344:
321:
2064:
243:
218:
350:
249:
4426:
1582:
1756:
loads it onto the RISC complex where one strand serves as a template for the production of RNAi products and the other is degraded. Insects with mutations leading to non-functional components of their RNAi pathway show increased viral loads for viruses they carry or increased susceptibility to viruses for which they are hosts. Similarly to humans, insect viruses have evolved mechanisms to avoid the RNAi pathway. As an example,
1487:
1425:
31:
1648:. DNA damage increases in mammalian cells with decreased Dicer expression as a result of decreased efficiency of DNA damage repair and other mechanisms. For example, siRNA from double strand breaks (produced by Dicer) may act as guides for protein complexes involved in the double strand break repair mechanisms and can also direct
1619:)) was found to be elevated in patients with insufficient Dicer levels. These non coding strands of RNA can loop forming dsRNA structures that would be degraded by Dicer in a healthy retina. However, with insufficient Dicer levels, the accumulation of alu RNA leads to the degeneration of RPE as a result of inflammation.
1469:
Current research suggests the PAZ domain is capable of binding the 2 nucleotide 3' overhang of dsRNA while the RNaseIII catalytic domains form a pseudo-dimer around the dsRNA to initiate cleavage of the strands. This results in a functional shortening of the dsRNA strand. The distance between the PAZ
1712:
encode such RNAi suppressing proteins. Inhibition of dicer is beneficial to the virus as dicer is able to cleave viral dsRNA and load the product onto RISC resulting in targeted degradation of viral mRNA; thus fighting the infection. Another potential mechanism for viral pathogenesis is the blockade
1577:
differ in the fact that siRNAs are typically specific to the mRNA sequence while miRNAs aren't completely complementary to the mRNA sequence. miRNAs can interact with targets that have similar sequences, which inhibits translation of different genes. In general, RNA interference is an essential part
1799:
The siRNA was shown to be delivered in two ways in mammalian species such as mice. One way would be to directly inject into the system, which would not require Dicer function. Another way would be to introduce it by plasmids that encode for short hairpin RNA, which are cleaved by Dicer into siRNA.
1755:
species, serve as the vectors for these viruses, they are not the intended host of the virus. Transmission occurs as a result of the female mosquito's need for vertebrate blood to develop her eggs. The RNAi pathway in insects is very similar to that of other animals; Dicer-2 cleaves viral RNA and
1470:
and RNaseIII domains is determined by the angle of the connector helix and influences the length of the micro RNA product. The dsRBD domain binds the dsRNA, although the specific binding site of the domain has not been defined. It is possible that this domain works as part of a complex with other
2856:
Rivera B, Nadaf J, Fahiminiya S, Apellaniz-Ruiz M, Saskin A, Chong AS, Sharma S, Wagener R, Revil T, Condello V, Harra Z, Hamel N, Sabbaghian N, Muchantef K, Thomas C, de Kock L, Hébert-Blouin MN, Bassenden AV, Rabenstein H, Mete O, Paschke R, Pusztaszeri MP, Paulus W, Berghuis A, Ragoussis J,
1640:
expression. Decreased dicer mRNA levels correlate with advanced tumor stage. However, high dicer expression in other cancers, like prostate and esophageal, has been shown to correlate with poor patient prognosis. This discrepancy between cancer types suggests unique RNAi regulatory processes
1352:. Bernstein sought to discover the enzyme responsible for generating small RNA fragments from double-stranded RNA. Dicer's ability to generate around 22 nucleotide RNA fragments was discovered by separating it from the RISC enzyme complex after initiating the RNAi pathway with dsRNA
1615:, did not have symptoms of macular degeneration as Dicer-knockout mice. This observation suggested a Dicer specific role in retinal health that was independent of the RNAi pathway and thus not a function of si/miRNA generation. A form of RNA called Alu RNA (the RNA transcripts of
57:
1861:
Rice and grapes also produce DCLs as the dicer mechanism is a common defense strategy of many organisms. Rice has evolved other functions for the five DCLs it produces and they play a more important role in function and development than in
1356:. This experiment showed that RISC was not responsible for generating the observable small nucleotide fragments. Subsequent experiments testing RNase III family enzymes abilities to create RNA fragments narrowed the search to
1811:
inhibitors. In general, small molecular inhibitors are difficult in terms of specificity along with unendurable side effects. Antibodies are as specific as siRNA, but it is limited by only being able to be used against
1631:
expression profiles in malignant cancers suggest a pivotal role of miRNA and thus dicer in cancer development and prognosis. miRNAs can function as tumor suppressors and therefore their altered expression may result in
357:
256:
1725:, Dicer-1 generates microRNAs (miRNAs) by processing pre-miRNA, Dicer-2 is responsible for producing small interfering RNAs (siRNAs) from long double-stranded RNA (dsRNA). Insects can use Dicer as a potent
1415:
Dicer is due to at least five different domains being present within human Dicer. These domains are important in Dicer activity regulation, dsRNA processing, and RNA interference protein factor functioning.
1411:, representing the conserved functional core that has subsequently been found in larger Dicer proteins in other organisms; for example, it is 219 kDa in humans. The difference in size from humans to
2857:
Nikiforov YE, Siebert R, Albrecht S, Turcotte R, Hasselblatt M, Fabian MR, Foulkes WD (2019) DGCR8 microprocessor defect characterizes familial multinodular goiter with schwannomatosis. J Clin Invest
1565:(siRNA) are produced and function in a similar manner to miRNA by cleaving double-stranded RNA with Dicer into smaller fragments, 21 to 23 nucleotides in length. Both miRNAs and siRNAs activate the
2283:
Matsuda S, Ichigotani Y, Okuda T, Irimura T, Nakatsugawa S, Hamaguchi M (Jan 2000). "Molecular cloning and characterization of a novel human gene (HERNA) which encodes a putative RNA-helicase".
1603:
is a prominent cause of blindness in developed countries. Dicer's role in this disease became apparent after it was discovered that affected patients showed decreased levels of Dicer in their
1803:
One of the advantages of using Dicer to produce siRNA therapeutically would be the specificity and diversity of targets it can affect compared to what is currently being used such as
1478:) in order to effectively position the RNaseIII domains and thus control the specificity of the sRNA products. The helicase domain has been implicated in processing long substrates.
1874:. Rice DCL expression can be affected by biological stress conditions, including drought, salinity and cold. Thus these stressors may decrease a plant's viral resistance. Unlike
1660:
silencing and in their absence, as when Dicer is knocked out/down, can lead to activated transposons that cause DNA damage. Accumulation of DNA damage may result in cells with
1842:, four dicer like proteins are made and are designated DCL1 to DCL4. DCL1 is involved with miRNA generation and sRNA production from inverted repeats. DCL2 creates siRNA from
3067:
1772:
3a encodes an RNase III enzyme similar to the RNase III domains of dicer which may compete for dsRNA substrate as well as degrade siRNA duplexes to prevent RISC loading.
3907:
3853:
1554:
to form the pre-miRNA, a process that occurs in the nucleus. These pre-miRNA are then exported to the cytoplasm, where they are cleaved by Dicer to form mature miRNA.
1607:(RPE). Mice with Dicer knocked out, lacking Dicer only in their RPE, exhibited similar symptoms. However, other mice lacking important RNAi pathway proteins like
1788:
had decreased expression levels of Dicer. The same study showed that lower Dicer expression correlated with lower patient survival length. Along with being a
1569:(RISC), which finds the complementary target mRNA sequence and cleaves the RNA using RNase. This in turn silences the particular gene by RNA interference.
969:
950:
179:
1836:
Plant genomes encode for dicer-like proteins with similar functions and protein domains as animal and insect dicer. For example, in the model organism
2415:
Hammond SM, Bernstein E, Beach D, Hannon GJ (Mar 2000). "An RNA-directed nuclease mediates post-transcriptional gene silencing in
Drosophila cells".
1846:
antisense transcripts which aid in viral immunity and defense. DCL3 generates siRNA which aids in chromatin modification and DCL4 is involved in
1578:
of normal processes within organisms such as humans, and it is an area being researched as a diagnostic and therapeutic tool for cancer targets.
3515:
2218:
Macrae IJ, Zhou K, Li F, Repic A, Brooks AN, Cande WZ, et al. (Jan 2006). "Structural basis for double-stranded RNA processing by Dicer".
1828:. Also, producing miRNA therapeutically lacks in specificity because only 6-8 nucleotide base pairing is required for miRNA to attach to mRNA.
2001:
3676:
1983:
3333:
1375:
DCL1b, one of four DICER proteins, is not involved in miRNA biogenesis but in dicing miRNA target transcripts. Thus, a novel mechanism for
3772:
54:
2965:
1208:
1201:
2019:
Bernstein E, Caudy AA, Hammond SM, Hannon GJ (2001). "Role for a bidentate ribonuclease in the initiation step of RNA interference".
