605:
599:
592:
588:
584:
580:
579:
578:
576:
572:
567:
558:
549:
547:
542:
538:
534:
529:
528:
523:
522:
514:gene product.
513:
510:
505:
499:
492:
486:
483:
479:
474:
472:
467:
466:
461:
453:
448:
442:
439:
428:
420:
416:
414:
410:
405:
401:
397:
392:
388:
383:
380:
376:
367:
358:
355:
351:
346:
343:
339:
335:
332:, containing
331:
326:
324:
320:
314:
311:
307:
306:
301:
297:
293:
289:
280:
275:
260:
257:
256:hyaluronidase
248:
247:hyaluronidase
243:
239:
236:
232:
229:and multiple
228:
220:
219:reading frame
216:
212:
209:
206:
203:
200:
196:
193:
190:
187:
184:
180:
176:
172:
171:cassette exon
168:
167:Exon skipping
165:
164:
163:
156:
148:
134:
133:
132:
131:
130:
126:
124:
120:
115:
113:
108:
104:
100:
96:
92:
87:
85:
80:
70:
68:
63:
60:
56:
52:
48:
41:
40:exon skipping
37:
32:
19:
4875:RNA splicing
4695:
4667: /
4653:RNA splicing
4451:
4447:
4437:
4400:
4396:
4386:
4351:
4347:
4337:
4300:
4296:
4286:
4251:
4247:
4237:
4204:
4200:
4194:
4168:10.3791/1622
4162:(34): 1622.
4159:
4155:
4145:
4110:
4106:
4066:
4062:
4052:
4009:
4005:
3995:
3960:
3956:
3946:
3929:10852/108838
3909:
3905:
3895:
3863:(1): 16–26.
3860:
3856:
3846:
3811:
3807:
3797:
3788:
3779:(3): 491–6.
3776:
3772:
3766:
3756:
3754:
3726:
3722:
3716:
3707:
3677:
3673:
3663:
3628:
3624:
3614:
3587:
3583:
3573:
3546:
3542:
3504:
3500:
3494:
3459:
3455:
3445:
3408:
3404:
3394:
3357:
3354:BMC Genomics
3353:
3343:
3308:
3304:
3294:
3259:
3255:
3200:(3): e4732.
3197:
3193:
3183:
3158:
3154:
3148:
3113:
3109:
3099:
3066:
3063:FEBS Letters
3062:
3056:
3021:
3017:
3007:
2977:(1): 29–30.
2974:
2970:
2964:
2934:(2): 235–8.
2931:
2927:
2920:
2890:(2): 232–4.
2887:
2883:
2877:
2832:
2828:
2818:
2773:
2769:
2724:(8): 340–7.
2721:
2717:
2707:
2662:
2658:
2648:
2616:(1): 37–42.
2613:
2609:
2564:
2560:
2550:
2523:
2519:
2479:
2475:
2430:(3): 814–9.
2427:
2423:
2413:
2378:
2374:
2364:
2337:
2333:
2283:
2280:PLOS Biology
2279:
2269:
2234:
2230:
2220:
2179:
2175:
2131:
2127:
2072:
2068:
2020:
2016:
2006:
1961:
1957:
1923:
1917:
1882:
1878:
1868:
1833:
1829:
1819:
1786:
1782:
1716:
1712:
1658:
1654:
1606:
1602:
1592:
1559:
1555:
1549:
1504:
1500:
1490:
1449:
1445:
1439:
1406:
1402:
1396:
1363:
1359:
1353:
1328:
1324:
1268:
1264:
1254:
1221:
1217:
1207:
1172:
1168:
1158:
1113:
1110:BMC Genomics
1109:
1099:
1066:
1062:
1018:
1014:
944:Intronerator
931:
922:
918:
907:
892:arrays from
889:
882:
876:
868:
853:
844:
837:
833:
824:
801:
792:
782:
778:
776:
764:
747:
736:
726:
715:
706:
693:
689:
685:
675:
669:
663:
651:
647:
631:
625:that causes
617:
603:
596:
590:
566:Fas receptor
563:
545:
525:
519:
517:
511:
508:
493:
490:
475:
463:
459:
457:
451:
440:
437:
425:
403:
390:
384:
375:trans-acting
372:
347:
327:
315:
303:
285:
274:RNA splicing
252:
224:
210:
204:
194:
188:
170:
166:
161:
127:
121:gene called
118:
116:
88:
76:
64:
54:
50:
46:
45:
4870:Spliceosome
4679:Spliceosome
4638:RNA editing
3507:(1): 7–10.