3483:
3028:"Intravenous injection of siRNA directed against hypoxia-inducible factors prolongs survival in a Lewis lung carcinoma cancer model"
2269:
1700:
as viruses that infect both plant and animal cells contain proteins designed to inhibit the RNAi response. In humans, the viruses
3500:
1636:. In analysis of lung and ovarian cancer, poor prognosis and decreased patient survival times correlate with decreased dicer and
3747:
1305:
4446:
2584:
2094:
1966:
3661:
3762:
3756:
3557:
3307:
1945:
1824:
uptake is the main obstacle of injection of siRNA. Injected SiRNA has poor stability in blood and causes stimulations of
3605:
1400:
2991:
Bronkhorst AW, van Rij RP (Aug 2014). "The long and short of antiviral defense: small RNA-based immunity in insects".
343:
4145:
3716:
3505:
1542:. These long sequences are cleaved into smaller precursor miRNA (pre-miRNA), which are usually 70 nucleotides with a
1784:
are present within the body based on the expression level of the enzyme. A study showed that many patients that had
1850:
metabolism and transcript silencing at the post-transcriptional level. Additionally, DCL 1 and 3 are important for
320:
3823:
3711:
1970:
1949:
3632:
3542:
3495:
3068:"Gene silencing by RNA interference is being used routinely to study gene function in cultured mammalian cells"
1652:
modifications. Additionally, miRNAs expression patterns change as a result of DNA damage caused by ionizing or
1014:
4471:
4301:
4054:
3995:
1892:
1566:
1499:
1376:
1349:
1309:
242:
217:
1866:. Additionally, expression patterns differ among the different plant cell types of rice while expression in
1763:
encodes for protein 1A which binds to dsRNA thus protecting it from dicer cleavage as well as RISC loading.
1438:
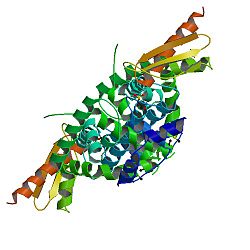
III domains are colored green, the PAZ domain yellow, the platform domain red, and the connector helix blue.
995:
4099:
3666:
3656:
159:
3094:"Cancer siRNA therapy by tumor selective delivery with ligand-targeted sterically stabilized nanoparticle"
4059:
3951:
3576:
356:
255:
4416:
3645:
3641:
3637:
3553:
3386:
1604:
349:
248:
2710:"DICER1 loss and Alu RNA induce age-related macular degeneration via the NLRP3 inflammasome and MyD88"
4456:
4402:
4389:
4376:
4363:
4350:
4337:
4324:
4286:
3813:
3369:
146:
2063:
4296:
4250:
4193:
3561:
3410:
3324:
2968:. National Center for Infections Disease, Center for Disease Control and Prevention. Archived from
1155:
1151:
1147:
1092:
1088:
1084:
167:
4198:
3430:
3300:
2318:
Hammond SM (Oct 2005). "Dicing and slicing: the core machinery of the RNA interference pathway".
2808:
Chiosea S, Jelezcova E, Chandran U, Acquafondata M, McHale T, Sobol RW, et al. (Nov 2006).
1184:
1163:
1159:
1121:
1100:
1096:
4451:
3986:
3625:
2526:
Vermeulen A, Behlen L, Reynolds A, Wolfson A, Marshall WS, Karpilow J, et al. (May 2005).
1817:
1507:
1337:
1277:
2918:"Phosphate and R2D2 restrict the substrate specificity of Dicer-2, an ATP-driven ribonuclease"
2368:"Phosphate and R2D2 restrict the substrate specificity of Dicer-2, an ATP-driven ribonuclease"
4219:
4138:
3921:
3818:
3610:
3571:
3435:
3357:
2969:
1907:
1825:
1733:
are responsible for the transmission of many viral diseases including the potentially deadly
1653:
1570:
1562:
1491:
1404:
1371:
1293:
231:
187:
4291:
4104:
3939:
3934:
3867:
3527:
3352:
3154:
2810:"Up-regulation of dicer, a component of the MicroRNA machinery, in prostate adenocarcinoma"
2672:
2613:
2424:
2227:
2028:
1838:
1600:
1450:
because it cleaves double-stranded RNA. In addition to two RNaseIII domains, it contains a
1430:
1391:
8:
4255:
3959:
3929:
3733:
3728:
3652:
3588:
3374:
2916:
Cenik ES, Fukunaga R, Lu G, Dutcher R, Wang Y, Tanaka Hall TM, et al. (April 2011).
2077:
1765:
1289:
150:, DCR1, Dicer, Dicer1e, HERNA, MNG1, RMSE2, K12H4.8-LIKE, dicer 1, ribonuclease III, GLOW
3158:
2676:
2617:
2602:"MicroRNAs and small interfering RNAs can inhibit mRNA expression by similar mechanisms"
2428:
2231:
2032:
1395:. The work was done by Ian MacRae while conducting research as a postdoctoral fellow in
4466:
4188:
4092:
3944:
3425:
3415:
3293:
3223:
3175:
3142:
2942:
2917:
2893:
2868:
2834:
2809:
2785:
2758:
2734:
2709:
2552:
2527:
2503:
2478:
2448:
2392:
2367:
2366:
Cenik ES, Fukunaga R, Lu G, Dutcher R, Wang Y, Tanaka Hall TM, et al. (Apr 2011).
2343:
2251:
2192:
2167:
2052:
1847:
1757:
1697:
191:
3118:
3093:
2636:
2601:
2296:
884:
879:
874:
869:
864:
859:
854:
849:
844:
839:
834:
829:
824:
819:
814:
809:
804:
799:
794:
778:
773:
768:
763:
758:
753:
748:
743:
738:
733:
728:
712:
707:
702:
697:
692:
687:
682:
677:
672:
667:
662:
657:
652:
647:
642:
637:
632:
627:
4461:
3893:
3838:
3806:
3681:
3400:
3274:
3265:
3253:
3244:
3215:
3180:
3123:
3049:
3008:
2947:
2898:
2839:
2790:
2739:
2690:
2641:
2580:
2557:
2508:
2440:
2397:
2335:
2300:
2243:
2197:
2145:
2100:
2090:
2044:
1726:
1543:
1471:
1292:(dsRNA) and pre-microRNA (pre-miRNA) into short double-stranded RNA fragments called
139:
47:
3227:
2347:
2255:
171:
4234:
4229:
4203:
4131:
3969:
3549:
3454:
3449:
3405:
3207:
3170:
3162:
3113:
3105:
3039:
3000:
2937:
2929:
2888:
2880:
2869:"The interplay between virus infection and the cellular RNA interference machinery"
2829:
2821:
2780:
2770:
2729:
2721:
2680:
2631:
2621:
2547:
2539:
2498:
2490:
2452:
2432:
2387:
2379:
2327:
2292:
2235:
2187:
2179:
2135:
2082:
2056:
2036:
1917:
1897:
1519:
1447:
1313:
436:
367:
311:
266:
2884:
2708:
Tarallo V, Hirano Y, Gelfand BD, Dridi S, Kerur N, Kim Y, et al. (May 2012).
2494:
2331:
4281:
4265:
4178:
4071:
3885:
3801:
3796:
3791:
3704:
3699:
3459:
3337:
3004:
2933:
2383:
1887:
1789:
1738:
1677:
1612:
1531:
1396:
1341:
411:
195:
3285:
2086:
2081:. Current Topics in Microbiology and Immunology. Vol. 320. pp. 77–97.
2006:
National Center for
Biotechnology Information, U.S. National Library of Medicine
1988:
National Center for
Biotechnology Information, U.S. National Library of Medicine
1386:
In terms of crystal structure, the first Dicer to be explored was that from the
4430:
4319:
4260:
4044:
4039:
4034:
3420:
3395:
3391:
3364:
3347:
2825:
2725:
2606:
Proceedings of the
National Academy of Sciences of the United States of America
2140:
2123:
1808:
1535:
1463:
1345:
512:
3211:
2183:
4440:
4224:
4183:
3785:
3694:
3444:
3092:
Schiffelers RM, Ansari A, Xu J, Zhou Q, Tang Q, Storm G, et al. (2004).
1821:
1751:
1633:
1503:
1408:
1366:
1325:
614:
3026:
Kamlah F, Eul BG, Li S, Lang N, Marsh LM, Seeger W, et al. (Mar 2009).
2626:
2579:. San Francisco, CA: Cold Spring Harbor Laboratory Press. pp. 641–648.
2239:
572:
450:
4173:
3912:
3843:
3566:
3487:
3219:
3184:
3127:
3053:
3012:
2951:
2902:
2843:
2794:
2743:
2694:
2645:
2561:
2512:
2444:
2401:
2339:
2304:
2247:
2201:
2149:
2104:
2048:
1746:
1742:
1616:
1539:
1353:
1321:
429:
208:
2775:
2528:"The contributions of dsRNA structure to Dicer specificity and efficiency"
1248:
1243:
18:
Enzyme that cleaves double-stranded RNA (dsRNA) into short dsRNA fragments
4397:
4332:
4168:
4087:
3858:
3752:
3615:
3581:
3519:
3109:
1878:, loss of function of DCL proteins causes developmental defects in rice.
1871:
1793:
1734:
1693:
1380:
1059:
1040:
3198:
Liu Q, Feng Y, Zhu Z (Aug 2009). "Dicer-like (DCL) proteins in plants".
3166:
3044:
3027:
2543:
4064:
2119:
1843:
1804:
1657:
1645:
1358:
328:
225:
175:
1696:
can trigger the RNAi cascade. It is likely dicer is involved in viral
116:
112:
108:
104:
100:
96:
92:
88:
84:
80:
76:
4371:
4345:
3977:
3776:
3689:
3316:
2436:
2040:
1792:, Dicer can be used for treating patients by injecting foreign siRNA
1769:
1705:
1680:
has been shown to be an autosomal dominant condition associated with
1661:
1649:
1459:
1387:
1317:
1301:
1285:
914:
395:
382:
294:
281:
183:
2807:
2685:
2660:
4425:
4022:
4017:
4012:
3833:
3468:
3320:
2285:
Biochimica et
Biophysica Acta (BBA) - Gene Structure and Expression
1902:
1730:
1709:
1681:
1581:
1495:
1451:
1297:
1232:
2168:"Structure of the human Dicer-TRBP complex by electron microscopy"
1486:
1466:, and two double stranded RNA binding domains (DUF283 and dsRBD).
4029:
4007:
4002:
3620:
3279:
3269:
3258:
3248:
1424:
1026:
981:
899:
895:
1490:
The enzyme dicer trims double stranded RNA or pri-miRNA to form
163:
4384:
4154:
4049:
3982:
3875:
3473:
3381:
1912:
1813:
1785:
1673:
1637:
1608:
1574:
1551:
1498:, respectively. These processed RNAs are incorporated into the
1443:
1272:
1216:
936:
2525:
1534:
of specific host mRNA sequences. miRNA is produced within the
1300:, respectively. These fragments are approximately 20–25
30:
4358:
3742:
3738:
2282:
1858:, DCL knockout does not cause severe developmental problems.
1781:
1701:
1628:
1547:
1527:
1435:
3264:
Overview of all the structural information available in the
3243:
Overview of all the structural information available in the
3143:"Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps"
1403:. A PAZ domain and two RNase III domains were discovered by
3723:
3600:
3593:
3537:
3532:
2414:
2117:
2018:
1922:
1455:
1281:
4123:
1775:
2707:
2124:"Transcriptional control of gene expression by microRNAs"
1523:
805:
negative regulation of transcription by RNA polymerase II
658:
endoribonuclease activity, producing 5'-phosphomonoesters
2479:"The dicey role of Dicer: implications for RNAi therapy"
2270:"Entrez Gene: DICER1 Dicer1, Dcr-1 homolog (Drosophila)"
2165:
845:
production of miRNAs involved in gene silencing by miRNA
419:
3091:
2118:
Khraiwesh B, Arif MA, Seumel GI, Ossowski S, Weigel D,
1434:, which catalyzes the cleavage of dsRNA to siRNAs. The
870:
negative regulation of tumor necrosis factor production
2915:
2365:
2166:
Lau PW, Potter CS, Carragher B, MacRae IJ (Oct 2009).
1713:
of dicer as a way to inhibit cellular miRNA pathways.
1641:
involving dicer differ amongst different tumor types.
4414:
584:
2076:
1729:. This finding is especially significant given that
774:
endoplasmic reticulum-Golgi intermediate compartment
835:
positive regulation of
Schwann cell differentiation
830:
RNA phosphodiester bond hydrolysis, endonucleolytic
3140:
2476:
1962:
1960:
1958:
1941:
1939:
1937:
3611:Fructose 6-P,2-kinase:fructose 2,6-bisphosphatase
3315:
2990:
2217:
1369:are present in many other organisms. In the moss
840:negative regulation of Schwann cell proliferation
366:
265:
4438:
3141:Chi SW, Zang JB, Mele A, Darnell RB (Jul 2009).
2866:
855:production of siRNA involved in RNA interference
3025:
1955:
1934:
1664:mutations and thus the development of a tumor.
1538:starting from primary miRNA (pri-miRNA) in the
2519:
1967:GRCm38: Ensembl release 89: ENSMUSG00000041415
1383:silencing of genes by miRNAs, was discovered.
4139:
3301:
2361:
2359:
2357:
2213:
2211:
2012:
3019:
2984:
2860:
2750:
2701:
2652:
2599:
2311:
2276:
3085:
2763:International Journal of Molecular Sciences
2477:Merritt WM, Bar-Eli M, Sood AK (Apr 2010).
2408:
2111:
2070:
1946:GRCh38: Ensembl release 89: ENSG00000100697
1749:. While mosquitoes, more specifically the
1585:Formation of miRNA used in RNA interference
1481:
4146:
4132:
3308:
3294:
3197:
2850:
2472:
2470:
2468:
2466:
2464:
2462:
2354:
2208:
1308:. Dicer facilitates the activation of the
810:peripheral nervous system myelin formation
3174:
3117:
3043:
2941:
2892:
2833:
2784:
2774:
2733:
2684:
2635:
2625:
2593:
2574:
2551:
2502:
2391:
2191:
2139:
3134:
3060:
2759:"The role of dicer in DNA damage repair"
1580:
1557:
1485:
1423:
3501:Ubiquitin carboxy-terminal hydrolase L1
2756:
2658:
2459:
2317:
1820:. On the other hand, low efficiency of
1776:Diagnostic and therapeutic applications
1594:
1428:One molecule of the Dicer protein from
4439:
3278:(Mouse Endoribonuclease Dicer) at the
3257:(Human Endoribonuclease Dicer) at the
2262:
1831:
1780:Dicer can be used to identify whether
1656:. RNAi mechanisms are responsible for
1442:Human dicer (also known as hsDicer or
880:negative regulation of gene expression
4127:
4081:either deoxy- or ribo-
3289:
3200:Functional & Integrative Genomics
2161:
2159:
1687:
1419:
1304:long with a two-base overhang on the
371:
332:
327:
270:
229:
224:
3662:Protein serine/threonine phosphatase
2600:Zeng Y, Yi R, Cullen BR (Aug 2003).
1522:is a process where the breakdown of
1336:Dicer was given its name in 2001 by
3763:Cyclic nucleotide phosphodiesterase
3757:Clostridium perfringens alpha toxin
3558:Tartrate-resistant acid phosphatase
2867:Berkhout B, Haasnoot J (May 2006).
1667:
13:
3606:Pyruvate dehydrogenase phosphatase
2156:
1401:University of California, Berkeley
820:positive regulation of myelination
14:
4483:
3506:4-hydroxybenzoyl-CoA thioesterase
3237:
2814:The American Journal of Pathology
1316:. RISC has a catalytic component
1275:that in humans is encoded by the
4424:
2062:
497:inferior ganglion of vagus nerve
355:
348:
342:
319:
254:
247:
241:
216:
29:
3824:N-acetylglucosamine-6-sulfatase
3712:Sphingomyelin phosphodiesterase
3191:
2958:
2909:
2801:
2661:"Vision: Dicer leaps into view"
2568:
1474:(TRBP in humans, R2D2, Loqs in
1312:(RISC), which is essential for
795:neuron projection morphogenesis
638:protein domain specific binding
3633:Inositol-phosphate phosphatase
3496:Palmitoyl protein thioesterase
1994:
1976:
1546:. Pri-miRNA are identified by
1407:. The protein size is 82
573:More reference expression data
1:
3996:RNA-induced silencing complex
2885:10.1016/j.febslet.2006.02.070
2577:Molecular Biology of the Gene
2495:10.1158/0008-5472.CAN-09-2536
2332:10.1016/j.febslet.2005.08.079
2297:10.1016/S0167-4781(99)00221-3
1928:
1716:
1567:RNA-induced silencing complex
1500:RNA-induced silencing complex
1377:regulation of gene expression
1350:Cold Spring Harbor Laboratory
1344:while conducting research in
1310:RNA-induced silencing complex
340:
239:
4447:Genes on human chromosome 14
4100:Serratia marcescens nuclease
3667:Dual-specificity phosphatase
3657:Protein tyrosine phosphatase
3005:10.1016/j.coviro.2014.03.010
2934:10.1016/j.molcel.2011.03.002
2384:10.1016/j.molcel.2011.03.002
1514:
1331:
713:deoxyribonuclease I activity
7:
4153:
3577:Fructose 1,6-bisphosphatase
2993:Current Opinion in Virology
2087:10.1007/978-3-540-75157-1_4
1881:
865:apoptotic DNA fragmentation
683:double-stranded RNA binding
10:
4488:
2826:10.2353/ajpath.2006.060480
2726:10.1016/j.cell.2012.03.036
2141:10.1016/j.cell.2009.12.023
2122:, et al. (Jan 2010).