2286:(9): E268.
735:, and that
521:Transformer
512:Transformer
496:Transformer
389:. Splicing
330:spliceosome
319:pyrimidines
4859:Categories
4303:: e82556.
3360:(1): 672.
3311:: 103–12.
2835:(1): 188.
1224:(1): 1–8.
1214:Roberts RJ
1116:(1): 637.
979:References
886:Affymetrix
690:C. elegans
623:retrovirus
537:Sex lethal
533:stop codon
509:Drosophila
491:Drosophila
438:Drosophila
413:SR protein
379:cis-acting
305:Drosophila
263:Mechanisms
179:transcript
107:calcitonin
103:calcitonin
95:endogenous
91:transcript
79:adenovirus
67:eukaryotes
4799:Cytosolic
3462:: 10–22.
3411:(5): 32.
1852:1365-313X
928:Databases
816:addiction
710:mutations
688:), worm (
571:apoptosis
404:enhancers
391:silencers
300:nematodes
227:promoters
183:pre-mRNAs
73:Discovery
4506:Archived
4478:23161672
4429:32976589
4378:28855263
4329:36519529
4278:20015017
4186:19956082
4137:18245441
4085:15610736
4036:12114529
3987:22705790
3938:36821799
3887:21215366
3838:21459101
3785:23020045
3751:19157711
3743:25083822
3704:24459410
3655:17128174
3647:11861299
3606:15048092
3584:Oncogene
3565:16364913
3521:18054115
3486:29324392
3437:21619627
3386:25109687
3335:24802673
3286:22943729
3278:26300000
3256:Oncogene
3234:19266097
3194:PLOS ONE
3175:17416541
3140:19918805
3091:30174458
3083:15792793
3048:17158149
2991:11743582
2956:44052050
2948:10835645
2912:19165121
2904:10835644
2869:17916237
2810:24244129
2748:24951248
2699:16717195
2640:19048051
2583:11526107
2542:14703516
2498:16109372
2454:23772719
2446:11857317
2405:11421359
2312:15340491
2261:18204002
2204:20445623
2158:18369186
2099:19861426
2047:19959365
1998:21685335
1909:18285363
1860:15341630
1811:14883495
1803:15956978
1753:18688268
1685:33239457
1633:13829976
1625:10892653
1584:13208589
1431:39704416
1388:44642349
1199:27712956
1150:52113302
1142:30153812
1083:18978789
1043:23576288
1035:12626338
957:See also
952:database
946:database
940:database
582:figure).
548:above).
454:pre-mRNA
431:Examples
199:upstream
101:hormone
59:splicing
36:isoforms
4779:PRPF40B
4774:PRPF40A
4764:PRPF38B
4759:PRPF38A
4574:Nuclear
4469:3531113
4420:7778957
4369:5630039
4320:9812405
4269:3427726
4229:2200121
4209:Bibcode
4201:Science
4177:3152247
4128:2731647
4044:8689111
4014:Bibcode
4006:Science
3978:3465671
3878:3038581
3829:3139704
3695:3898681
3477:6728079
3428:4137096
3377:3219073
3326:4123867
3225:2648985
3202:Bibcode
3131:2855871
3039:1802581
2999:2724843
2860:2082043
2837:Bibcode
2801:3820534
2778:Bibcode
2739:4112133
2690:1482503
2667:Bibcode
2631:2561970
2396:1370132
2356:8769651
2252:2279241
2212:2398858
2184:Bibcode
2149:2327353
2090:2779669
2038:2834840
1989:3131313
1966:Bibcode
1900:2367711
1744:2467475
1721:Bibcode
1676:7851563
1576:6786756
1541:6952224
1509:Bibcode