1644:Dicer is also involved in
1605:retinal pigment epithelium
1589:
1209:Chr 12: 104.65 – 104.72 Mb
4310:
4302:Michaelis–Menten kinetics
4274:
4243:
4212:
4161:
4080:
3968:
3920:
3906:
3884:
3866:
3852:
3832:
3814:Galactosamine-6 sulfatase
3771:
3675:
3514:
3482:
3370:6-phosphogluconolactonase
3332:
3212:10.1007/s10142-009-0111-5
2966:"Mosquito-borne Diseases"
2184:10.1016/j.str.2009.08.013
2002:"Mouse PubMed Reference:"
1984:"Human PubMed Reference:"
1796:to cause gene silencing.
1622:
1362:CG4792, now named Dicer.
1269:helicase with RNase motif
1247:
1242:
1238:
1231:
1215:
1196:
1181:
1177:
1144:
1140:
1133:
1118:
1114:
1081:
1077:
1070:
1057:
1053:
1038:
1034:
1025:
1012:
1008:
993:
989:
980:
967:
963:
948:
944:
935:
920:
913:
909:
893:
703:ribonuclease III activity
653:endoribonuclease activity
613:
609:
597:
592:
583:
570:
547:medullary collecting duct
519:
510:
481:mucosa of paranasal sinus
457:
448:
418:
410:
406:
389:
376:
339:
318:
309:
305:
288:
275:
238:
215:
206:
202:
157:
154:
144:
137:
132:
73:
68:
51:
46:
41:
37:
28:
23:
4194:Diffusion-limited enzyme
3562:Purple acid phosphatases
1482:Role in RNA interference
1202:Chr 14: 95.09 – 95.16 Mb
2757:Tang KF, Ren H (2012).
2627:10.1073/pnas.1630797100
2240:10.1126/science.1121638
875:NIK/NF-kappaB signaling
850:miRNA metabolic process
3987:Microprocessor complex
3626:Beta-propeller phytase
3098:Nucleic Acids Research
2659:Meister G (Mar 2011).
1586:
1511:
1502:(RISC), which targets
1439:
1288:family, Dicer cleaves
1265:endoribonuclease Dicer
4287:Eadie–Hofstee diagram
4220:Allosteric regulation
3922:Endodeoxyribonuclease
3819:Iduronate-2-sulfatase
3572:Glucose 6-phosphatase
3358:Butyrylcholinesterase
2776:10.3390/ijms131216769
1908:Small interfering RNA
1826:non-specific immunity
1654:ultraviolet radiation
1584:
1563:Small interfering RNA
1558:Small Interfering RNA
1492:small interfering RNA
1489:
1427:
1405:X-ray crystallography
1372:Physcomitrella patens
1324:capable of degrading
1294:small interfering RNA
779:extracellular exosome
688:endonuclease activity
543:mandibular prominence
373:12 E|12 54.83 cM
334:Chromosome 12 (mouse)
232:Chromosome 14 (human)
4472:RNA-binding proteins
4297:Lineweaver–Burk plot
4105:Micrococcal nuclease
3940:Deoxyribonuclease IV
3935:Deoxyribonuclease II
3868:Exodeoxyribonuclease
3528:Alkaline phosphatase
3353:Acetylcholinesterase
1839:Arabidopsis thaliana
1601:macular degeneration
1595:Macular degeneration
1431:Giardia intestinalis
1392:Giardia intestinalis
1284:. Being part of the
815:pre-miRNA processing
754:RISC-loading complex
69:List of PDB id codes
42:Available structures
3960:UvrABC endonuclease
3930:Deoxyribonuclease I
3653:Protein phosphatase
3589:Protein phosphatase
3387:Bile salt-dependent
3375:PAF acetylhydrolase
3167:10.1038/nature08170
3159:2009Natur.460..479C
3045:10.1038/cgt.2008.71
3032:Cancer Gene Therapy
2677:2011Natur.471..308M
2618:2003PNAS..100.9779Z
2544:10.1261/rna.7272305
2429:2000Natur.404..293H
2232:2006Sci...311..195M
2033:2001Natur.409..363B
1832:Dicer-like proteins
1766:Heliothis virescens
1290:double-stranded RNA
4256:Enzyme superfamily
4189:Enzyme promiscuity
4093:Mung bean nuclease
3952:Restriction enzyme
3945:Restriction enzyme
3110:10.1093/nar/gnh140
2972:on 31 January 2014
2575:Watson JD (2008).
1848:trans-acting siRNA
1688:Viral pathogenesis
1587:
1512:
1472:regulator proteins
1446:) is classified a
1440:
1420:Functional domains
1015:ENSMUSG00000041415
788:Biological process
722:Cellular component
693:hydrolase activity
628:nucleotide binding
621:Molecular function
461:tail of epididymis
196:DICER1 - orthologs
4412:
4411:
4121:
4120:
4117:
4116:
4113:
4112:
3902:
3901:
3894:Oligonucleotidase
3839:deoxyribonuclease
3807:Steroid sulfatase
3682:Phosphodiesterase
3411:Hormone-sensitive
3072:Life Technologies
2586:978-0-8053-9592-1
2096:978-3-540-75156-4
1818:surface receptors
1544:hairpin structure
1258:
1257:
1254:
1253:
1227:
1226:
1192:
1191:
1171:
1170:
1129:
1128:
1108:
1107:
1066:
1065:
1047:
1046:
1021:
1020:
1002:
1001:
976:
975:
957:
956:
905:
904:
825:nerve development
678:nuclease activity
648:metal ion binding
643:pre-miRNA binding
633:helicase activity
605:
604:
601:
600:
579:
578:
566:
565:
504:
503:
477:corpus epididymis
402:
401:
301:
300:
128:
127:
124:
123:
52:Ortholog search:
4479:
4457:RNA interference
4429:
4428:
4420:
4292:Hanes–Woolf plot
4235:Enzyme activator
4230:Enzyme inhibitor
4204:Enzyme catalysis
4148:
4141:
4134:
4125:
4124:
3970:Endoribonuclease
3956:
3950:
3918:
3917:
3864:
3863:
3850:
3849:
3550:Acid phosphatase
3431:Monoacylglycerol
3341:ester hydrolases
3310:
3303:
3296:
3287:
3286:
3232:
3231:
3195:
3189:
3188:
3178:
3153:(7254): 479–86.
3138:
3132:
3131:
3121:
3089:
3083:
3082:
3080:
3078:
3064:
3058:
3057:
3047:
3023:
3017:
3016:
2988:
2982:
2981:
2979:
2977:
2962:
2956:
2955:
2945:
2913:
2907:
2906:
2896:
2879:(12): 2896–902.
2864:
2858:
2854:
2848:
2847:
2837:
2805:
2799:
2798:
2788:
2778:
2769:(12): 16769–78.
2754:
2748:
2747:
2737:
2705:
2699:
2698:
2688:
2656:
2650:
2649:
2639:
2629:
2597:
2591:
2590:
2572:
2566:
2565:
2555:
2523:
2517:
2516:
2506:
2474:
2457:
2456:
2437:10.1038/35005107
2412:
2406:
2405:
2395:
2363:
2352:
2351:
2315:
2309:
2308:
2280:
2274:
2273:
2266:
2260:
2259:
2215:
2206:
2205:
2195:
2163:
2154:
2153:
2143:
2115:
2109:
2108:
2079:RNA Interference
2074:
2068:
2067:
2066:
2060:
2041:10.1038/35053110
2016:
2010:
2009:
1998:
1992:
1991:
1980:
1974:
1964:
1953:
1943:
1918:Ribonuclease III
1898:RNA interference
1668:Other conditions
1520:RNA interference
1448:Ribonuclease III
1314:RNA interference
1263:, also known as
1240:
1239:
1211:
1204:
1187:
1175:
1174:
1166:
1138:
1137:
1134:RefSeq (protein)
1124:
1112:
1111:
1103:
1075:
1074:
1051:
1050:
1032:
1031:
1006:
1005:
987:
986:
961:
960:
942:
941:
911:
910:
611:
610:
590:
589:
575:
559:primitive streak
539:genital tubercle
527:secondary oocyte
515:
513:Top expressed in
508:
507:
465:caput epididymis
453:
451:Top expressed in
446:
445:
425:
424:
408:
407:
398:
385:
374:
359:
352:
346:
335:
323:
307:
306:
297:
284:
273:
258:
251:
245:
234:
220:
204:
203:
198:
149:
142:
119:
66:
65:
60:
39:
38:
33:
21:
20:
4487:
4486:
4482:
4481:
4480:
4478:
4477:
4476:
4437:
4436:
4435:
4423:
4415:
4413:
4408:
4320:Oxidoreductases
4306:
4282:Enzyme kinetics
4270:
4266:List of enzymes
4239:
4208:
4179:Catalytic triad
4157:
4152:
4122:
4109:
4076:
3964:
3954:
3948:
3911:
3898:
3886:Exoribonuclease
3880:
3857:
3841:
3837:
3828:
3802:Arylsulfatase L
3797:Arylsulfatase B
3792:Arylsulfatase A
3767:
3680:
3671:
3510:
3478:
3340:
3328:
3314:
3240:
3235:
3196:
3192:
3139:
3135:
3090:
3086:
3076:
3074:
3066:
3065:
3061:
3024:
3020:
2989:
2985:
2975:
2973:
2964:
2963:
2959:
2914:
2910:
2865:
2861:
2855:
2851:
2806:
2802:
2755:
2751:
2706:
2702:
2686:10.1038/471308a
2671:(7338): 308–9.
2657:
2653:
2612:(17): 9779–84.
2598:
2594:
2587:
2573:
2569:
2524:
2520:
2483:Cancer Research
2475:
2460:
2423:(6775): 293–6.
2413:
2409:
2364:
2355:
2316:
2312:
2281:
2277:
2268:
2267:
2263:
2226:(5758): 195–8.
2216:
2209:
2178:(10): 1326–32.
2164:
2157:
2116:
2112:
2097:
2075:
2071:
2061:
2027:(6818): 363–6.
2017:
2013:
2000:
1999:
1995:
1982:
1981:
1977:
1965:
1956:
1944:
1935:
1931:
1888:gene expression
1884:
1834:
1809:small molecular
1790:diagnostic tool
1778:
1739:West Nile virus
1719:
1690:
1678:schwannomatosis
1670:
1625:
1597:
1592:
1560:
1550:and cleaved by
1532:gene expression
1526:molecules into
1517:
1484:
1454:domain, a PAZ (
1422:
1413:G. intestinalis
1397:Jennifer Doudna
1342:Emily Bernstein
1334:
1249:View/Edit Mouse
1244:View/Edit Human
1207:
1200:
1197:Location (UCSC)
1183:
1162:
1158:
1154:
1150:
1146:
1120:
1099:
1095:
1091:
1087:
1083:
996:ENSG00000100697
889:
783:
717:
663:protein binding
571:
562:
557:
553:
551:renal corpuscle
549:
545:
541:
537:
533:
529:
525:
511:
500:
495:
493:parietal pleura
491:
487:
485:visceral pleura
483:
479:
475:
473:trabecular bone
471:
467:
463:
449:
393:
380:
372:
362:
361:
360:
353:
333:
310:Gene location (
292:
279:
271:
261:
260:
259:
252:
230:
207:Gene location (
158:
145:
138:
75:
53:
19:
12:
11:
5:
4485:
4475:
4474:
4469:
4464:
4459:
4454:
4449:
4434:
4433:
4410:
4409:
4407:
4406:
4393:
4380:
4367:
4354:
4341:
4328:
4314:
4312:
4308:
4307:
4305:
4304:
4299:
4294:
4289:
4284:
4278:
4276:
4272:
4271:
4269:
4268:
4263:
4258:
4253:
4247:
4245:
4244:Classification
4241:
4240:
4238:
4237:
4232:
4227:
4222:
4216:
4214:
4210:
4209:
4207:
4206:
4201:
4196:
4191:
4186:
4181:
4176:
4171:
4165:
4163:
4159:
4158:
4151:
4150:
4143:
4136:
4128:
4119:
4118:
4115:
4114:
4111:
4110:
4108:
4107:
4102:
4097:
4096:
4095:
4084:
4082:
4078:
4077:
4075:
4074:
4069:
4068:
4067:
4062:
4057:
4052:
4042:
4037:
4032:
4027:
4026:
4025:
4020:
4015:
4010:
4000:
3999:
3998:
3989:
3974:
3972:
3966:
3965:
3963:
3962:
3957:
3942:
3937:
3932:
3926:
3924:
3915:
3904:
3903:
3900:
3899:
3897:
3896:
3890:
3888:
3882:
3881:
3879:
3878:
3872:
3870:
3861:
3847:
3830:
3829:
3827:
3826:
3821:
3816:
3811:
3810:
3809:
3804:
3799:
3794:
3781:
3779:
3769:
3768:
3766:
3765:
3760:
3750:
3745:
3736:
3731:
3726:
3721:
3720:
3719:
3709:
3708:
3707:
3702:
3692:
3686:
3684:
3673:
3672:
3670:
3669:
3664:
3659:
3650:
3649:
3648:
3630:
3629:
3628:
3618:
3613:
3608:
3603:
3598:
3597:
3596:
3586:
3585:
3584:
3574:
3569:
3564:
3547:
3546:
3545:
3540:
3535:
3524:
3522:
3512:
3511:
3509:
3508:
3503:
3498:
3492:
3490:
3480:
3479:
3477:
3476:
3471:
3465:
3464:
3463:
3462:
3457:
3452:
3441:
3440:
3439:
3438:
3436:Diacylglycerol
3433:
3428:
3423:
3418:
3413:
3408:
3403:
3398:
3389:
3378:
3377:
3372:
3367:
3365:Pectinesterase
3362:
3361:
3360:
3355:
3348:Cholinesterase
3344:
3342:
3330:
3329:
3313:
3312:
3305:
3298:
3290:
3284:
3283:
3262:
3239:
3238:External links
3236:
3234:
3233:
3190:
3133:
3084:
3059:
3038:(3): 195–205.
3018:
2983:
2957:
2928:(2): 172–184.
2922:Molecular Cell
2908:
2859:
2849:
2820:(5): 1812–20.
2800:
2749:
2700:
2651:
2592:
2585:
2567:
2518:
2458:
2407:
2372:Molecular Cell
2353:
2326:(26): 5822–9.
2310:
2291:(1–2): 163–9.
2275:
2261:
2207:
2155:
2110:
2095:
2069:
2011:
1993:
1975:
1954:
1932:
1930:
1927:
1926:
1925:
1920:
1915:
1910:
1905:
1900:
1895:
1890:
1883:
1880:
1854:flowering. In
1833:
1830:
1777:
1774:
1718:
1715:
1689:
1686:
1684:in this gene.