1482:4318349
1474:7207587
1454:Bibcode
1345:3017190
1273:Bibcode
1246:2099968
1190:6526280
1133:6114036
1091:9228930
963:Exitron
938:AspicDB
894:ExonHit
869:in vivo
849:lariats
845:in vivo
806:gene –
756:SF2/ASF
718:RNA-Seq
703:Disease
292:introns
215:introns
99:thyroid
4769:PRPF39
4754:PRPF31
4749:PRPF19
4744:PRPF18
4729:PRPF4B
4665:Intron
4498:SciVee
4476:
4466:
4427:
4417:
4376:
4366:
4327:
4317:
4276:
4266:
4227:
4184:
4174:
4135:
4125:
4083:
4042:
4034:
3985:
3975:
3936:
3885:
3875:
3836:
3826:
3783:
3749:
3741:
3702:
3692:
3653:
3645:
3604:
3563:
3519:
3484:
3474:
3435:
3425:
3384:
3374:
3333:
3323:
3284:
3276:
3232:
3222:
3173:
3138:
3128:
3089:
3081:
3046:
3036:
2997:
2989:
2954:
2946:
2910:
2902:
2867:
2857:
2808:
2798:
2746:
2736:
2697:
2687:
2638:
2628:
2581:
2540:
2496:
2452:
2444:
2424:Cancer
2403:
2393:
2354:
2310:
2303:514884
2300:
2259:
2249:
2210:
2202:
2176:Nature
2156:
2146:
2097:
2087:
2045:
2035:
1996:
1986:
1930:
1907:
1897:
1858:
1850:
1809:
1801:
1751:
1741:
1683:
1673:
1631:
1623:
1582:
1574:
1539:
1532:346051
1529:
1480:
1472:
1446:Nature
1429:
1423:729004
1421:
1386:
1380:719751
1378:
1343:
1305:269380
1303:
1296:431482
1293:
1244:
1238:902310
1236:
1197:
1187:
1148:
1140:
1130:
1089:
1081:
1041:
1033:
950:ProSAS
908:CLIP (
771:methyl
738:DNMT3A
728:DNMT3A
621:, the
606:exon 2
334:snRNPs
321:– the
298:. (In
4822:DCP1B
4817:DCP1A
4739:PRPF8
4734:PRPF6
4724:PRPF4
4719:PRPF3
4714:PLRG1
4684:minor
4674:snRNP
4297:eLife
4040:S2CID
3747:S2CID
3651:S2CID
3625:Blood
3282:S2CID
3087:S2CID
2995:S2CID
2952:S2CID
2908:S2CID
2450:S2CID
2208:S2CID
1807:S2CID
1661:(4).
1629:S2CID
1580:S2CID
1478:S2CID
1427:S2CID
1384:S2CID
1242:S2CID
1146:S2CID
1087:S2CID
1039:S2CID
1011:(PDF)
898:Jivan
879:exons
808:ΔFosB
784:MST1R
575:TIA-1
462:gene
296:exons
201:exon.
142:Modes
123:Dscam
53:, or
49:, or
4842:EDC4
4837:EDC3
4832:DCPS
4827:DCP2
4669:Exon
4626:CFII
4616:PAB2
4606:CstF
4601:CPSF
4474:PMID
4425:PMID
4374:PMID
4325:PMID
4274:PMID
4225:PMID
4182:PMID
4133:PMID
4081:PMID
4032:PMID
3983:PMID
3934:PMID
3883:PMID
3857:Cell
3834:PMID
3781:PMID
3739:PMID
3700:PMID
3643:PMID
3602:PMID
3561:PMID
3517:PMID
3482:PMID
3433:PMID
3382:PMID
3331:PMID
3274:PMID
3230:PMID
3171:PMID
3136:PMID
3079:PMID
3044:PMID
2987:PMID
2944:PMID
2900:PMID
2865:PMID
2806:PMID
2744:PMID
2695:PMID
2636:PMID
2579:PMID
2538:PMID
2494:PMID
2442:PMID
2401:PMID
2352:PMID
2308:PMID
2257:PMID
2200:PMID
2154:PMID
2095:PMID
2043:PMID
1994:PMID
1928:ISBN
1905:PMID
1856:PMID
1848:ISSN
1799:PMID
1749:PMID
1681:PMID
1621:PMID
1603:Cell
1572:PMID
1556:Cell
1537:PMID
1470:PMID
1419:PMID
1403:Cell
1376:PMID
1360:Cell
1341:PMID
1301:PMID
1234:PMID
1218:Cell
1195:PMID
1138:PMID
1079:PMID
1031:PMID
912:and
902:cDNA
890:e.g.