1669:
1666:
1624:
1621:
1596:
1593:
1591:
1588:
1559:
1556:
1516:
1513:
1483:
1480:
1421:
1418:
1399:'s lab at the
1346:Gregory Hannon
1333:
1330:
1320:, which is an
1256:
1255:
1252:
1251:
1246:
1236:
1235:
1229:
1228:
1225:
1224:
1222:
1220:
1213:
1212:
1205:
1198:
1194:
1193:
1190:
1189:
1179:
1178:
1172:
1169:
1168:
1142:
1141:
1135:
1131:
1130:
1127:
1126:
1116:
1115:
1109:
1106:
1105:
1079:
1078:
1072:
1068:
1067:
1064:
1063:
1055:
1054:
1048:
1045:
1044:
1036:
1035:
1029:
1023:
1022:
1019:
1018:
1010:
1009:
1003:
1000:
999:
991:
990:
984:
978:
977:
974:
973:
965:
964:
958:
955:
954:
946:
945:
939:
933:
932:
927:
922:
918:
917:
907:
906:
903:
902:
891:
890:
888:
887:
885:tube formation
882:
877:
872:
867:
862:
860:gene silencing
857:
852:
847:
842:
837:
832:
827:
822:
817:
812:
807:
802:
800:RNA processing
797:
791:
789:
785:
784:
782:
781:
776:
771:
766:
761:
756:
751:
746:
741:
736:
731:
725:
723:
719:
718:
716:
715:
710:
705:
700:
695:
690:
685:
680:
675:
670:
665:
660:
655:
650:
645:
640:
635:
630:
624:
622:
618:
617:
607:
606:
603:
602:
599:
598:
595:
594:
587:
581:
580:
577:
576:
568:
567:
564:
563:
561:
560:
556:
555:abdominal wall
552:
548:
544:
540:
536:
535:tail of embryo
532:
531:primary oocyte
528:
524:
520:
517:
516:
505:
502:
501:
499:
498:
494:
490:
486:
482:
478:
474:
470:
466:
462:
458:
455:
454:
442:
441:
433:
422:
416:
415:
412:RNA expression
404:
403:
400:
399:
391:
387:
386:
378:
375:
370:
364:
363:
354:
347:
341:
337:
336:
331:
325:
324:
316:
315:
303:
302:
299:
298:
290:
286:
285:
277:
274:
269:
263:
262:
253:
246:
240:
236:
235:
228:
222:
221:
213:
212:
200:
199:
156:
152:
151:
143:
135:
134:
130:
129:
126:
125:
122:
121:
71:
70:
62:
61:
50:
44:
43:
35:
34:
26:
25:
17:
9:
6:
4:
3:
2:
4484:
4473:
4470:
4468:
4465:
4463:
4460:
4458:
4455:
4453:
4452:Ribonucleases
4450:
4448:
4445:
4444:
4442:
4432:
4427:
4422:
4421:
4418:
4404:
4400:
4399:
4394:
4391:
4387:
4386:
4381:
4378:
4374:
4373:
4368:
4365:
4361:
4360:
4355:
4352:
4348:
4347:
4342:
4339:
4335:
4334:
4329:
4326:
4322:
4321:
4316:
4315:
4313:
4309:
4303:
4300:
4298:
4295:
4293:
4290:
4288:
4285:
4283:
4280:
4279:
4277:
4273:
4267:
4264:
4262:
4261:Enzyme family
4259:
4257:
4254:
4252:
4249:
4248:
4246:
4242:
4236:
4233:
4231:
4228:
4226:
4225:Cooperativity
4223:
4221:
4218:
4217:
4215:
4211:
4205:
4202:
4200:
4197:
4195:
4192:
4190:
4187:
4185:
4184:Oxyanion hole
4182:
4180:
4177:
4175:
4172:
4170:
4167:
4166:
4164:
4160:
4156:
4149:
4144:
4142:
4137:
4135:
4130:
4129:
4126:
4106:
4103:
4101:
4098:
4094:
4091:
4090:
4089:
4086:
4085:
4083:
4079:
4073:
4070:
4066:
4063:
4061:
4058:
4056:
4053:
4051:
4048:
4047:
4046:
4043:
4041:
4038:
4036:
4033:
4031:
4028:
4024:
4021:
4019:
4016:
4014:
4011:
4009:
4006:
4005:
4004:
4001:
3997:
3993:
3990:
3988:
3984:
3981:
3980:
3979:
3976:
3975:
3973:
3971:
3967:
3961:
3958:
3953:
3946:
3943:
3941:
3938:
3936:
3933:
3931:
3928:
3927:
3925:
3923:
3919:
3916:
3914:
3909:
3905:
3895:
3892:
3891:
3889:
3887:
3883:
3877:
3874:
3873:
3871:
3869:
3865:
3862:
3860:
3855:
3851:
3848:
3845:
3840:
3835:
3831:
3825:
3822:
3820:
3817:
3815:
3812:
3808:
3805:
3803:
3800:
3798:
3795:
3793:
3790:
3789:
3788:
3787:
3786:arylsulfatase
3783:
3782:
3780:
3778:
3774:
3770:
3764:
3761:
3758:
3754:
3751:
3749:
3746:
3744:
3740:
3737:
3735:
3732:
3730:
3727:
3725:
3722:
3718:
3715:
3714:
3713:
3710:
3706:
3703:
3701:
3698:
3697:
3696:
3695:Phospholipase
3693:
3691:
3688:
3687:
3685:
3683:
3678:
3674:
3668:
3665:
3663:
3660:
3658:
3654:
3651:
3647:
3643:
3639:
3636:
3635:
3634:
3631:
3627:
3624:
3623:
3622:
3619:
3617:
3614:
3612:
3609:
3607:
3604:
3602:
3599:
3595:
3592:
3591:
3590:
3587:
3583:
3580:
3579:
3578:
3575:
3573:
3570:
3568:
3565:
3563:
3559:
3555:
3551:
3548:
3544:
3541:
3539:
3536:
3534:
3531:
3530:
3529:
3526:
3525:
3523:
3521:
3517:
3513:
3507:
3504:
3502:
3499:
3497:
3494:
3493:
3491:
3489:
3485:
3481:
3475:
3472:
3470:
3467:
3466:
3461:
3458:
3456:
3453:
3451:
3448:
3447:
3446:
3445:Phospholipase
3443:
3442:
3437:
3434:
3432:
3429:
3427:
3424:
3422:
3419:
3417:
3414:
3412:
3409:
3407:
3404:
3402:
3399:
3397:
3393:
3390:
3388:
3385:
3384:
3383:
3380:
3379:
3376:
3373:
3371:
3368:
3366:
3363:
3359:
3356:
3354:
3351:
3350:
3349:
3346:
3345:
3343:
3339:
3335:
3331:
3326:
3322:
3318:
3311:
3306:
3304:
3299:
3297:
3292:
3291:
3288:
3281:
3277:
3276:
3271:
3267:
3263:
3260:
3256:
3255:
3250:
3246:
3242:
3241:
3229:
3225:
3221:
3217:
3213:
3209:
3206:(3): 277–86.
3205:
3201:
3194:
3186:
3182:
3177:
3172:
3168:
3164:
3160:
3156:
3152:
3148:
3144:
3137:
3129:
3125:
3120:
3115:
3111:
3107:
3103:
3099:
3095:
3088:
3073:
3069:
3063:
3055:
3051:
3046:
3041:
3037:
3033:
3029:
3022:
3014:
3010:
3006:
3002:
2998:
2994:
2987:
2971:
2967:
2961:
2953:
2949:
2944:
2939:
2935:
2931:
2927:
2923:
2919:
2912:
2904:
2900:
2895:
2890:
2886:
2882:
2878:
2874:
2870:
2863:
2853:
2845:
2841:
2836:
2831:
2827:
2823:
2819:
2815:
2811:
2804:
2796:
2792:
2787:
2782:
2777:
2772:
2768:
2764:
2760:
2753:
2745:
2741:
2736:
2731:
2727:
2723:
2720:(4): 847–59.
2719:
2715:
2711:
2704:
2696:
2692:
2687:
2682:
2678:
2674:
2670:
2666:
2662:
2655:
2647:
2643:
2638:
2633:
2628:
2623:
2619:
2615:
2611:
2607:
2603:
2596:
2588:
2582:
2578:
2571:
2563:
2559:
2554:
2549:
2545:
2541:
2538:(5): 674–82.
2537:
2533:
2529:
2522:
2514:
2510:
2505:
2500:
2496:
2492:
2489:(7): 2571–4.
2488:
2484:
2480:
2473:
2471:
2469:
2467:
2465:
2463:
2454:
2450:
2446:
2442:
2438:
2434:
2430:
2426:
2422:
2418:
2411:
2403:
2399:
2394:
2389:
2385:
2381:
2378:(2): 172–84.
2377:
2373:
2369:
2362:
2360:
2358:
2349:
2345:
2341:
2337:
2333:
2329:
2325:
2321:
2314:
2306:
2302:
2298:
2294:
2290:
2286:
2279:
2271:
2265:
2257:
2253:
2249:
2245:
2241:
2237:
2233:
2229:
2225:
2221:
2214:
2212:
2203:
2199:
2194:
2189:
2185:
2181:
2177:
2173:
2169:
2162:
2160:
2151:
2147:
2142:
2137:
2134:(1): 111–22.
2133:
2129:
2125:
2121:
2114:
2106:
2102:
2098:
2092:
2088:
2084:
2080:
2073:
2065:
2058:
2054:
2050:
2046:
2042:
2038:
2034:
2030:
2026:
2022:
2015:
2007:
2003:
1997:
1989:
1985:
1979:
1972:
1968:
1963:
1961:
1959:
1951:
1947:
1942:
1940:
1938:
1933:
1924:
1921:
1919:
1916:
1914:
1911:
1909:
1906:
1904:
1901:
1899:
1896:
1894:
1891:
1889:
1886:
1885:
1879:
1877:
1873:
1869:
1865:
1859:
1857:
1853:
1849:
1845:
1841:
1840:
1829:
1827:
1823:
1822:intracellular
1819:
1815:
1810:
1806:
1801:
1797:
1795:
1794:intravenously
1791:
1787:
1783:
1773:
1771:
1768:
1767:
1762:
1760:
1754:
1753:
1752:Aedes aegypti
1748:
1744:
1740:
1736:
1732:
1728:
1724:
1714:
1711:
1707:
1703:
1699:
1695:
1692:Infection by
1685:
1683:
1679:
1675:
1672:Multinodular
1665:
1663:
1659:
1655:
1651:
1647:
1642:
1639:
1635:
1634:tumorigenesis
1630:
1620:
1618:
1614:
1610:
1606:
1602:
1583:
1579:
1576:
1572:
1568:
1564:
1555:
1553:
1549:
1545:
1541:
1537:
1533:
1529:
1525:
1521:
1509:
1505:
1504:messenger RNA
1501:
1497:
1493:
1488:
1479:
1477:
1473:
1467:
1465:
1461:
1457:
1453:
1449:
1445:
1437:
1433:
1432:
1426:
1417:
1414:
1410:
1406:
1402:
1398:
1394:
1393:
1389:
1384:
1382:
1378:
1374:
1373:
1368:
1363:
1361:
1360:
1355:
1351:
1347:
1343:
1339:
1329:
1327:
1326:messenger RNA
1323:
1319:
1315:
1311:
1307:
1303:
1299:
1295:
1291:
1287:
1283:
1280:
1279:
1274:
1270:
1266:
1262:
1250:
1245:
1241:
1237:
1234:
1230:
1223:
1221:
1218:
1214:
1210:
1206:
1203:
1199:
1195:
1188:
1186:
1180:
1176:
1173:
1167:
1165:
1161:
1157:
1153:
1149:
1143:
1139:
1136:
1132:
1125:
1123:
1117:
1113:
1110:
1104:
1102:
1098:
1094:
1090:
1086:
1080:
1076:
1073:
1071:RefSeq (mRNA)
1069:
1062:
1061:
1056:
1052:
1049:
1043:
1042:
1037:
1033:
1030:
1028:
1024:
1017:
1016:
1011:
1007:
1004:
998:
997:
992:
988:
985:
983:
979:
972:
971:
966:
962:
959:
953:
952:
947:
943:
940:
938:
934:
931:
928:
926:
923:
919:
916:
912:
908:
901:
897:
892:
886:
883:
881:
878:
876:
873:
871:
868:
866:
863:
861:
858:
856:
853:
851:
848:
846:
843:
841:
838:
836:
833:
831:
828:
826:
823:
821:
818:
816:
813:
811:
808:
806:
803:
801:
798:
796:
793:
792:
790:
787:
786:
780:
777:
775:
772:
770:
767:
765:
762:
760:
757:
755:
752:
750:
747:
745:
742:
740:
737:
735:
732:
730:
727:
726:
724:
721:
720:
714:
711:
709:
706:
704:
701:
699:
696:
694:
691:
689:
686:
684:
681:
679:
676:
674:
671:
669:
668:siRNA binding
666:
664:
661:
659:
656:
654:
651:
649:
646:
644:
641:
639:
636:
634:
631:
629:
626:
625:
623:
620:
619:
616:
615:Gene ontology
612:
608:
596:
591:
588:
586:
582:
574:
569:
558:
554:
550:
546:
542:
538:
534:
530:
526:
522:
521:
518:
514:
509:
506:
496:
492:
488:
484:
480:
476:
472:
468:
464:
460:
459:
456:
452:
447:
444:
443:
440:
438:
434:
432:
431:
427:
426:
423:
421:
417:
413:
409:
405:
397:
392:
388:
384:
379:
369:
365:
358:
351:
345:
338:
330:
326:
322:
317:
313:
308:
304:
296:
291:
287:
283:
278:
268:
264:
257:
250:
244:
237:
233:
227:
223:
219:
214:
210:
205:
201:
197:
193:
189:
185:
181:
177:
173:
169:
165:
161:
153:
148:
141:
136:
131:
120:
118:
114:
110:
106:
102:
98:
94:
90:
86:
82:
78:
72:
67:
64:
63:
59:
56:
49:
45:
40:
36:
32:
27:
22:
16:
4398:Translocases
4395:
4382:
4369:
4356:
4343:
4333:Transferases
4330:
4317:
4174:Binding site
3991:
3955:}}
3949:{{
3913:Endonuclease
3844:ribonuclease
3784:
3567:Nucleotidase
3488:Thioesterase
3273:
3252:
3203:
3199:
3193:
3150:
3146:
3136:
3104:(19): e149.
3101:
3097:
3087:
3075:. Retrieved
3071:
3062:
3035:
3031:
3021:
2996:
2992:
2986:
2974:. Retrieved
2970:the original
2960:
2925:
2921:
2911:
2876:
2873:FEBS Letters
2872:
2862:
2852:
2817:
2813:
2803:
2766:
2762:
2752:
2717:
2713:
2703:
2668:
2664:
2654:
2609:
2605:
2595:
2576:
2570:
2535:
2531:
2521:
2486:
2482:
2420:
2416:
2410:
2375:
2371:
2323:
2320:FEBS Letters
2319:
2313:
2288:
2284:
2278:
2264:
2223:
2219:
2175:
2171:
2131:
2127:
2113:
2078:
2072:
2024:
2020:
2014:
2005:
1996:
1987:
1978:
1875:
1867:
1863:
1860:
1855:
1851:
1837:
1835:
1802:
1798:
1779:
1764:
1759:Drosophila C
1758:
1750:
1747:yellow fever
1743:dengue fever
1722:
1720:
1691:
1671:
1643:
1626:
1617:alu elements
1599:Age related
1598:
1561:
1518:
1475:
1468:
1441:
1429:
1412:
1390:
1385:
1370:
1364:
1357:
1354:transfection
1340:PhD student
1335:
1322:endonuclease
1276:
1268:
1264:
1260:
1259:
1182:
1156:NP_001278557
1152:NP_001258211
1148:NP_001182502
1145:
1119:
1093:NM_001291628
1089:NM_001271282
1085:NM_001195573
1082:
1058:
1039:
1013:
994:
968:
949:
929:
924:
739:RISC complex
435:
428:
394:104,718,211
381:104,654,001
155:External IDs
74:
15:
4169:Active site
4088:Nuclease S1
3859:Exonuclease
3753:Lecithinase
3582:Calcineurin
3520:Phosphatase
3426:Lipoprotein
3416:Endothelial
1876:Arabidopsis
1872:homogeneous
1868:Arabidopsis
1864:Arabidopsis
1856:Arabidopsis
1852:Arabidopsis
1735:arboviruses
1694:RNA viruses
1508:translation
1506:to prevent
1338:Stony Brook
749:ARC complex
744:growth cone
708:DNA binding
698:ATP binding
673:RNA binding
489:oral cavity
469:optic nerve
293:95,158,010
280:95,086,228
133:Identifiers
4441:Categories
4372:Isomerases
4346:Hydrolases
4213:Regulation
3401:Pancreatic
3338:Carboxylic
1973:, May 2017
1952:, May 2017
1929:References
1844:cis-acting
1805:antibodies
1731:mosquitoes
1723:Drosophila
1717:In insects
1658:transposon
1646:DNA repair
1476:Drosophila
1381:epigenetic
1359:Drosophila
1348:'s lab at
1302:base pairs
439:(ortholog)
176:HomoloGene
4467:EC 3.1.26
4251:EC number
3978:RNase III
3836:(includes
3777:Sulfatase
3690:Autotaxin
3554:Prostatic
3406:Lysosomal
3321:esterases
3317:Hydrolase
2999:: 19–28.
2172:Structure
1770:ascovirus
1727:antiviral
1706:influenza
1682:mutations
1662:oncogenic
1650:chromatin
1530:inhibits
1515:Micro RNA
1462:/Zwille)
1460:Argonaute
1388:protozoan
1367:orthologs
1332:Discovery
1318:Argonaute
1286:RNase III
1185:NP_683750
1164:NP_803187
1160:NP_085124
1122:NM_148948
1101:NM_177438
1097:NM_030621
915:Orthologs
729:cytoplasm
184:GeneCards
4462:MicroRNA
4275:Kinetics
4199:Cofactor
4162:Activity
4072:RNase T1
3834:Nuclease
3469:Cutinase
3228:28801338
3220:19221817
3185:19536157
3128:15520458
3077:23 April
3054:18818708
3013:24732439
2976:22 April
2952:21419681
2903:16563388
2844:17071602
2795:23222681
2744:22541070
2695:21412326
2646:12902540
2562:15811921
2513:20179193
2445:10749213
2402:21419681
2348:14495726
2340:16214139
2305:10786632
2256:23785494
2248:16410517
2202:19836333
2150:20085706
2105:18268840
2049:11201747
1969:–
1948:–
1903:microRNA
1882:See also
1870:is more
1710:vaccinia
1698:immunity
1627:Altered
1496:microRNA
1452:helicase
1328:(mRNA).