883:e.g.
804:FOSB
767:DNMT
627:AIDS
591:ure6
541:U2AF
342:U2AF
310:mRNA
294:and
175:exon
4621:CFI
4611:PAP
4496:at
4464:PMC
4456:doi
4415:PMC
4405:doi
4364:PMC
4356:doi
4315:PMC
4305:doi
4264:PMC
4256:doi
4217:doi
4205:249
4172:PMC
4164:doi
4123:PMC
4115:doi
4071:doi
4022:doi
4010:297
3973:PMC
3965:doi
3924:hdl
3914:doi
3873:PMC
3865:doi
3861:144
3824:PMC
3816:doi
3731:doi
3690:PMC
3682:doi
3633:doi
3592:doi
3551:doi
3509:doi
3472:PMC
3464:doi
3423:PMC
3413:doi
3372:PMC
3362:doi
3321:PMC
3313:doi
3309:107
3264:doi
3220:PMC
3210:doi
3163:doi
3126:PMC
3118:doi
3114:220
3071:doi
3067:579
3034:PMC
3026:doi
2979:doi
2936:doi
2892:doi
2855:PMC
2845:doi
2796:PMC
2786:doi
2734:PMC
2726:doi
2685:PMC
2675:doi
2663:103
2626:PMC
2618:doi
2569:doi
2565:276
2528:doi
2524:279
2484:doi
2432:doi
2391:PMC
2383:doi
2375:RNA
2342:doi
2298:PMC
2288:doi
2247:PMC
2239:doi
2192:doi
2180:465
2144:PMC
2136:doi
2128:RNA
2085:PMC
2077:doi
2069:RNA
2033:PMC
2025:doi
1984:PMC
1974:doi
1962:108
1895:PMC
1887:doi
1838:doi
1791:doi
1739:PMC
1729:doi
1671:PMC
1663:doi
1611:doi
1607:101
1564:doi
1527:PMC
1517:doi
1462:doi
1450:290
1411:doi
1368:doi
1333:doi
1291:PMC
1281:doi
1226:doi
1185:PMC
1177:doi
1128:PMC
1118:doi
1071:doi
1023:doi
896:or
856:EST
793:Ron
779:Ron
632:tat
619:HIV
604:tat
586:b).
546:dsx
465:dsx
452:dsx
441:dsx
288:DNA
169:or
4861::
4689:U1
4472:.
4462:.
4452:41
4450:.
4446:.
4423:.
4413:.
4401:49
4399:.
4395:.
4372:.
4362:.
4352:27
4350:.
4346:.
4323:.
4313:.
4301:11
4299:.
4295:.
4272:.
4262:.
4252:13
4250:.
4246:.
4223:.
4215:.
4203:.
4180:.
4170:.
4160:34
4158:.
4154:.
4131:.
4121:.
4111:22
4109:.
4105:.
4093:^
4079:.
4067:16
4065:.
4061:.
4038:.
4030:.
4020:.
4008:.
4004:.
3981:.
3971:.
3961:19
3959:.
3955:.
3932:.
3922:.
3908:.
3904:.
3881:.
3871:.
3859:.
3855:.
3832:.
3822:.
3812:61
3810:.
3806:.
3787:.
3777:19
3775:.
3753:.
3745:.
3737:.
3727:40
3725:.
3706:.
3698:.
3688:.
3678:15
3676:.
3672:.
3649:.
3641:.
3629:99
3627:.
3623:.
3600:.
3588:23
3586:.
3582:.
3559:.
3547:20
3545:.
3541:.
3529:^
3515:.
3505:24
3503:.
3480:.
3470:.
3460:69
3458:.
3454:.
3431:.
3421:.
3407:.
3403:.
3380:.
3370:.
3358:15
3356:.
3352:.
3329:.
3319:.
3307:.
3303:.
3280:.
3272:.
3260:35
3258:.
3254:.
3242:^
3228:.
3218:.
3208:.
3196:.
3192:.
3169:.
3159:39
3157:.
3134:.
3124:.
3112:.
3108:.
3085:.
3077:.
3065:.
3042:.
3032:.
3022:35
3020:.
3016:.
2993:.
2985:.
2975:30
2973:.
2950:.
2942:.