1298:microRNA
1271:, is an
1233:Wikidata
894:Sources:
764:dendrite
272:14q32.13
4431:Biology
4385:Ligases
4155:Enzymes
4045:RNase E
4040:RNase Z
4035:RNase A
4030:RNase P
4003:RNase H
3621:Phytase
3421:Hepatic
3396:Lingual
3392:Gastric
3280:PDBe-KB
3270:UniProt
3259:PDBe-KB
3249:UniProt
3176:2733940
3155:Bibcode
2943:3115569
2894:7094296
2835:1780192
2786:3546719
2735:3351582
2673:Bibcode
2614:Bibcode
2553:1370754
2504:3170915
2453:9091863
2425:Bibcode
2393:3115569
2228:Bibcode
2220:Science
2193:2880462
2120:Reski R
2057:4371481
2029:Bibcode
1971:Ensembl
1950:Ensembl
1814:ligands
1590:Disease
1540:nucleus
1027:UniProt
982:Ensembl
921:Species
900:QuickGO
769:nucleus
734:cytosol
414:pattern
172:2177178
140:Aliases
4417:Portal
4359:Lyases
3983:Drosha
3908:3.1.21
3876:RecBCD
3854:3.1.11
3474:PETase
3382:Lipase
3275:Q8R418
3254:Q9UPY3
3226:
3218:
3183:
3173:
3147:Nature
3126:
3119:528817
3116:
3052:
3011:
2950:
2940:
2901:
2891:
2842:
2832:
2793:
2783:
2742:
2732:
2693:
2665:Nature
2644:
2637:187842
2634:
2583:
2560:
2550:
2511:
2501:
2451:
2443:
2417:Nature
2400:
2390:
2346:
2338:
2303:
2254:
2246:
2200:
2190:
2148:
2103:
2093:
2055:
2047:
2021:Nature
1913:Drosha
1786:cancer
1782:tumors
1708:, and
1674:goiter
1638:drosha
1623:Cancer
1609:Drosha
1575:miRNAs
1571:siRNAs
1552:Drosha
1464:domain
1444:DICER1
1379:, the
1365:Dicer
1306:3′-end
1278:DICER1
1273:enzyme
1219:search
1217:PubMed
1060:Q8R418
1041:Q9UPY3
970:192119
937:Entrez
585:BioGPS
523:zygote
188:DICER1
164:606241
147:DICER1
24:DICER1
4311:Types
3992:Dicer
3947:;see
3773:3.1.6
3743:PDE4B
3739:PDE4A
3677:3.1.4
3646:IMPA3
3642:IMPA2
3638:IMPA1
3516:3.1.3
3484:3.1.2
3334:3.1.1
3224:S2CID
2449:S2CID
2344:S2CID
2252:S2CID
2053:S2CID
1761:virus
1702:HIV-1
1676:with
1629:miRNA
1613:Pasha
1548:DGCR8
1528:miRNA
1436:RNase
1261:Dicer
951:23405
930:Mouse
925:Human
896:Amigo
437:Mouse
430:Human
377:Start
312:Mouse
276:Start
209:Human
180:13251
4403:list
4396:EC7
4390:list
4383:EC6
4377:list
4370:EC5
4364:list
4357:EC4
4351:list
4344:EC3
4338:list
4331:EC2
4325:list
4318:EC1
3910:-31:
3856:-16:
3842:and
3748:PDE5
3734:PDE3
3729:PDE2
3724:PDE1
3616:PTEN
3601:OCRL
3594:PP2A
3543:ALPP
3538:ALPL
3533:ALPI
3327:3.1)
3268:for
3247:for
3216:PMID
3181:PMID
3124:PMID
3079:2014
3050:PMID
3009:PMID
2978:2014
2948:PMID
2899:PMID
2840:PMID
2791:PMID
2740:PMID
2714:Cell
2691:PMID
2642:PMID
2581:ISBN
2558:PMID
2509:PMID
2441:PMID
2398:PMID
2336:PMID
2301:PMID
2289:1490
2244:PMID
2198:PMID
2146:PMID
2128:Cell
2101:PMID
2091:ISBN
2045:PMID
1923:mRNA
1893:RISC
1745:and
1611:and
1573:and
1536:cell
1456:Piwi
1296:and
1282:gene
759:axon
420:Bgee
368:Band
329:Chr.
267:Band
226:Chr.
160:OMIM
117:4WYQ
113:4NHA
109:4NH6
105:4NH5
101:4NH3
97:4NGG
93:4NGF
89:4NGD
85:4NGC
81:4NGB
77:2EB1
58:RCSB
55:PDBe
4065:4/5
3266:PDB
3245:PDB
3208:doi
3171:PMC
3163:doi
3151:460
3114:PMC
3106:doi
3040:doi
3001:doi
2938:PMC
2930:doi
2889:PMC
2881:doi
2877:580
2830:PMC
2822:doi
2818:169
2781:PMC
2771:doi
2730:PMC
2722:doi
2718:149
2681:doi
2669:471
2632:PMC
2622:doi
2610:100
2548:PMC
2540:doi
2532:RNA
2499:PMC
2491:doi
2433:doi
2421:404
2388:PMC
2380:doi
2328:doi
2324:579
2293:doi
2236:doi
2224:311
2188:PMC
2180:doi
2136:doi
2132:140
2083:doi
2037:doi
2025:409
1816:or
1807:or
1721:In
1524:RNA
1494:or
1409:kDa
1267:or
593:n/a
390:End
289:End
192:OMA
168:MGI
48:PDB
4443::
4023:2C
4018:2B
4013:2A
3994::
3985::
3775::
3655::
3644:,
3640:,
3556:)/
3518::
3486::
3455:A2
3450:A1
3336::
3325:EC
3319::
3272::
3251::
3222:.
3214:.
3202:.
3179:.
3169:.
3161:.
3149:.
3145:.
3122:.
3112:.
3102:32
3100:.
3096:.
3070:.
3048:.
3036:16
3034:.
3030:.
3007:.
2995:.
2946:.
2936:.
2926:42
2924:.
2920:.
2897:.
2887:.
2875:.
2871:.
2838:.
2828:.
2816:.
2812:.
2789:.
2779:.
2767:13
2765:.
2761:.
2738:.
2728:.
2716:.
2712:.
2689:.
2679:.
2667:.
2663:.
2640:.
2630:.
2620:.
2608:.
2604:.
2556:.
2546:.
2536:11
2534:.
2530:.
2507:.
2497:.
2487:70
2485:.
2481:.
2461:^
2447:.
2439:.
2431:.
2419:.
2396:.
2386:.
2376:42
2374:.
2370:.
2356:^
2342:.
2334:.
2322:.
2299:.
2287:.
2250:.
2242:.
2234:.
2222:.
2210:^
2196:.
2186:.
2176:17
2174:.
2170:.
2158:^
2144:.
2130:.
2126:.
2099:.
2089:.
2051:.
2043:.
2035:.
2023:.
2004:.
1986:.
1957:^
1936:^
1741:,
1737::
1704:,
898:/
396:bp
383:bp
295:bp
282:bp
190:;
186::
182:;
178::
174:;
170::
166:;
162::
115:,
111:,
107:,
103:,
99:,
95:,
91:,
87:,
83:,
79:,
4419::
4405:)
4401:(
4392:)
4388:(
4379:)
4375:(
4366:)
4362:(
4353:)
4349:(
4340:)
4336:(
4327:)
4323:(
4147:e
4140:t
4133:v
4060:3
4055:2
4050:1
4008:1
3846:)
3759:)
3755:(
3741:/
3717:1
3705:D
3700:C
3679::
3560:/
3552:(
3460:B
3394:/
3323:(
3309:e
3302:t
3295:v
3282:.
3261:.
3230:.
3210::
3204:9
3187:.
3165::
3157::
3130:.
3108::
3081:.
3056:.
3042::
3015:.
3003::
2997:7
2980:.
2954:.
2932::
2905:.
2883::
2846:.
2824::
2797:.
2773::
2746:.
2724::
2697:.
2683::
2675::
2648:.
2624::
2616::
2589:.
2564:.
2542::
2515:.
2493::
2455:.
2435::
2427::
2404:.
2382::
2350:.
2330::
2307:.
2295::
2272:.
2258:.
2238::
2230::
2204:.
2182::
2152:.
2138::
2107:.
2085::
2059:.
2039::
2031::
2008:.
1990:.
1510:.
1458:/
314:)
211:)
194::
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.