2932:25
2930:.
2906:.
2898:.
2888:25
2886:.
2863:.
2853:.
2843:.
2831:.
2827:.
2804:.
2794:.
2784:.
2772:.
2768:.
2756:^
2742:.
2732:.
2722:30
2720:.
2716:.
2693:.
2683:.
2673:.
2661:.
2657:.
2634:.
2624:.
2612:.
2608:.
2591:^
2577:.
2563:.
2559:.
2536:.
2522:.
2518:.
2506:^
2492:.
2480:19
2478:.
2474:.
2462:^
2448:.
2440:.
2428:94
2426:.
2422:.
2399:.
2389:.
2377:.
2373:.
2350:.
2338:10
2336:.
2332:.
2320:^
2306:.
2296:.
2282:.
2278:.
2255:.
2245:.
2235:18
2233:.
2229:.
2206:.
2198:.
2190:.
2178:.
2166:^
2152:.
2142:.
2132:14
2130:.
2126:.
2107:^
2093:.
2083:.
2073:15
2071:.
2067:.
2055:^
2041:.
2031:.
2021:35
2019:.
2015:.
1992:.
1982:.
1972:.
1960:.
1956:.
1942:^
1903:.
1893:.
1883:36
1881:.
1877:.
1854:.
1846:.
1834:39
1832:.
1828:.
1805:.
1797:.
1785:.
1761:^
1747:.
1737:.
1727:.
1715:.
1711:.
1693:^
1679:.
1669:.
1659:95
1657:.
1653:.
1641:^
1627:.
1619:.
1605:.
1601:.
1578:.
1570:.
1560:24
1558:.
1535:.
1525:.
1515:.
1505:79
1503:.
1499:.
1476:.
1468:.
1460:.
1448:.
1425:.
1417:.
1407:15
1405:.
1382:.
1374:.
1364:15
1362:.
1339:.
1329:55
1327:.
1313:^
1299:.
1289:.
1279:.
1269:74
1267:.
1263:.
1240:.
1232:.
1222:12
1220:.
1193:.
1183:.
1173:42
1171:.
1167:.
1144:.
1136:.
1126:.
1114:19
1112:.
1108:.
1085:.
1077:.
1067:40
1065:.
1051:^
1037:.
1029:.
1019:72
1017:.
1013:.
986:^
822:.
799:.
787:)
762:.
338:U2
4558:e
4551:t
4544:v
4480:.
4458::
4431:.
4407::
4380:.
4358::
4331:.
4307::
4280:.
4258::
4231:.
4219::
4211::
4188:.
4166::
4139:.
4117::
4087:.
4073::
4046:.
4024::
4016::
3989:.
3967::
3940:.
3926::
3916::
3910:7
3889:.
3867::
3840:.
3818::
3733::
3684::
3657:.
3635::
3608:.
3594::
3567:.
3553::
3523:.
3511::
3488:.
3466::
3439:.
3415::
3409:3
3388:.
3364::
3337:.
3315::
3288:.
3266::
3236:.
3212::
3204::
3198:4
3177:.
3165::
3142:.
3120::
3093:.
3073::
3050:.
3028::
3001:.
2981::
2958:.
2938::
2914:.
2894::
2871:.
2847::
2839::
2833:7
2812:.
2788::
2780::
2774:9
2750:.
2728::
2701:.
2677::
2669::
2642:.
2620::
2614:1
2585:.
2571::
2544:.
2530::
2500:.
2486::
2456:.
2434::
2407:.
2385::
2379:7
2358:.
2344::
2314:.
2290::
2284:2
2263:.
2241::
2214:.
2194::
2186::
2160:.
2138::
2101:.
2079::
2049:.
2027::
2000:.
1976::
1968::
1936:.
1911:.
1889::
1862:.
1840::
1813:.
1793::
1787:6
1755:.
1731::
1723::
1717:4
1687:.
1665::
1635:.
1613::
1586:.
1566::
1543:.
1519::
1511::
1484:.
1464::
1456::
1433:.
1413::
1390:.
1370::
1347:.
1335::
1307:.
1283::
1275::
1248:.
1228::
1201:.
1179::
1152:.
1120::
1093:.
1073::
1045:.
1025::
881:(
781:(
407:(
394:(
185:.
110:(
42:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.